Soumyadeep Roy
Sample Efficient Active Algorithms for Offline Reinforcement Learning
Feb 01, 2026Abstract:Offline reinforcement learning (RL) enables policy learning from static data but often suffers from poor coverage of the state-action space and distributional shift problems. This problem can be addressed by allowing limited online interactions to selectively refine uncertain regions of the learned value function, which is referred to as Active Reinforcement Learning (ActiveRL). While there has been good empirical success, no theoretical analysis is available in the literature. We fill this gap by developing a rigorous sample-complexity analysis of ActiveRL through the lens of Gaussian Process (GP) uncertainty modeling. In this respect, we propose an algorithm and using GP concentration inequalities and information-gain bounds, we derive high-probability guarantees showing that an $ε$-optimal policy can be learned with ${\mathcal{O}}(1/ε^2)$ active transitions, improving upon the $Ω(1/ε^2(1-γ)^4)$ rate of purely offline methods. Our results reveal that ActiveRL achieves near-optimal information efficiency, that is, guided uncertainty reduction leads to accelerated value-function convergence with minimal online data. Our analysis builds on GP concentration inequalities and information-gain bounds, bridging Bayesian nonparametric regression and reinforcement learning theories. We conduct several experiments to validate the algorithm and theoretical findings.
"Where does it hurt?" -- Dataset and Study on Physician Intent Trajectories in Doctor Patient Dialogues
Aug 26, 2025



Abstract:In a doctor-patient dialogue, the primary objective of physicians is to diagnose patients and propose a treatment plan. Medical doctors guide these conversations through targeted questioning to efficiently gather the information required to provide the best possible outcomes for patients. To the best of our knowledge, this is the first work that studies physician intent trajectories in doctor-patient dialogues. We use the `Ambient Clinical Intelligence Benchmark' (Aci-bench) dataset for our study. We collaborate with medical professionals to develop a fine-grained taxonomy of physician intents based on the SOAP framework (Subjective, Objective, Assessment, and Plan). We then conduct a large-scale annotation effort to label over 5000 doctor-patient turns with the help of a large number of medical experts recruited using Prolific, a popular crowd-sourcing platform. This large labeled dataset is an important resource contribution that we use for benchmarking the state-of-the-art generative and encoder models for medical intent classification tasks. Our findings show that our models understand the general structure of medical dialogues with high accuracy, but often fail to identify transitions between SOAP categories. We also report for the first time common trajectories in medical dialogue structures that provide valuable insights for designing `differential diagnosis' systems. Finally, we extensively study the impact of intent filtering for medical dialogue summarization and observe a significant boost in performance. We make the codes and data, including annotation guidelines, publicly available at https://github.com/DATEXIS/medical-intent-classification.
Evaluation of LLMs in Medical Text Summarization: The Role of Vocabulary Adaptation in High OOV Settings
May 27, 2025



Abstract:Large Language Models (LLMs) recently achieved great success in medical text summarization by simply using in-context learning. However, these recent efforts do not perform fine-grained evaluations under difficult settings where LLMs might fail. They typically report performance scores over the entire dataset. Through our benchmarking study, we show that LLMs show a significant performance drop for data points with high concentration of out-of-vocabulary (OOV) words or with high novelty. Vocabulary adaptation is an intuitive solution to this vocabulary mismatch issue where the LLM vocabulary gets updated with certain expert domain (here, medical) words or subwords. An interesting finding from our study is that Llama-3.1, even with a vocabulary size of around 128K tokens, still faces over-fragmentation issue with medical words. To that end, we show vocabulary adaptation helps improve the LLM summarization performance even in difficult settings. Through extensive experimentation of multiple vocabulary adaptation strategies, two continual pretraining strategies, and three benchmark medical summarization datasets, we gain valuable insights into the role of vocabulary adaptation strategies for customizing LLMs to the medical domain. We also performed a human evaluation study with medical experts where they found that vocabulary adaptation results in more relevant and faithful summaries. Our codebase is made publicly available at https://github.com/gb-kgp/LLM-MedicalSummarization-Benchmark.
Adaptive BPE Tokenization for Enhanced Vocabulary Adaptation in Finetuning Pretrained Language Models
Oct 04, 2024



Abstract:In this work, we show a fundamental limitation in vocabulary adaptation approaches that use Byte-Pair Encoding (BPE) tokenization scheme for fine-tuning pretrained language models (PLMs) to expert domains. Current approaches trivially append the target domain-specific vocabulary at the end of the PLM vocabulary. This approach leads to a lower priority score and causes sub-optimal tokenization in BPE that iteratively uses merge rules to tokenize a given text. To mitigate this issue, we propose AdaptBPE where the BPE tokenization initialization phase is modified to first perform the longest string matching on the added (target) vocabulary before tokenizing at the character level. We perform an extensive evaluation of AdaptBPE versus the standard BPE over various classification and summarization tasks; AdaptBPE improves by 3.57% (in terms of accuracy) and 1.87% (in terms of Rouge-L), respectively. AdaptBPE for MEDVOC works particularly well when reference summaries have high OOV concentration or are longer in length. We also conduct a human evaluation, revealing that AdaptBPE generates more relevant and more faithful summaries as compared to MEDVOC. We make our codebase publicly available at https://github.com/gb-kgp/adaptbpe.
Unlocking Efficiency: Adaptive Masking for Gene Transformer Models
Aug 13, 2024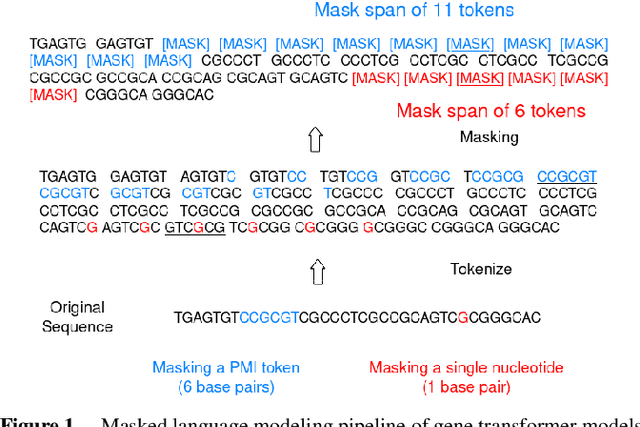
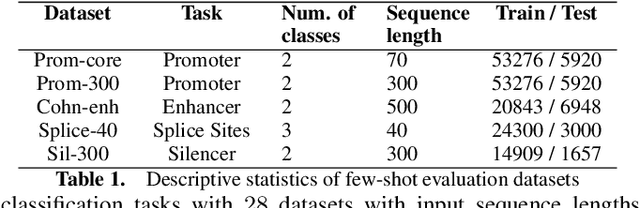
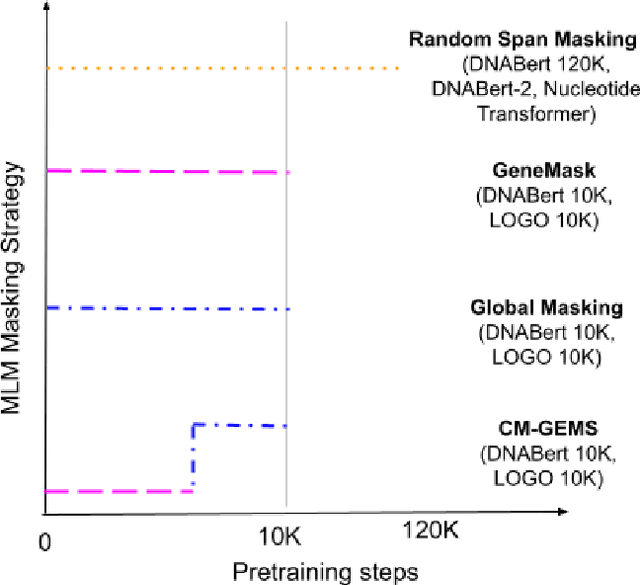
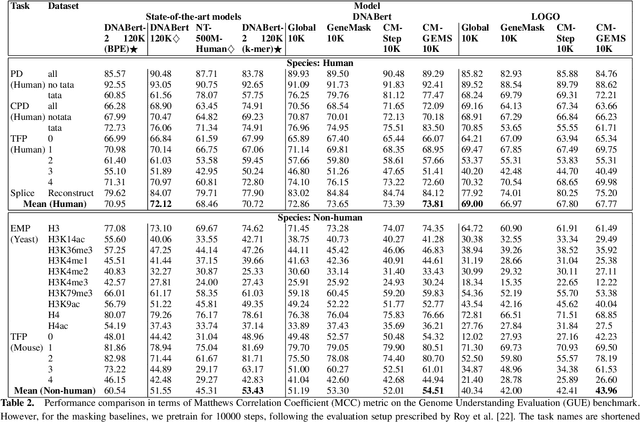
Abstract:Gene transformer models such as Nucleotide Transformer, DNABert, and LOGO are trained to learn optimal gene sequence representations by using the Masked Language Modeling (MLM) training objective over the complete Human Reference Genome. However, the typical tokenization methods employ a basic sliding window of tokens, such as k-mers, that fail to utilize gene-centric semantics. This could result in the (trivial) masking of easily predictable sequences, leading to inefficient MLM training. Time-variant training strategies are known to improve pretraining efficiency in both language and vision tasks. In this work, we focus on using curriculum masking where we systematically increase the difficulty of masked token prediction task by using a Pointwise Mutual Information-based difficulty criterion, as gene sequences lack well-defined semantic units similar to words or sentences of NLP domain. Our proposed Curriculum Masking-based Gene Masking Strategy (CM-GEMS) demonstrates superior representation learning capabilities compared to baseline masking approaches when evaluated on downstream gene sequence classification tasks. We perform extensive evaluation in both few-shot (five datasets) and full dataset settings (Genomic Understanding Evaluation benchmark consisting of 27 tasks). Our findings reveal that CM-GEMS outperforms state-of-the-art models (DNABert-2, Nucleotide transformer, DNABert) trained at 120K steps, achieving similar results in just 10K and 1K steps. We also demonstrate that Curriculum-Learned LOGO (a 2-layer DNABert-like model) can achieve nearly 90% of the state-of-the-art model performance of 120K steps. We will make the models and codes publicly available at https://github.com/roysoumya/curriculum-GeneMask.
MEDVOC: Vocabulary Adaptation for Fine-tuning Pre-trained Language Models on Medical Text Summarization
May 07, 2024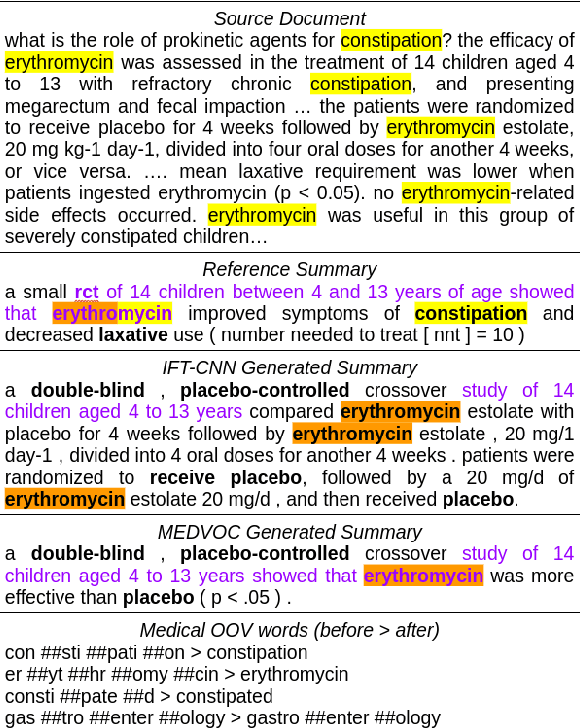
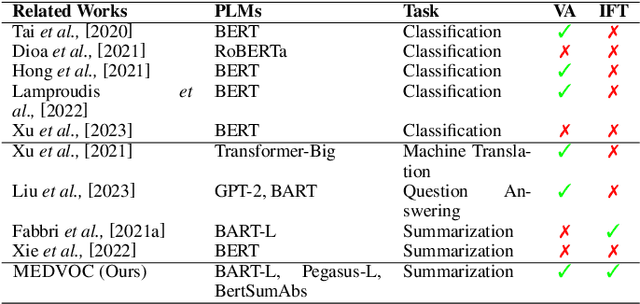
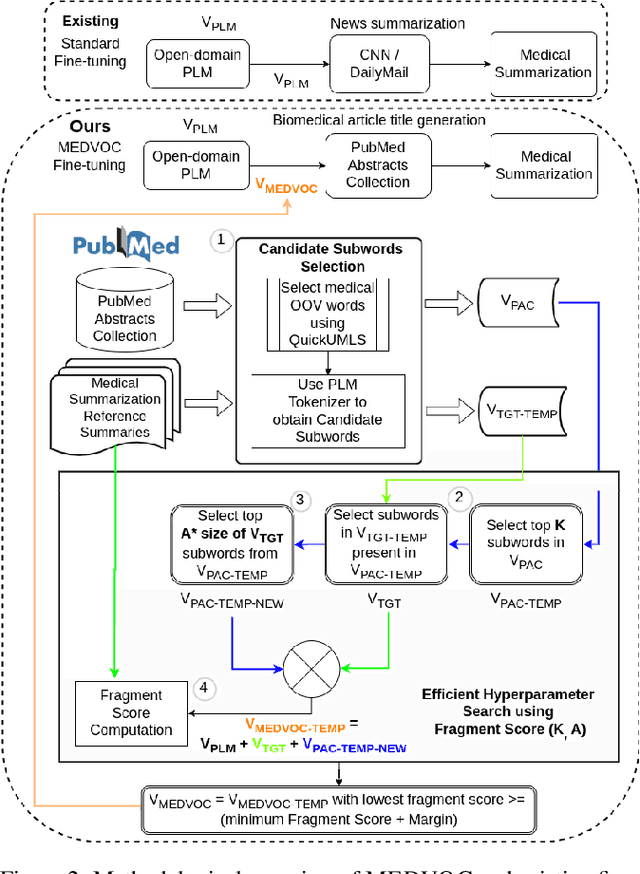
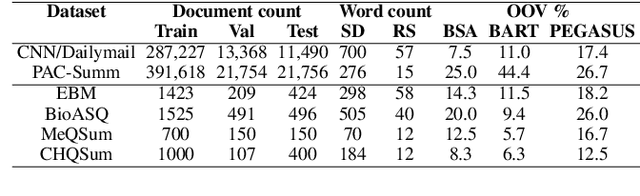
Abstract:This work presents a dynamic vocabulary adaptation strategy, MEDVOC, for fine-tuning pre-trained language models (PLMs) like BertSumAbs, BART, and PEGASUS for improved medical text summarization. In contrast to existing domain adaptation approaches in summarization, MEDVOC treats vocabulary as an optimizable parameter and optimizes the PLM vocabulary based on fragment score conditioned only on the downstream task's reference summaries. Unlike previous works on vocabulary adaptation (limited only to classification tasks), optimizing vocabulary based on summarization tasks requires an extremely costly intermediate fine-tuning step on large summarization datasets. To that end, our novel fragment score-based hyperparameter search very significantly reduces this fine-tuning time -- from 450 days to less than 2 days on average. Furthermore, while previous works on vocabulary adaptation are often primarily tied to single PLMs, MEDVOC is designed to be deployable across multiple PLMs (with varying model vocabulary sizes, pre-training objectives, and model sizes) -- bridging the limited vocabulary overlap between the biomedical literature domain and PLMs. MEDVOC outperforms baselines by 15.74% in terms of Rouge-L in zero-shot setting and shows gains of 17.29% in high Out-Of-Vocabulary (OOV) concentrations. Our human evaluation shows MEDVOC generates more faithful medical summaries (88% compared to 59% in baselines). We make the codebase publicly available at https://github.com/gb-kgp/MEDVOC.
Beyond Accuracy: Investigating Error Types in GPT-4 Responses to USMLE Questions
Apr 20, 2024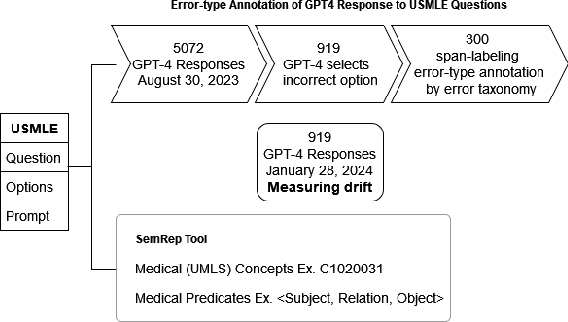
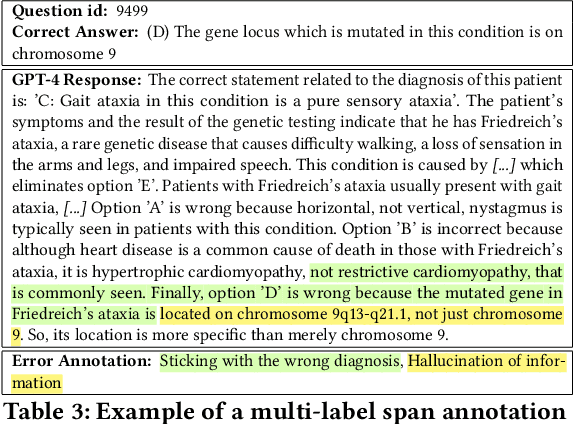
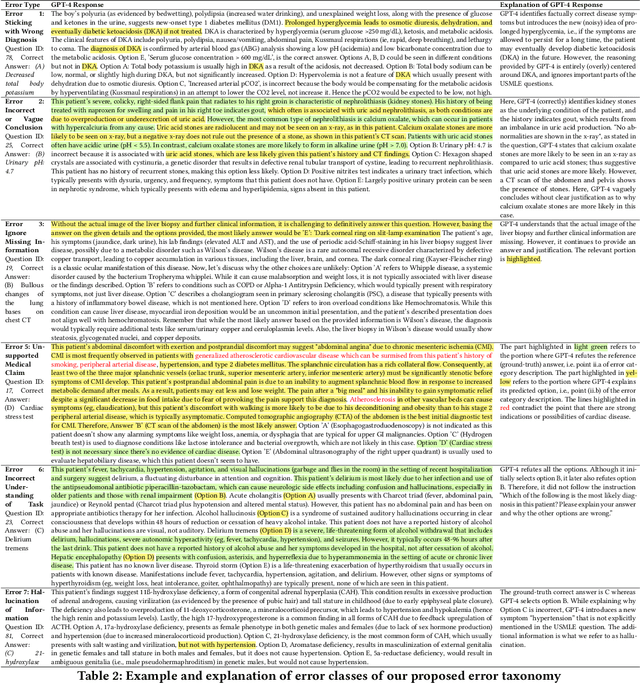
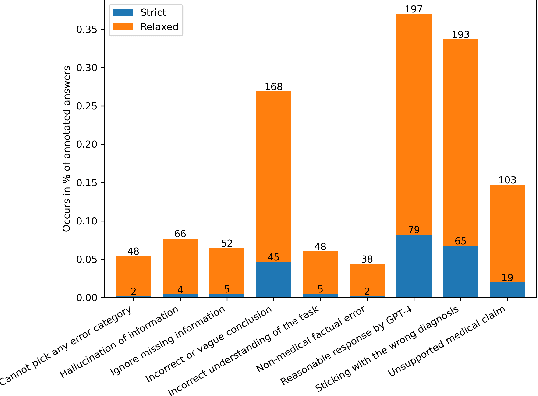

Abstract:GPT-4 demonstrates high accuracy in medical QA tasks, leading with an accuracy of 86.70%, followed by Med-PaLM 2 at 86.50%. However, around 14% of errors remain. Additionally, current works use GPT-4 to only predict the correct option without providing any explanation and thus do not provide any insight into the thinking process and reasoning used by GPT-4 or other LLMs. Therefore, we introduce a new domain-specific error taxonomy derived from collaboration with medical students. Our GPT-4 USMLE Error (G4UE) dataset comprises 4153 GPT-4 correct responses and 919 incorrect responses to the United States Medical Licensing Examination (USMLE) respectively. These responses are quite long (258 words on average), containing detailed explanations from GPT-4 justifying the selected option. We then launch a large-scale annotation study using the Potato annotation platform and recruit 44 medical experts through Prolific, a well-known crowdsourcing platform. We annotated 300 out of these 919 incorrect data points at a granular level for different classes and created a multi-label span to identify the reasons behind the error. In our annotated dataset, a substantial portion of GPT-4's incorrect responses is categorized as a "Reasonable response by GPT-4," by annotators. This sheds light on the challenge of discerning explanations that may lead to incorrect options, even among trained medical professionals. We also provide medical concepts and medical semantic predications extracted using the SemRep tool for every data point. We believe that it will aid in evaluating the ability of LLMs to answer complex medical questions. We make the resources available at https://github.com/roysoumya/usmle-gpt4-error-taxonomy .
GeneMask: Fast Pretraining of Gene Sequences to Enable Few-Shot Learning
Jul 29, 2023Abstract:Large-scale language models such as DNABert and LOGO aim to learn optimal gene representations and are trained on the entire Human Reference Genome. However, standard tokenization schemes involve a simple sliding window of tokens like k-mers that do not leverage any gene-based semantics and thus may lead to (trivial) masking of easily predictable sequences and subsequently inefficient Masked Language Modeling (MLM) training. Therefore, we propose a novel masking algorithm, GeneMask, for MLM training of gene sequences, where we randomly identify positions in a gene sequence as mask centers and locally select the span around the mask center with the highest Normalized Pointwise Mutual Information (NPMI) to mask. We observe that in the absence of human-understandable semantics in the genomics domain (in contrast, semantic units like words and phrases are inherently available in NLP), GeneMask-based models substantially outperform the SOTA models (DNABert and LOGO) over four benchmark gene sequence classification datasets in five few-shot settings (10 to 1000-shot). More significantly, the GeneMask-based DNABert model is trained for less than one-tenth of the number of epochs of the original SOTA model. We also observe a strong correlation between top-ranked PMI tokens and conserved DNA sequence motifs, which may indicate the incorporation of latent genomic information. The codes (including trained models) and datasets are made publicly available at https://github.com/roysoumya/GeneMask.
Knowledge-Aware Neural Networks for Medical Forum Question Classification
Sep 27, 2021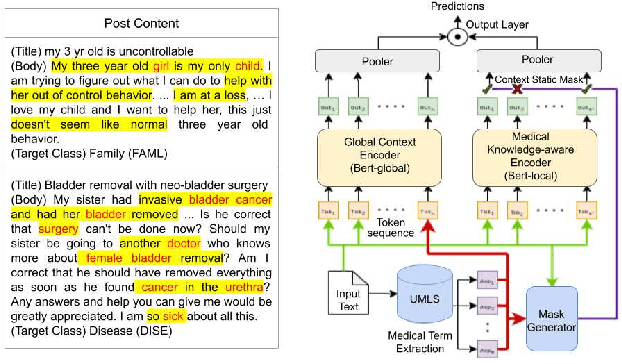
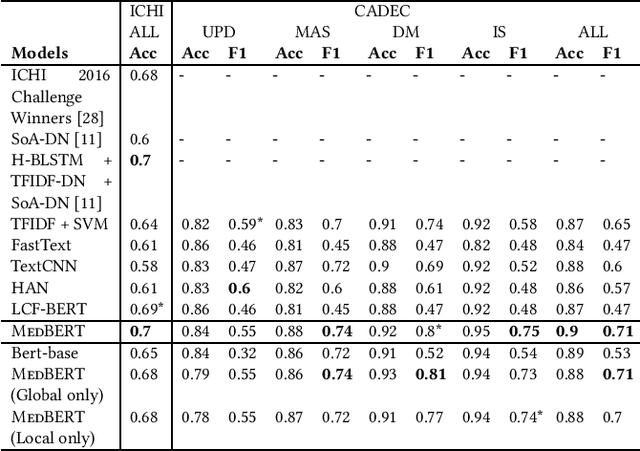
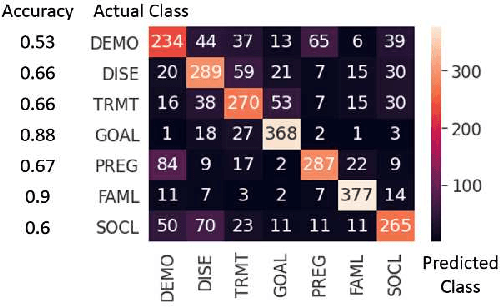
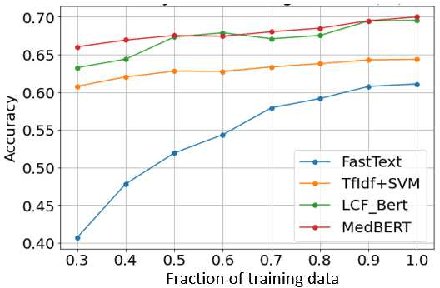
Abstract:Online medical forums have become a predominant platform for answering health-related information needs of consumers. However, with a significant rise in the number of queries and the limited availability of experts, it is necessary to automatically classify medical queries based on a consumer's intention, so that these questions may be directed to the right set of medical experts. Here, we develop a novel medical knowledge-aware BERT-based model (MedBERT) that explicitly gives more weightage to medical concept-bearing words, and utilize domain-specific side information obtained from a popular medical knowledge base. We also contribute a multi-label dataset for the Medical Forum Question Classification (MFQC) task. MedBERT achieves state-of-the-art performance on two benchmark datasets and performs very well in low resource settings.
An Integrated Approach for Improving Brand Consistency of Web Content: Modeling, Analysis and Recommendation
Nov 20, 2020
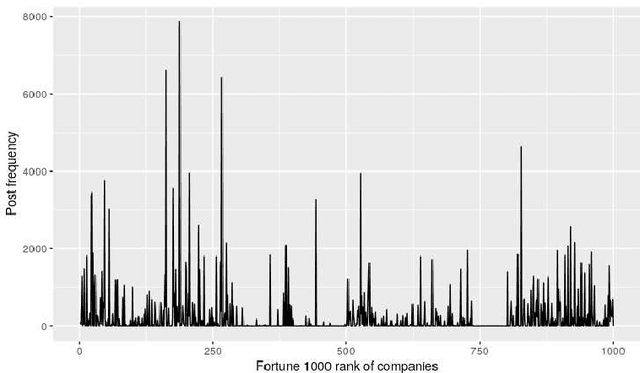
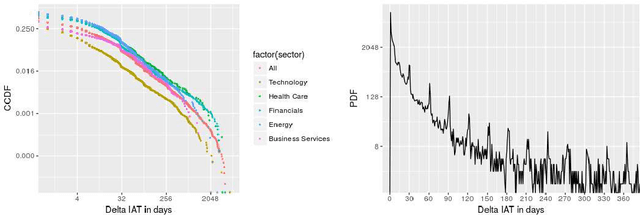
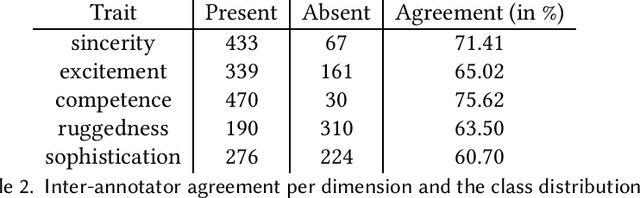
Abstract:A consumer-dependent (business-to-consumer) organization tends to present itself as possessing a set of human qualities, which is termed as the brand personality of the company. The perception is impressed upon the consumer through the content, be it in the form of advertisement, blogs or magazines, produced by the organization. A consistent brand will generate trust and retain customers over time as they develop an affinity towards regularity and common patterns. However, maintaining a consistent messaging tone for a brand has become more challenging with the virtual explosion in the amount of content which needs to be authored and pushed to the Internet to maintain an edge in the era of digital marketing. To understand the depth of the problem, we collect around 300K web page content from around 650 companies. We develop trait-specific classification models by considering the linguistic features of the content. The classifier automatically identifies the web articles which are not consistent with the mission and vision of a company and further helps us to discover the conditions under which the consistency cannot be maintained. To address the brand inconsistency issue, we then develop a sentence ranking system that outputs the top three sentences that need to be changed for making a web article more consistent with the company's brand personality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge