Xinheng Lyu
PathMem: Toward Cognition-Aligned Memory Transformation for Pathology MLLMs
Mar 10, 2026Abstract:Computational pathology demands both visual pattern recognition and dynamic integration of structured domain knowledge, including taxonomy, grading criteria, and clinical evidence. In practice, diagnostic reasoning requires linking morphological evidence with formal diagnostic and grading criteria. Although multimodal large language models (MLLMs) demonstrate strong vision language reasoning capabilities, they lack explicit mechanisms for structured knowledge integration and interpretable memory control. As a result, existing models struggle to consistently incorporate pathology-specific diagnostic standards during reasoning. Inspired by the hierarchical memory process of human pathologists, we propose PathMem, a memory-centric multimodal framework for pathology MLLMs. PathMem organizes structured pathology knowledge as a long-term memory (LTM) and introduces a Memory Transformer that models the dynamic transition from LTM to working memory (WM) through multimodal memory activation and context-aware knowledge grounding, enabling context-aware memory refinement for downstream reasoning. PathMem achieves SOTA performance across benchmarks, improving WSI-Bench report generation (12.8% WSI-Precision, 10.1% WSI-Relevance) and open-ended diagnosis by 9.7% and 8.9% over prior WSI-based models.
HySurvPred: Multimodal Hyperbolic Embedding with Angle-Aware Hierarchical Contrastive Learning and Uncertainty Constraints for Survival Prediction
Mar 18, 2025Abstract:Multimodal learning that integrates histopathology images and genomic data holds great promise for cancer survival prediction. However, existing methods face key limitations: 1) They rely on multimodal mapping and metrics in Euclidean space, which cannot fully capture the hierarchical structures in histopathology (among patches from different resolutions) and genomics data (from genes to pathways). 2) They discretize survival time into independent risk intervals, which ignores its continuous and ordinal nature and fails to achieve effective optimization. 3) They treat censorship as a binary indicator, excluding censored samples from model optimization and not making full use of them. To address these challenges, we propose HySurvPred, a novel framework for survival prediction that integrates three key modules: Multimodal Hyperbolic Mapping (MHM), Angle-aware Ranking-based Contrastive Loss (ARCL) and Censor-Conditioned Uncertainty Constraint (CUC). Instead of relying on Euclidean space, we design the MHM module to explore the inherent hierarchical structures within each modality in hyperbolic space. To better integrate multimodal features in hyperbolic space, we introduce the ARCL module, which uses ranking-based contrastive learning to preserve the ordinal nature of survival time, along with the CUC module to fully explore the censored data. Extensive experiments demonstrate that our method outperforms state-of-the-art methods on five benchmark datasets. The source code is to be released.
WSI-LLaVA: A Multimodal Large Language Model for Whole Slide Image
Dec 03, 2024



Abstract:Recent advancements in computational pathology have produced patch-level Multi-modal Large Language Models (MLLMs), but these models are limited by their inability to analyze whole slide images (WSIs) comprehensively and their tendency to bypass crucial morphological features that pathologists rely on for diagnosis. To address these challenges, we first introduce WSI-Bench, a large-scale morphology-aware benchmark containing 180k VQA pairs from 9,850 WSIs across 30 cancer types, designed to evaluate MLLMs' understanding of morphological characteristics crucial for accurate diagnosis. Building upon this benchmark, we present WSI-LLaVA, a novel framework for gigapixel WSI understanding that employs a three-stage training approach: WSI-text alignment, feature space alignment, and task-specific instruction tuning. To better assess model performance in pathological contexts, we develop two specialized WSI metrics: WSI-Precision and WSI-Relevance. Experimental results demonstrate that WSI-LLaVA outperforms existing models across all capability dimensions, with a significant improvement in morphological analysis, establishing a clear correlation between morphological understanding and diagnostic accuracy.
PathMMU: A Massive Multimodal Expert-Level Benchmark for Understanding and Reasoning in Pathology
Feb 05, 2024



Abstract:The emergence of large multimodal models has unlocked remarkable potential in AI, particularly in pathology. However, the lack of specialized, high-quality benchmark impeded their development and precise evaluation. To address this, we introduce PathMMU, the largest and highest-quality expert-validated pathology benchmark for LMMs. It comprises 33,573 multimodal multi-choice questions and 21,599 images from various sources, and an explanation for the correct answer accompanies each question. The construction of PathMMU capitalizes on the robust capabilities of GPT-4V, utilizing approximately 30,000 gathered image-caption pairs to generate Q\&As. Significantly, to maximize PathMMU's authority, we invite six pathologists to scrutinize each question under strict standards in PathMMU's validation and test sets, while simultaneously setting an expert-level performance benchmark for PathMMU. We conduct extensive evaluations, including zero-shot assessments of 14 open-sourced and three closed-sourced LMMs and their robustness to image corruption. We also fine-tune representative LMMs to assess their adaptability to PathMMU. The empirical findings indicate that advanced LMMs struggle with the challenging PathMMU benchmark, with the top-performing LMM, GPT-4V, achieving only a 51.7\% zero-shot performance, significantly lower than the 71.4\% demonstrated by human pathologists. After fine-tuning, even open-sourced LMMs can surpass GPT-4V with a performance of over 60\%, but still fall short of the expertise shown by pathologists. We hope that the PathMMU will offer valuable insights and foster the development of more specialized, next-generation LLMs for pathology.
PathAsst: Redefining Pathology through Generative Foundation AI Assistant for Pathology
May 24, 2023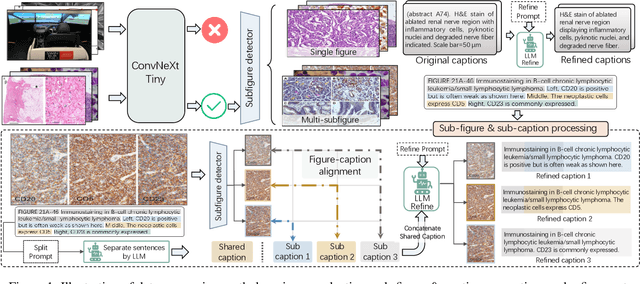
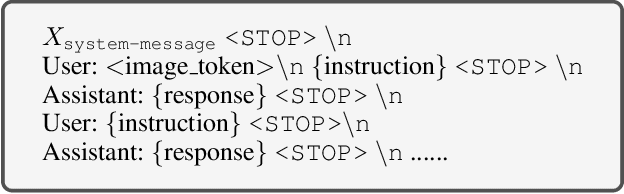

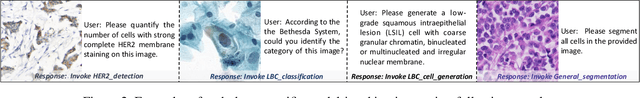
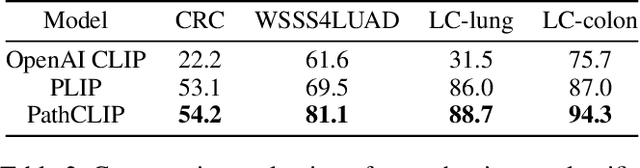
Abstract:As advances in large language models (LLMs) and multimodal techniques continue to mature, the development of general-purpose multimodal large language models (MLLMs) has surged, with significant applications in natural image interpretation. However, the field of pathology has largely remained untapped in this regard, despite the growing need for accurate, timely, and personalized diagnostics. To bridge the gap in pathology MLLMs, we present the PathAsst in this study, which is a generative foundation AI assistant to revolutionize diagnostic and predictive analytics in pathology. To develop PathAsst, we collect over 142K high-quality pathology image-text pairs from a variety of reliable sources, including PubMed, comprehensive pathology textbooks, reputable pathology websites, and private data annotated by pathologists. Leveraging the advanced capabilities of ChatGPT/GPT-4, we generate over 180K instruction-following samples. Furthermore, we devise additional instruction-following data, specifically tailored for the invocation of the pathology-specific models, allowing the PathAsst to effectively interact with these models based on the input image and user intent, consequently enhancing the model's diagnostic capabilities. Subsequently, our PathAsst is trained based on Vicuna-13B language model in coordination with the CLIP vision encoder. The results of PathAsst show the potential of harnessing the AI-powered generative foundation model to improve pathology diagnosis and treatment processes. We are committed to open-sourcing our meticulously curated dataset, as well as a comprehensive toolkit designed to aid researchers in the extensive collection and preprocessing of their own datasets. Resources can be obtained at https://github.com/superjamessyx/Generative-Foundation-AI-Assistant-for-Pathology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge