Shigeki Aoki
Department of Radiology, Juntendo University School of Medicine
Parameter-Efficient Fine-Tuning of 3D DDPM for MRI Image Generation Using Tensor Networks
Jul 24, 2025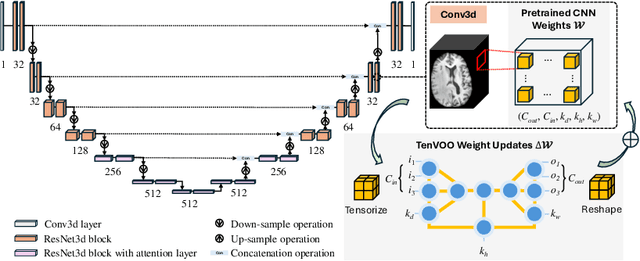

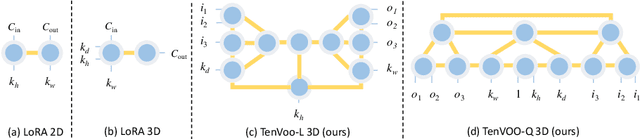

Abstract:We address the challenge of parameter-efficient fine-tuning (PEFT) for three-dimensional (3D) U-Net-based denoising diffusion probabilistic models (DDPMs) in magnetic resonance imaging (MRI) image generation. Despite its practical significance, research on parameter-efficient representations of 3D convolution operations remains limited. To bridge this gap, we propose Tensor Volumetric Operator (TenVOO), a novel PEFT method specifically designed for fine-tuning DDPMs with 3D convolutional backbones. Leveraging tensor network modeling, TenVOO represents 3D convolution kernels with lower-dimensional tensors, effectively capturing complex spatial dependencies during fine-tuning with few parameters. We evaluate TenVOO on three downstream brain MRI datasets-ADNI, PPMI, and BraTS2021-by fine-tuning a DDPM pretrained on 59,830 T1-weighted brain MRI scans from the UK Biobank. Our results demonstrate that TenVOO achieves state-of-the-art performance in multi-scale structural similarity index measure (MS-SSIM), outperforming existing approaches in capturing spatial dependencies while requiring only 0.3% of the trainable parameters of the original model. Our code is available at: https://github.com/xiaovhua/tenvoo
Identifying Suspicious Regions of Covid-19 by Abnormality-Sensitive Activation Mapping
Mar 27, 2023Abstract:This paper presents a fully-automated method for the identification of suspicious regions of a coronavirus disease (COVID-19) on chest CT volumes. One major role of chest CT scanning in COVID-19 diagnoses is identification of an inflammation particular to the disease. This task is generally performed by radiologists through an interpretation of the CT volumes, however, because of the heavy workload, an automatic analysis method using a computer is desired. Most computer-aided diagnosis studies have addressed only a portion of the elements necessary for the identification. In this work, we realize the identification method through a classification task by using a 2.5-dimensional CNN with three-dimensional attention mechanisms. We visualize the suspicious regions by applying a backpropagation based on positive gradients to attention-weighted features. We perform experiments on an in-house dataset and two public datasets to reveal the generalization ability of the proposed method. The proposed architecture achieved AUCs of over 0.900 for all the datasets, and mean sensitivity $0.853 \pm 0.036$ and specificity $0.870 \pm 0.040$. The method can also identify notable lesions pointed out in the radiology report as suspicious regions.
COVID-19 Infection Segmentation from Chest CT Images Based on Scale Uncertainty
Jan 09, 2022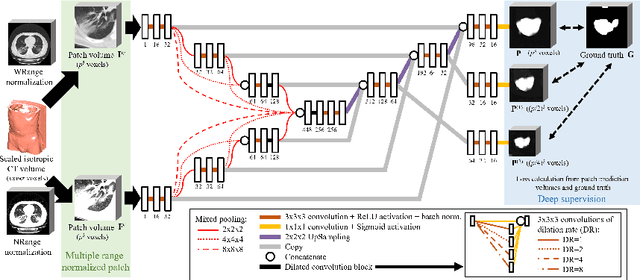
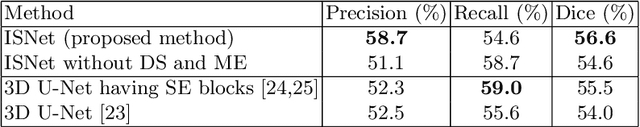
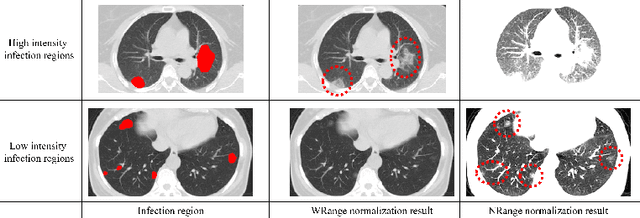

Abstract:This paper proposes a segmentation method of infection regions in the lung from CT volumes of COVID-19 patients. COVID-19 spread worldwide, causing many infected patients and deaths. CT image-based diagnosis of COVID-19 can provide quick and accurate diagnosis results. An automated segmentation method of infection regions in the lung provides a quantitative criterion for diagnosis. Previous methods employ whole 2D image or 3D volume-based processes. Infection regions have a considerable variation in their sizes. Such processes easily miss small infection regions. Patch-based process is effective for segmenting small targets. However, selecting the appropriate patch size is difficult in infection region segmentation. We utilize the scale uncertainty among various receptive field sizes of a segmentation FCN to obtain infection regions. The receptive field sizes can be defined as the patch size and the resolution of volumes where patches are clipped from. This paper proposes an infection segmentation network (ISNet) that performs patch-based segmentation and a scale uncertainty-aware prediction aggregation method that refines the segmentation result. We design ISNet to segment infection regions that have various intensity values. ISNet has multiple encoding paths to process patch volumes normalized by multiple intensity ranges. We collect prediction results generated by ISNets having various receptive field sizes. Scale uncertainty among the prediction results is extracted by the prediction aggregation method. We use an aggregation FCN to generate a refined segmentation result considering scale uncertainty among the predictions. In our experiments using 199 chest CT volumes of COVID-19 cases, the prediction aggregation method improved the dice similarity score from 47.6% to 62.1%.
* Accepted paper as a oral presentation at CILP2021, 10th MICCAI CLIP Workshop
The utility of a convolutional neural network for generating a myelin volume index map from rapid simultaneous relaxometry imaging
Apr 24, 2019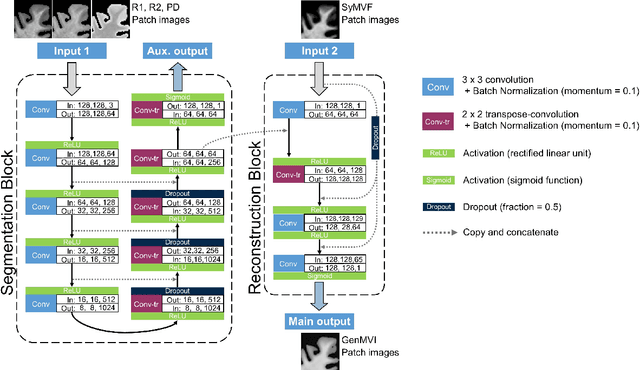
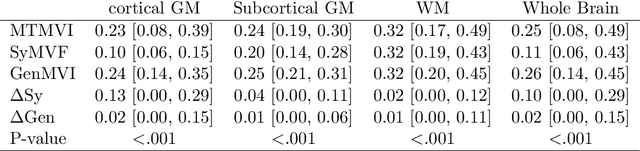
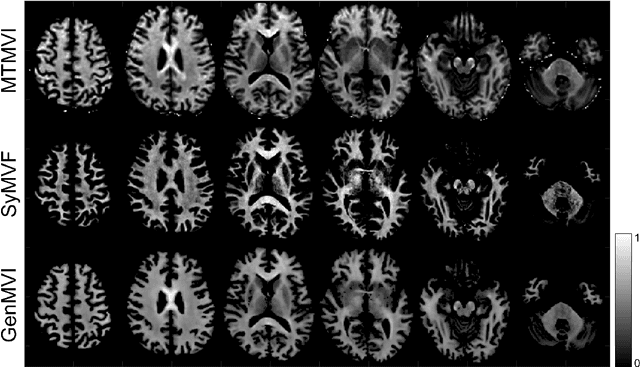
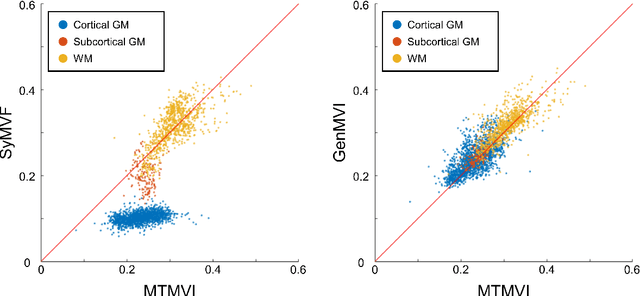
Abstract:Background and Purpose: A current algorithm to obtain a synthetic myelin volume fraction map (SyMVF) from rapid simultaneous relaxometry imaging (RSRI) has a potential problem, that it does not incorporate information from surrounding pixels. The purpose of this study was to develop a method that utilizes a convolutional neural network (CNN) to overcome this problem. Methods: RSRI and magnetization transfer images from 20 healthy volunteers were included. A CNN was trained to reconstruct RSRI-related metric maps into a myelin volume-related index (generated myelin volume index: GenMVI) map using the myelin volume index map calculated from magnetization transfer images (MTMVI) as reference. The SyMVF and GenMVI maps were statistically compared by testing how well they correlated with the MTMVI map. The correlations were evaluated based on: (i) averaged values obtained from 164 atlas-based ROIs, and (ii) pixel-based comparison for ROIs defined in four different tissue types (cortical and subcortical gray matter, white matter, and whole brain). Results: For atlas-based ROIs, the overall correlation with the MTMVI map was higher for the GenMVI map than for the SyMVF map. In the pixel-based comparison, correlation with the MTMVI map was stronger for the GenMVI map than for the SyMVF map, and the difference in the distribution for the volunteers was significant (Wilcoxon sign-rank test, P<.001) in all tissue types. Conclusion: The proposed method is useful, as it can incorporate more specific information about local tissue properties than the existing method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge