Sunyi Zheng
Large-scale cervical precancerous screening via AI-assisted cytology whole slide image analysis
Jul 28, 2024



Abstract:Cervical Cancer continues to be the leading gynecological malignancy, posing a persistent threat to women's health on a global scale. Early screening via cytology Whole Slide Image (WSI) diagnosis is critical to prevent this Cancer progression and improve survival rate, but pathologist's single test suffers inevitable false negative due to the immense number of cells that need to be reviewed within a WSI. Though computer-aided automated diagnostic models can serve as strong complement for pathologists, their effectiveness is hampered by the paucity of extensive and detailed annotations, coupled with the limited interpretability and robustness. These factors significantly hinder their practical applicability and reliability in clinical settings. To tackle these challenges, we develop an AI approach, which is a Scalable Technology for Robust and Interpretable Diagnosis built on Extensive data (STRIDE) of cervical cytology. STRIDE addresses the bottleneck of limited annotations by integrating patient-level labels with a small portion of cell-level labels through an end-to-end training strategy, facilitating scalable learning across extensive datasets. To further improve the robustness to real-world domain shifts of cytology slide-making and imaging, STRIDE employs color adversarial samples training that mimic staining and imaging variations. Lastly, to achieve pathologist-level interpretability for the trustworthiness in clinical settings, STRIDE can generate explanatory textual descriptions that simulates pathologists' diagnostic processes by cell image feature and textual description alignment. Conducting extensive experiments and evaluations in 183 medical centers with a dataset of 341,889 WSIs and 0.1 billion cells from cervical cytology patients, STRIDE has demonstrated a remarkable superiority over previous state-of-the-art techniques.
WSI-VQA: Interpreting Whole Slide Images by Generative Visual Question Answering
Jul 08, 2024



Abstract:Whole slide imaging is routinely adopted for carcinoma diagnosis and prognosis. Abundant experience is required for pathologists to achieve accurate and reliable diagnostic results of whole slide images (WSI). The huge size and heterogeneous features of WSIs make the workflow of pathological reading extremely time-consuming. In this paper, we propose a novel framework (WSI-VQA) to interpret WSIs by generative visual question answering. WSI-VQA shows universality by reframing various kinds of slide-level tasks in a question-answering pattern, in which pathologists can achieve immunohistochemical grading, survival prediction, and tumor subtyping following human-machine interaction. Furthermore, we establish a WSI-VQA dataset which contains 8672 slide-level question-answering pairs with 977 WSIs. Besides the ability to deal with different slide-level tasks, our generative model which is named Wsi2Text Transformer (W2T) outperforms existing discriminative models in medical correctness, which reveals the potential of our model to be applied in the clinical scenario. Additionally, we also visualize the co-attention mapping between word embeddings and WSIs as an intuitive explanation for diagnostic results. The dataset and related code are available at https://github.com/cpystan/WSI-VQA.
ADR: Attention Diversification Regularization for Mitigating Overfitting in Multiple Instance Learning based Whole Slide Image Classification
Jun 18, 2024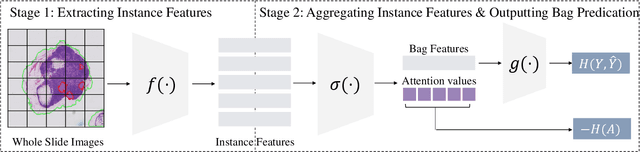
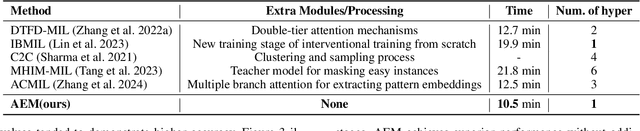

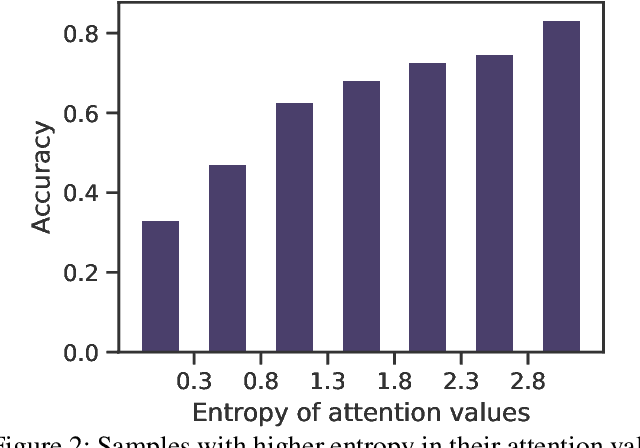
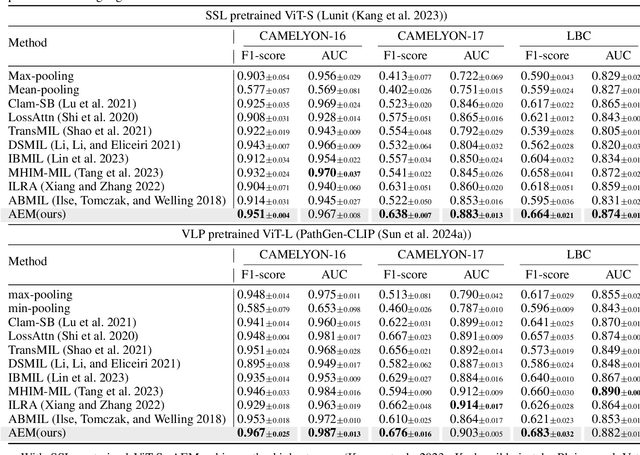
Abstract:Multiple Instance Learning (MIL) has demonstrated effectiveness in analyzing whole slide images (WSIs), yet it often encounters overfitting challenges in real-world applications. This paper reveals the correlation between MIL's performance and the entropy of attention values. Based on this observation, we propose Attention Diversity Regularization (ADR), a simple but effective technique aimed at promoting high entropy in attention values. Specifically, ADR introduces a negative Shannon entropy loss for attention values into the regular MIL framework. Compared to existing methods aimed at alleviating overfitting, which often necessitate additional modules or processing steps, our ADR approach requires no such extras, demonstrating simplicity and efficiency. We evaluate our ADR on three WSI classification tasks. ADR achieves superior performance over the state-of-the-art on most of them. We also show that ADR can enhance heatmaps, aligning them better with pathologists' diagnostic criteria. The source code is available at \url{https://github.com/dazhangyu123/ADR}.
PathMMU: A Massive Multimodal Expert-Level Benchmark for Understanding and Reasoning in Pathology
Feb 05, 2024



Abstract:The emergence of large multimodal models has unlocked remarkable potential in AI, particularly in pathology. However, the lack of specialized, high-quality benchmark impeded their development and precise evaluation. To address this, we introduce PathMMU, the largest and highest-quality expert-validated pathology benchmark for LMMs. It comprises 33,573 multimodal multi-choice questions and 21,599 images from various sources, and an explanation for the correct answer accompanies each question. The construction of PathMMU capitalizes on the robust capabilities of GPT-4V, utilizing approximately 30,000 gathered image-caption pairs to generate Q\&As. Significantly, to maximize PathMMU's authority, we invite six pathologists to scrutinize each question under strict standards in PathMMU's validation and test sets, while simultaneously setting an expert-level performance benchmark for PathMMU. We conduct extensive evaluations, including zero-shot assessments of 14 open-sourced and three closed-sourced LMMs and their robustness to image corruption. We also fine-tune representative LMMs to assess their adaptability to PathMMU. The empirical findings indicate that advanced LMMs struggle with the challenging PathMMU benchmark, with the top-performing LMM, GPT-4V, achieving only a 51.7\% zero-shot performance, significantly lower than the 71.4\% demonstrated by human pathologists. After fine-tuning, even open-sourced LMMs can surpass GPT-4V with a performance of over 60\%, but still fall short of the expertise shown by pathologists. We hope that the PathMMU will offer valuable insights and foster the development of more specialized, next-generation LLMs for pathology.
Benchmarking PathCLIP for Pathology Image Analysis
Jan 05, 2024
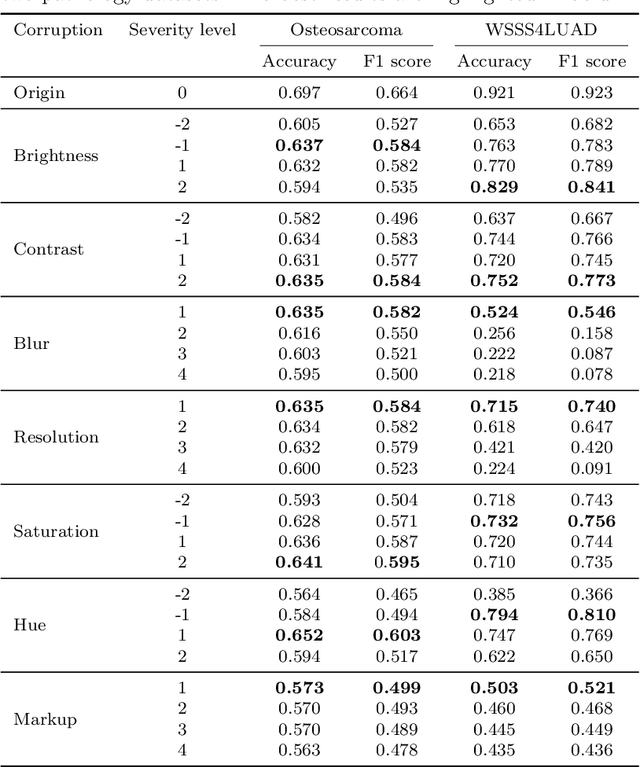

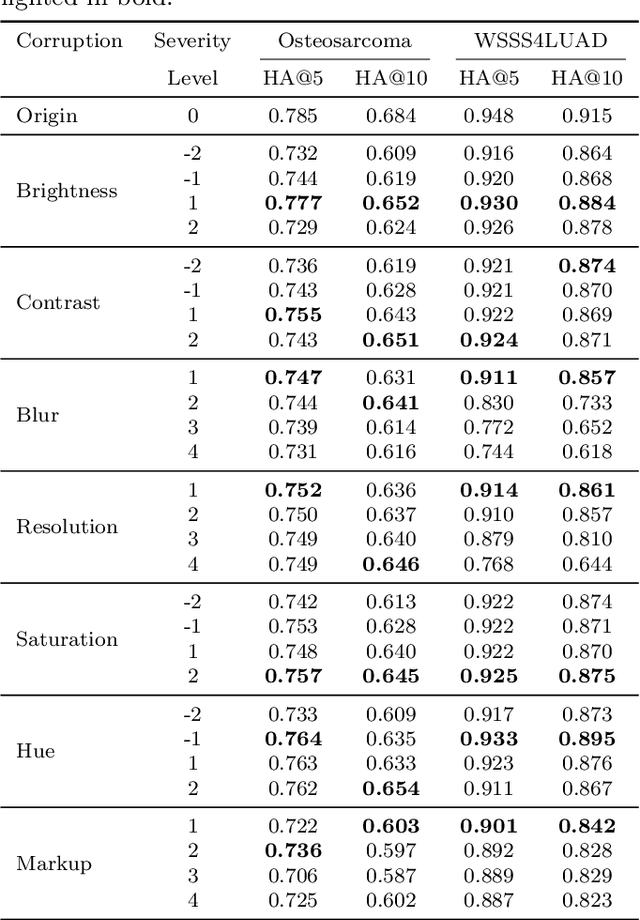
Abstract:Accurate image classification and retrieval are of importance for clinical diagnosis and treatment decision-making. The recent contrastive language-image pretraining (CLIP) model has shown remarkable proficiency in understanding natural images. Drawing inspiration from CLIP, PathCLIP is specifically designed for pathology image analysis, utilizing over 200,000 image and text pairs in training. While the performance the PathCLIP is impressive, its robustness under a wide range of image corruptions remains unknown. Therefore, we conduct an extensive evaluation to analyze the performance of PathCLIP on various corrupted images from the datasets of Osteosarcoma and WSSS4LUAD. In our experiments, we introduce seven corruption types including brightness, contrast, Gaussian blur, resolution, saturation, hue, and markup at four severity levels. Through experiments, we find that PathCLIP is relatively robustness to image corruptions and surpasses OpenAI-CLIP and PLIP in zero-shot classification. Among the seven corruptions, blur and resolution can cause server performance degradation of the PathCLIP. This indicates that ensuring the quality of images is crucial before conducting a clinical test. Additionally, we assess the robustness of PathCLIP in the task of image-image retrieval, revealing that PathCLIP performs less effectively than PLIP on Osteosarcoma but performs better on WSSS4LUAD under diverse corruptions. Overall, PathCLIP presents impressive zero-shot classification and retrieval performance for pathology images, but appropriate care needs to be taken when using it. We hope this study provides a qualitative impression of PathCLIP and helps understand its differences from other CLIP models.
Multi-modal Learning with Missing Modality in Predicting Axillary Lymph Node Metastasis
Jan 03, 2024



Abstract:Multi-modal Learning has attracted widespread attention in medical image analysis. Using multi-modal data, whole slide images (WSIs) and clinical information, can improve the performance of deep learning models in the diagnosis of axillary lymph node metastasis. However, clinical information is not easy to collect in clinical practice due to privacy concerns, limited resources, lack of interoperability, etc. Although patient selection can ensure the training set to have multi-modal data for model development, missing modality of clinical information can appear during test. This normally leads to performance degradation, which limits the use of multi-modal models in the clinic. To alleviate this problem, we propose a bidirectional distillation framework consisting of a multi-modal branch and a single-modal branch. The single-modal branch acquires the complete multi-modal knowledge from the multi-modal branch, while the multi-modal learns the robust features of WSI from the single-modal. We conduct experiments on a public dataset of Lymph Node Metastasis in Early Breast Cancer to validate the method. Our approach not only achieves state-of-the-art performance with an AUC of 0.861 on the test set without missing data, but also yields an AUC of 0.842 when the rate of missing modality is 80\%. This shows the effectiveness of the approach in dealing with multi-modal data and missing modality. Such a model has the potential to improve treatment decision-making for early breast cancer patients who have axillary lymph node metastatic status.
MI-Gen: Multiple Instance Generation of Pathology Reports for Gigapixel Whole-Slide Images
Nov 27, 2023



Abstract:Whole slide images are the foundation of digital pathology for the diagnosis and treatment of carcinomas. Writing pathology reports is laborious and error-prone for inexperienced pathologists. To reduce the workload and improve clinical automation, we investigate how to generate pathology reports given whole slide images. On the data end, we curated the largest WSI-text dataset (TCGA-PathoText). In specific, we collected nearly 10000 high-quality WSI-text pairs for visual-language models by recognizing and cleaning pathology reports which narrate diagnostic slides in TCGA. On the model end, we propose the multiple instance generative model (MI-Gen) which can produce pathology reports for gigapixel WSIs. We benchmark our model on the largest subset of TCGA-PathoText. Experimental results show our model can generate pathology reports which contain multiple clinical clues. Furthermore, WSI-text prediction can be seen as an approach of visual-language pre-training, which enables our model to be transferred to downstream diagnostic tasks like carcinoma grading and phenotyping. We observe that simple semantic extraction from the pathology reports can achieve the best performance (0.838 of F1 score) on BRCA subtyping without adding extra parameters or tricky fine-tuning. Our collected dataset and related code will all be publicly available.
Long-MIL: Scaling Long Contextual Multiple Instance Learning for Histopathology Whole Slide Image Analysis
Nov 21, 2023Abstract:Histopathology image analysis is the golden standard of clinical diagnosis for Cancers. In doctors daily routine and computer-aided diagnosis, the Whole Slide Image (WSI) of histopathology tissue is used for analysis. Because of the extremely large scale of resolution, previous methods generally divide the WSI into a large number of patches, then aggregate all patches within a WSI by Multi-Instance Learning (MIL) to make the slide-level prediction when developing computer-aided diagnosis tools. However, most previous WSI-MIL models using global-attention without pairwise interaction and any positional information, or self-attention with absolute position embedding can not well handle shape varying large WSIs, e.g. testing WSIs after model deployment may be larger than training WSIs, since the model development set is always limited due to the difficulty of histopathology WSIs collection. To deal with the problem, in this paper, we propose to amend position embedding for shape varying long-contextual WSI by introducing Linear Bias into Attention, and adapt it from 1-d long sequence into 2-d long-contextual WSI which helps model extrapolate position embedding to unseen or under-fitted positions. We further utilize Flash-Attention module to tackle the computational complexity of Transformer, which also keep full self-attention performance compared to previous attention approximation work. Our method, Long-contextual MIL (Long-MIL) are evaluated on extensive experiments including 4 dataset including WSI classification and survival prediction tasks to validate the superiority on shape varying WSIs. The source code will be open-accessed soon.
Test-Time Training for Semantic Segmentation with Output Contrastive Loss
Nov 14, 2023
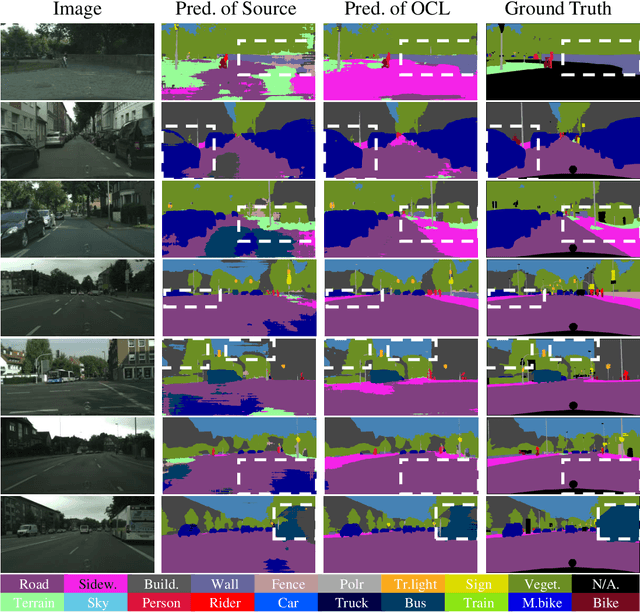
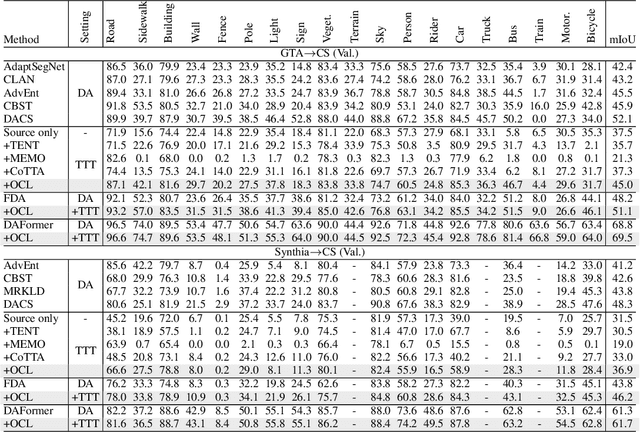
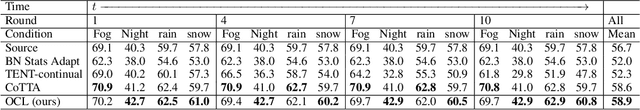

Abstract:Although deep learning-based segmentation models have achieved impressive performance on public benchmarks, generalizing well to unseen environments remains a major challenge. To improve the model's generalization ability to the new domain during evaluation, the test-time training (TTT) is a challenging paradigm that adapts the source-pretrained model in an online fashion. Early efforts on TTT mainly focus on the image classification task. Directly extending these methods to semantic segmentation easily experiences unstable adaption due to segmentation's inherent characteristics, such as extreme class imbalance and complex decision spaces. To stabilize the adaptation process, we introduce contrastive loss (CL), known for its capability to learn robust and generalized representations. Nevertheless, the traditional CL operates in the representation space and cannot directly enhance predictions. In this paper, we resolve this limitation by adapting the CL to the output space, employing a high temperature, and simplifying the formulation, resulting in a straightforward yet effective loss function called Output Contrastive Loss (OCL). Our comprehensive experiments validate the efficacy of our approach across diverse evaluation scenarios. Notably, our method excels even when applied to models initially pre-trained using domain adaptation methods on test domain data, showcasing its resilience and adaptability.\footnote{Code and more information could be found at~ \url{https://github.com/dazhangyu123/OCL}}
Attention-Challenging Multiple Instance Learning for Whole Slide Image Classification
Nov 13, 2023Abstract:Overfitting remains a significant challenge in the application of Multiple Instance Learning (MIL) methods for Whole Slide Image (WSI) analysis. Visualizing heatmaps reveals that current MIL methods focus on a subset of predictive instances, hindering effective model generalization. To tackle this, we propose Attention-Challenging MIL (ACMIL), aimed at forcing the attention mechanism to capture more challenging predictive instances. ACMIL incorporates two techniques, Multiple Branch Attention (MBA) to capture richer predictive instances and Stochastic Top-K Instance Masking (STKIM) to suppress simple predictive instances. Evaluation on three WSI datasets outperforms state-of-the-art methods. Additionally, through heatmap visualization, UMAP visualization, and attention value statistics, this paper comprehensively illustrates ACMIL's effectiveness in overcoming the overfitting challenge. The source code is available at \url{https://github.com/dazhangyu123/ACMIL}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge