Yuan Long
Reproducibility Assessment of Magnetic Resonance Spectroscopy of Pregenual Anterior Cingulate Cortex across Sessions and Vendors via the Cloud Computing Platform CloudBrain-MRS
Mar 06, 2025



Abstract:Given the need to elucidate the mechanisms underlying illnesses and their treatment, as well as the lack of harmonization of acquisition and post-processing protocols among different magnetic resonance system vendors, this work is to determine if metabolite concentrations obtained from different sessions, machine models and even different vendors of 3 T scanners can be highly reproducible and be pooled for diagnostic analysis, which is very valuable for the research of rare diseases. Participants underwent magnetic resonance imaging (MRI) scanning once on two separate days within one week (one session per day, each session including two proton magnetic resonance spectroscopy (1H-MRS) scans with no more than a 5-minute interval between scans (no off-bed activity)) on each machine. were analyzed for reliability of within- and between- sessions using the coefficient of variation (CV) and intraclass correlation coefficient (ICC), and for reproducibility of across the machines using correlation coefficient. As for within- and between- session, all CV values for a group of all the first or second scans of a session, or for a session were almost below 20%, and most of the ICCs for metabolites range from moderate (0.4-0.59) to excellent (0.75-1), indicating high data reliability. When it comes to the reproducibility across the three scanners, all Pearson correlation coefficients across the three machines approached 1 with most around 0.9, and majority demonstrated statistical significance (P<0.01). Additionally, the intra-vendor reproducibility was greater than the inter-vendor ones.
An artificially intelligent magnetic resonance spectroscopy quantification method: Comparison between QNet and LCModel on the cloud computing platform CloudBrain-MRS
Mar 06, 2025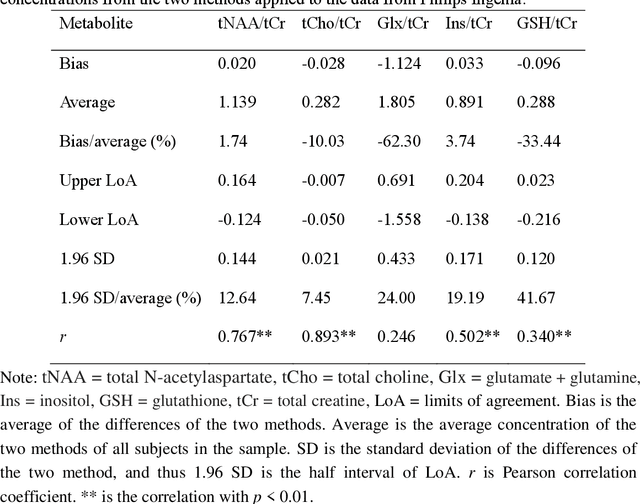

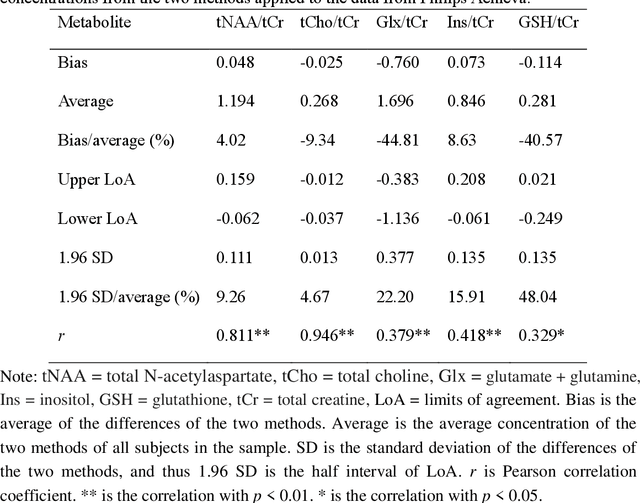
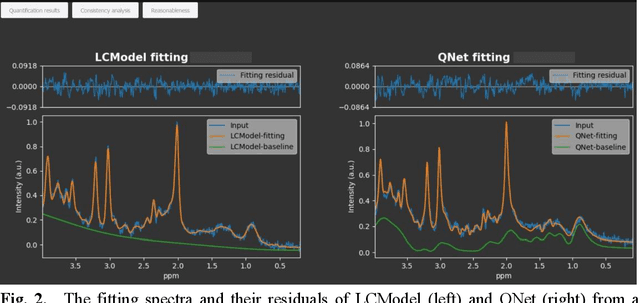
Abstract:Objctives: This work aimed to statistically compare the metabolite quantification of human brain magnetic resonance spectroscopy (MRS) between the deep learning method QNet and the classical method LCModel through an easy-to-use intelligent cloud computing platform CloudBrain-MRS. Materials and Methods: In this retrospective study, two 3 T MRI scanners Philips Ingenia and Achieva collected 61 and 46 in vivo 1H magnetic resonance (MR) spectra of healthy participants, respectively, from the brain region of pregenual anterior cingulate cortex from September to October 2021. The analyses of Bland-Altman, Pearson correlation and reasonability were performed to assess the degree of agreement, linear correlation and reasonability between the two quantification methods. Results: Fifteen healthy volunteers (12 females and 3 males, age range: 21-35 years, mean age/standard deviation = 27.4/3.9 years) were recruited. The analyses of Bland-Altman, Pearson correlation and reasonability showed high to good consistency and very strong to moderate correlation between the two methods for quantification of total N-acetylaspartate (tNAA), total choline (tCho), and inositol (Ins) (relative half interval of limits of agreement = 3.04%, 9.3%, and 18.5%, respectively; Pearson correlation coefficient r = 0.775, 0.927, and 0.469, respectively). In addition, quantification results of QNet are more likely to be closer to the previous reported average values than those of LCModel. Conclusion: There were high or good degrees of consistency between the quantification results of QNet and LCModel for tNAA, tCho, and Ins, and QNet generally has more reasonable quantification than LCModel.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge