Yuichiro Hayashi
WSI-INR: Implicit Neural Representations for Lesion Segmentation in Whole-Slide Images
Mar 04, 2026Abstract:Whole-slide images (WSIs) are fundamental for computational pathology, where accurate lesion segmentation is critical for clinical decision making. Existing methods partition WSIs into discrete patches, disrupting spatial continuity and treating multi-resolution views as independent samples, which leads to spatially fragmented segmentation and reduced robustness to resolution variations. To address the issues, we propose WSI-INR, a novel patch-free framework based on Implicit Neural Representations (INRs). WSI-INR models the WSI as a continuous implicit function mapping spatial coordinates directly to tissue semantics features, outputting segmentation results while preserving intrinsic spatial information across the entire slide. In the WSI-INR, we incorporate multi-resolution hash grid encoding to regard different resolution levels as varying sampling densities of the same continuous tissue, achieving a consistent feature representation across resolutions. In addition, by jointly training a shared INR decoder, WSI-INR can capture general priors across different cases. Experimental results showed that WSI-INR maintains robust segmentation performance across resolutions; at Base/4, our resolution-specific optimization improves Dice score by +26.11%, while U-Net and TransUNet decrease by 54.28% and 36.18%, respectively. Crucially, this work enables INRs to segment highly heterogeneous pathological lesions beyond structurally consistent anatomical tissues, offering a fresh perspective for pathological analysis.
A Bayesian Approach to Weakly-supervised Laparoscopic Image Segmentation
Oct 11, 2024Abstract:In this paper, we study weakly-supervised laparoscopic image segmentation with sparse annotations. We introduce a novel Bayesian deep learning approach designed to enhance both the accuracy and interpretability of the model's segmentation, founded upon a comprehensive Bayesian framework, ensuring a robust and theoretically validated method. Our approach diverges from conventional methods that directly train using observed images and their corresponding weak annotations. Instead, we estimate the joint distribution of both images and labels given the acquired data. This facilitates the sampling of images and their high-quality pseudo-labels, enabling the training of a generalizable segmentation model. Each component of our model is expressed through probabilistic formulations, providing a coherent and interpretable structure. This probabilistic nature benefits accurate and practical learning from sparse annotations and equips our model with the ability to quantify uncertainty. Extensive evaluations with two public laparoscopic datasets demonstrated the efficacy of our method, which consistently outperformed existing methods. Furthermore, our method was adapted for scribble-supervised cardiac multi-structure segmentation, presenting competitive performance compared to previous methods. The code is available at https://github.com/MoriLabNU/Bayesian_WSS.
Identifying Suspicious Regions of Covid-19 by Abnormality-Sensitive Activation Mapping
Mar 27, 2023Abstract:This paper presents a fully-automated method for the identification of suspicious regions of a coronavirus disease (COVID-19) on chest CT volumes. One major role of chest CT scanning in COVID-19 diagnoses is identification of an inflammation particular to the disease. This task is generally performed by radiologists through an interpretation of the CT volumes, however, because of the heavy workload, an automatic analysis method using a computer is desired. Most computer-aided diagnosis studies have addressed only a portion of the elements necessary for the identification. In this work, we realize the identification method through a classification task by using a 2.5-dimensional CNN with three-dimensional attention mechanisms. We visualize the suspicious regions by applying a backpropagation based on positive gradients to attention-weighted features. We perform experiments on an in-house dataset and two public datasets to reveal the generalization ability of the proposed method. The proposed architecture achieved AUCs of over 0.900 for all the datasets, and mean sensitivity $0.853 \pm 0.036$ and specificity $0.870 \pm 0.040$. The method can also identify notable lesions pointed out in the radiology report as suspicious regions.
Semi-automated Virtual Unfolded View Generation Method of Stomach from CT Volumes
Jan 14, 2022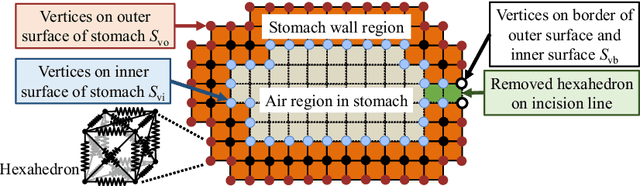
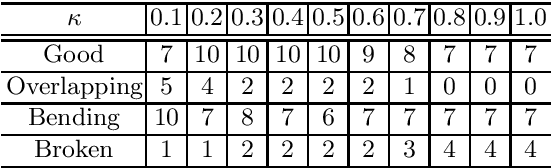
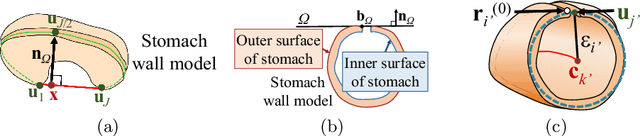

Abstract:CT image-based diagnosis of the stomach is developed as a new way of diagnostic method. A virtual unfolded (VU) view is suitable for displaying its wall. In this paper, we propose a semi-automated method for generating VU views of the stomach. Our method requires minimum manual operations. The determination of the unfolding forces and the termination of the unfolding process are automated. The unfolded shape of the stomach is estimated based on its radius. The unfolding forces are determined so that the stomach wall is deformed to the expected shape. The iterative deformation process is terminated if the difference of the shapes between the deformed shape and expected shape is small. Our experiments using 67 CT volumes showed that our proposed method can generate good VU views for 76.1% cases.
* Accepted paper as a poster presentation at MICCAI 2013 (International Conference on Medical Image Computing and Computer-Assisted Intervention), Nagoya, Japan
COVID-19 Infection Segmentation from Chest CT Images Based on Scale Uncertainty
Jan 09, 2022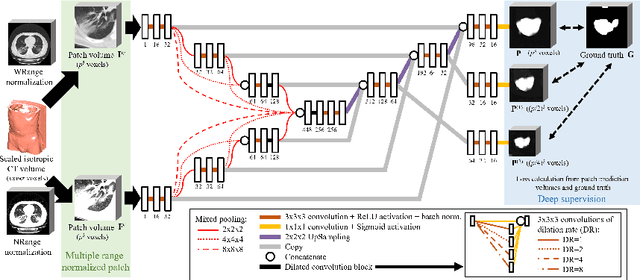
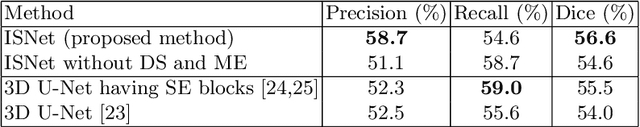
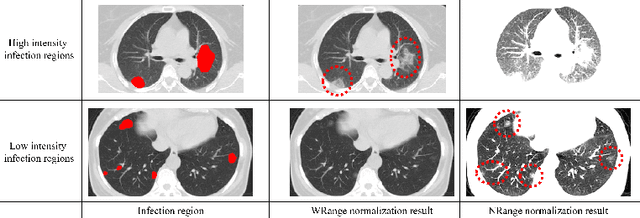

Abstract:This paper proposes a segmentation method of infection regions in the lung from CT volumes of COVID-19 patients. COVID-19 spread worldwide, causing many infected patients and deaths. CT image-based diagnosis of COVID-19 can provide quick and accurate diagnosis results. An automated segmentation method of infection regions in the lung provides a quantitative criterion for diagnosis. Previous methods employ whole 2D image or 3D volume-based processes. Infection regions have a considerable variation in their sizes. Such processes easily miss small infection regions. Patch-based process is effective for segmenting small targets. However, selecting the appropriate patch size is difficult in infection region segmentation. We utilize the scale uncertainty among various receptive field sizes of a segmentation FCN to obtain infection regions. The receptive field sizes can be defined as the patch size and the resolution of volumes where patches are clipped from. This paper proposes an infection segmentation network (ISNet) that performs patch-based segmentation and a scale uncertainty-aware prediction aggregation method that refines the segmentation result. We design ISNet to segment infection regions that have various intensity values. ISNet has multiple encoding paths to process patch volumes normalized by multiple intensity ranges. We collect prediction results generated by ISNets having various receptive field sizes. Scale uncertainty among the prediction results is extracted by the prediction aggregation method. We use an aggregation FCN to generate a refined segmentation result considering scale uncertainty among the predictions. In our experiments using 199 chest CT volumes of COVID-19 cases, the prediction aggregation method improved the dice similarity score from 47.6% to 62.1%.
* Accepted paper as a oral presentation at CILP2021, 10th MICCAI CLIP Workshop
Lung infection and normal region segmentation from CT volumes of COVID-19 cases
Jan 09, 2022Abstract:This paper proposes an automated segmentation method of infection and normal regions in the lung from CT volumes of COVID-19 patients. From December 2019, novel coronavirus disease 2019 (COVID-19) spreads over the world and giving significant impacts to our economic activities and daily lives. To diagnose the large number of infected patients, diagnosis assistance by computers is needed. Chest CT is effective for diagnosis of viral pneumonia including COVID-19. A quantitative analysis method of condition of the lung from CT volumes by computers is required for diagnosis assistance of COVID-19. This paper proposes an automated segmentation method of infection and normal regions in the lung from CT volumes using a COVID-19 segmentation fully convolutional network (FCN). In diagnosis of lung diseases including COVID-19, analysis of conditions of normal and infection regions in the lung is important. Our method recognizes and segments lung normal and infection regions in CT volumes. To segment infection regions that have various shapes and sizes, we introduced dense pooling connections and dilated convolutions in our FCN. We applied the proposed method to CT volumes of COVID-19 cases. From mild to severe cases of COVID-19, the proposed method correctly segmented normal and infection regions in the lung. Dice scores of normal and infection regions were 0.911 and 0.753, respectively.
* Accepted paper as a poster presentation at SPIE Medical Imaging 2021
Precise Estimation of Renal Vascular Dominant Regions Using Spatially Aware Fully Convolutional Networks, Tensor-Cut and Voronoi Diagrams
Aug 05, 2019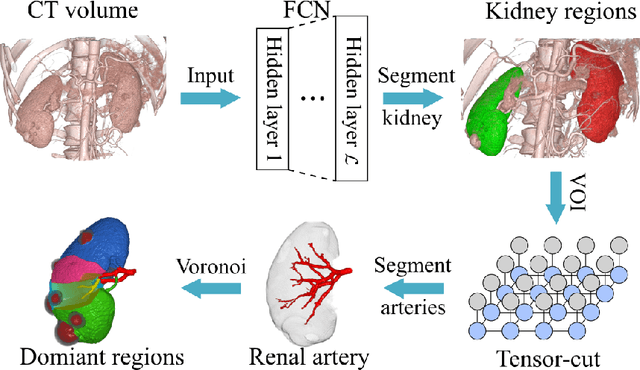
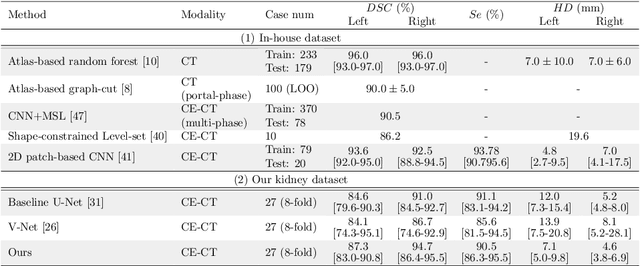
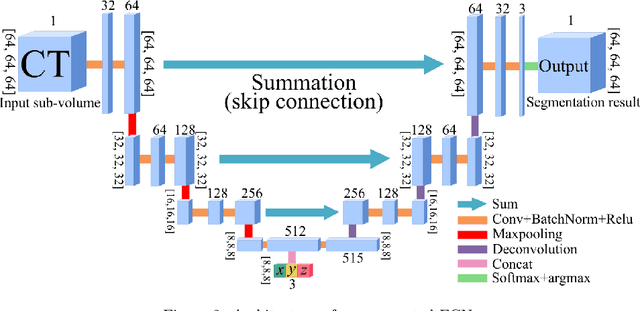
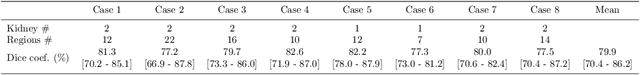
Abstract:This paper presents a new approach for precisely estimating the renal vascular dominant region using a Voronoi diagram. To provide computer-assisted diagnostics for the pre-surgical simulation of partial nephrectomy surgery, we must obtain information on the renal arteries and the renal vascular dominant regions. We propose a fully automatic segmentation method that combines a neural network and tensor-based graph-cut methods to precisely extract the kidney and renal arteries. First, we use a convolutional neural network to localize the kidney regions and extract tiny renal arteries with a tensor-based graph-cut method. Then we generate a Voronoi diagram to estimate the renal vascular dominant regions based on the segmented kidney and renal arteries. The accuracy of kidney segmentation in 27 cases with 8-fold cross validation reached a Dice score of 95%. The accuracy of renal artery segmentation in 8 cases obtained a centerline overlap ratio of 80%. Each partition region corresponds to a renal vascular dominant region. The final dominant-region estimation accuracy achieved a Dice coefficient of 80%. A clinical application showed the potential of our proposed estimation approach in a real clinical surgical environment. Further validation using large-scale database is our future work.
A multi-scale pyramid of 3D fully convolutional networks for abdominal multi-organ segmentation
Jun 06, 2018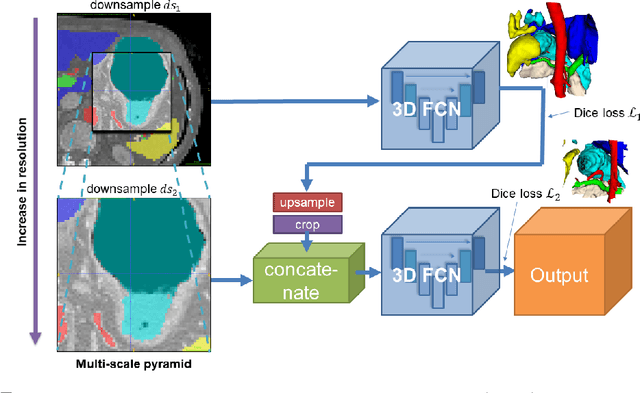
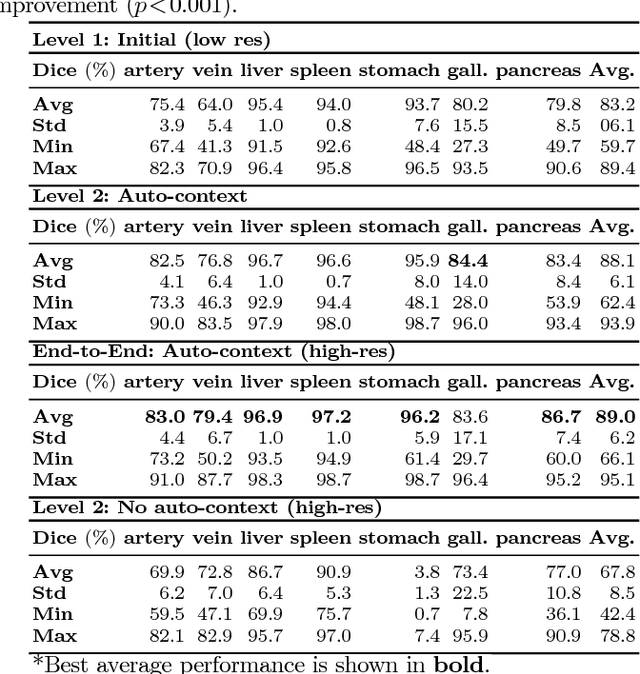
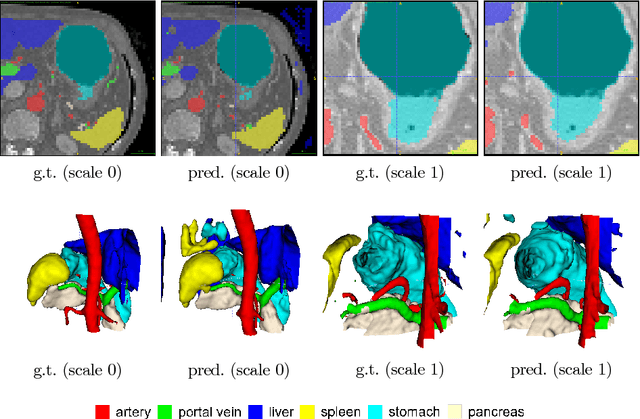
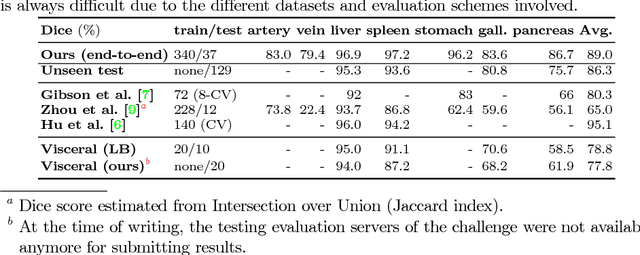
Abstract:Recent advances in deep learning, like 3D fully convolutional networks (FCNs), have improved the state-of-the-art in dense semantic segmentation of medical images. However, most network architectures require severely downsampling or cropping the images to meet the memory limitations of today's GPU cards while still considering enough context in the images for accurate segmentation. In this work, we propose a novel approach that utilizes auto-context to perform semantic segmentation at higher resolutions in a multi-scale pyramid of stacked 3D FCNs. We train and validate our models on a dataset of manually annotated abdominal organs and vessels from 377 clinical CT images used in gastric surgery, and achieve promising results with close to 90% Dice score on average. For additional evaluation, we perform separate testing on datasets from different sources and achieve competitive results, illustrating the robustness of the model and approach.
Deep learning and its application to medical image segmentation
Mar 23, 2018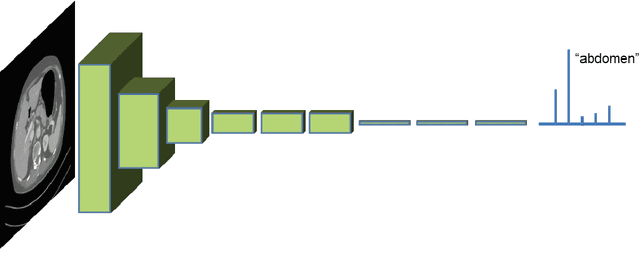
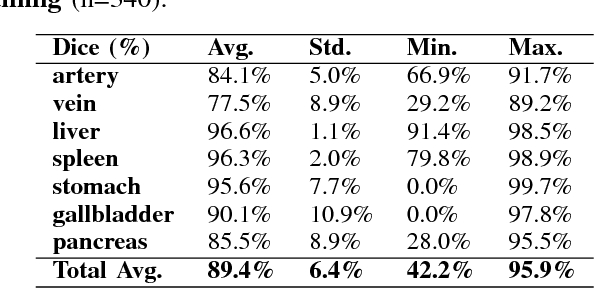
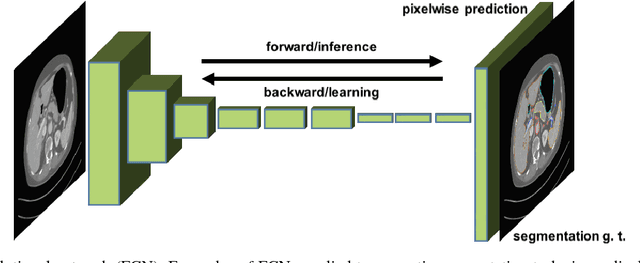
Abstract:One of the most common tasks in medical imaging is semantic segmentation. Achieving this segmentation automatically has been an active area of research, but the task has been proven very challenging due to the large variation of anatomy across different patients. However, recent advances in deep learning have made it possible to significantly improve the performance of image recognition and semantic segmentation methods in the field of computer vision. Due to the data driven approaches of hierarchical feature learning in deep learning frameworks, these advances can be translated to medical images without much difficulty. Several variations of deep convolutional neural networks have been successfully applied to medical images. Especially fully convolutional architectures have been proven efficient for segmentation of 3D medical images. In this article, we describe how to build a 3D fully convolutional network (FCN) that can process 3D images in order to produce automatic semantic segmentations. The model is trained and evaluated on a clinical computed tomography (CT) dataset and shows state-of-the-art performance in multi-organ segmentation.
* Accepted for publication in the journal of the Japanese Society of Medical Imaging Technology (JAMIT)
An application of cascaded 3D fully convolutional networks for medical image segmentation
Mar 20, 2018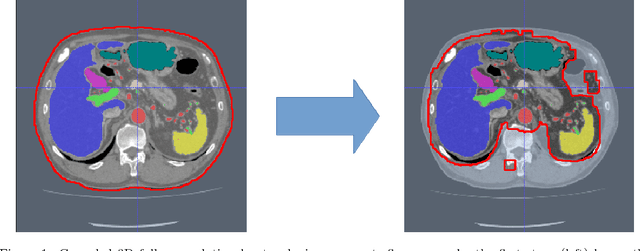
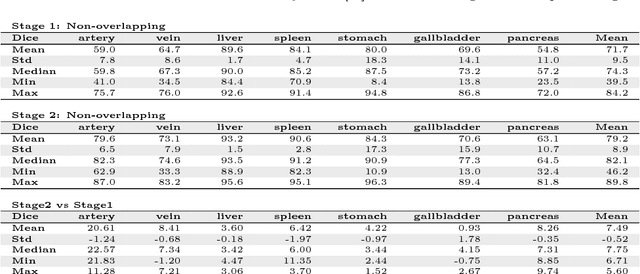
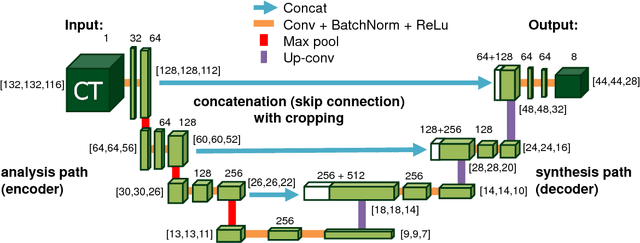
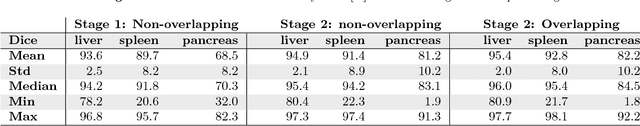
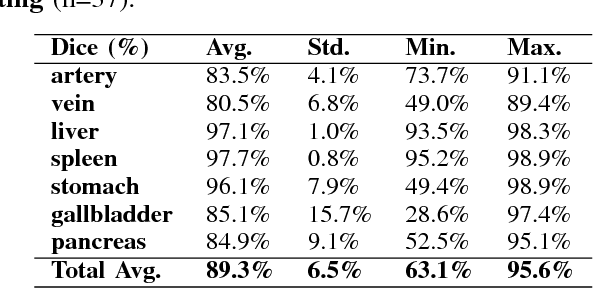
Abstract:Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: https://github.com/holgerroth/3Dunet_abdomen_cascade.
* Preprint accepted for publication in Computerized Medical Imaging and Graphics. Substantial extension of arXiv:1704.06382; Corrected references to figure numbers in this version
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge