Shahab Aslani
A computationally frugal open-source foundation model for thoracic disease detection in lung cancer screening programs
Jul 02, 2025Abstract:Low-dose computed tomography (LDCT) imaging employed in lung cancer screening (LCS) programs is increasing in uptake worldwide. LCS programs herald a generational opportunity to simultaneously detect cancer and non-cancer-related early-stage lung disease. Yet these efforts are hampered by a shortage of radiologists to interpret scans at scale. Here, we present TANGERINE, a computationally frugal, open-source vision foundation model for volumetric LDCT analysis. Designed for broad accessibility and rapid adaptation, TANGERINE can be fine-tuned off the shelf for a wide range of disease-specific tasks with limited computational resources and training data. Relative to models trained from scratch, TANGERINE demonstrates fast convergence during fine-tuning, thereby requiring significantly fewer GPU hours, and displays strong label efficiency, achieving comparable or superior performance with a fraction of fine-tuning data. Pretrained using self-supervised learning on over 98,000 thoracic LDCTs, including the UK's largest LCS initiative to date and 27 public datasets, TANGERINE achieves state-of-the-art performance across 14 disease classification tasks, including lung cancer and multiple respiratory diseases, while generalising robustly across diverse clinical centres. By extending a masked autoencoder framework to 3D imaging, TANGERINE offers a scalable solution for LDCT analysis, departing from recent closed, resource-intensive models by combining architectural simplicity, public availability, and modest computational requirements. Its accessible, open-source lightweight design lays the foundation for rapid integration into next-generation medical imaging tools that could transform LCS initiatives, allowing them to pivot from a singular focus on lung cancer detection to comprehensive respiratory disease management in high-risk populations.
Deep Learning for Vascular Segmentation and Applications in Phase Contrast Tomography Imaging
Nov 22, 2023Abstract:Automated blood vessel segmentation is vital for biomedical imaging, as vessel changes indicate many pathologies. Still, precise segmentation is difficult due to the complexity of vascular structures, anatomical variations across patients, the scarcity of annotated public datasets, and the quality of images. We present a thorough literature review, highlighting the state of machine learning techniques across diverse organs. Our goal is to provide a foundation on the topic and identify a robust baseline model for application to vascular segmentation in a new imaging modality, Hierarchical Phase Contrast Tomography (HiP CT). Introduced in 2020 at the European Synchrotron Radiation Facility, HiP CT enables 3D imaging of complete organs at an unprecedented resolution of ca. 20mm per voxel, with the capability for localized zooms in selected regions down to 1mm per voxel without sectioning. We have created a training dataset with double annotator validated vascular data from three kidneys imaged with HiP CT in the context of the Human Organ Atlas Project. Finally, utilising the nnU Net model, we conduct experiments to assess the models performance on both familiar and unseen samples, employing vessel specific metrics. Our results show that while segmentations yielded reasonably high scores such as clDice values ranging from 0.82 to 0.88, certain errors persisted. Large vessels that collapsed due to the lack of hydrostatic pressure (HiP CT is an ex vivo technique) were segmented poorly. Moreover, decreased connectivity in finer vessels and higher segmentation errors at vessel boundaries were observed. Such errors obstruct the understanding of the structures by interrupting vascular tree connectivity. Through our review and outputs, we aim to set a benchmark for subsequent model evaluations using various modalities, especially with the HiP CT imaging database.
A hybrid CNN-RNN approach for survival analysis in a Lung Cancer Screening study
Mar 19, 2023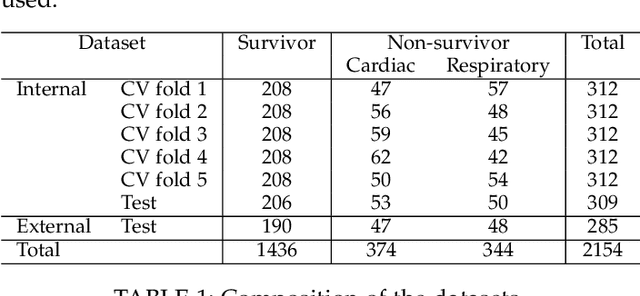

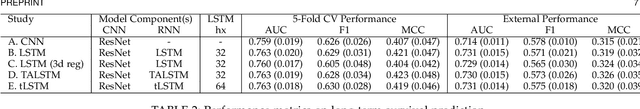
Abstract:In this study, we present a hybrid CNN-RNN approach to investigate long-term survival of subjects in a lung cancer screening study. Subjects who died of cardiovascular and respiratory causes were identified whereby the CNN model was used to capture imaging features in the CT scans and the RNN model was used to investigate time series and thus global information. The models were trained on subjects who underwent cardiovascular and respiratory deaths and a control cohort matched to participant age, gender, and smoking history. The combined model can achieve an AUC of 0.76 which outperforms humans at cardiovascular mortality prediction. The corresponding F1 and Matthews Correlation Coefficient are 0.63 and 0.42 respectively. The generalisability of the model is further validated on an 'external' cohort. The same models were applied to survival analysis with the Cox Proportional Hazard model. It was demonstrated that incorporating the follow-up history can lead to improvement in survival prediction. The Cox neural network can achieve an IPCW C-index of 0.75 on the internal dataset and 0.69 on an external dataset. Delineating imaging features associated with long-term survival can help focus preventative interventions appropriately, particularly for under-recognised pathologies thereby potentially reducing patient morbidity.
Optimising Chest X-Rays for Image Analysis by Identifying and Removing Confounding Factors
Aug 22, 2022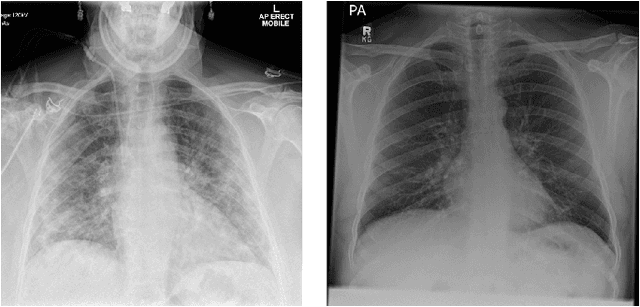
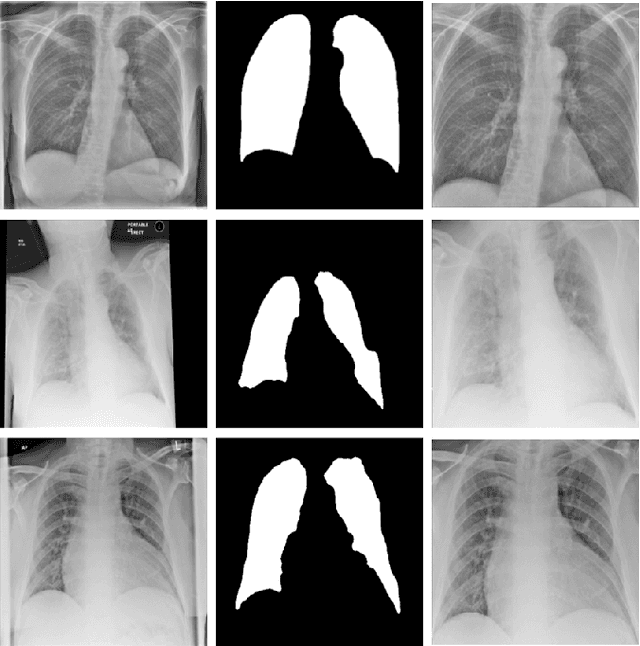
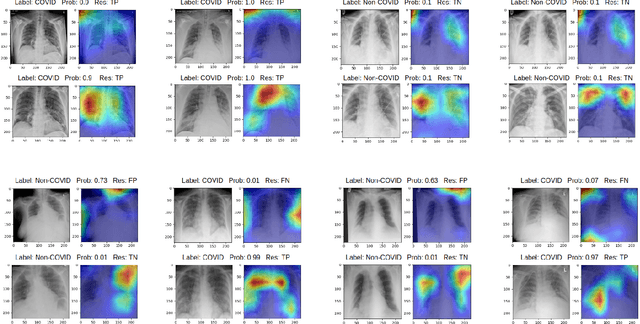
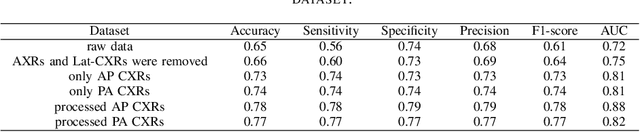
Abstract:During the COVID-19 pandemic, the sheer volume of imaging performed in an emergency setting for COVID-19 diagnosis has resulted in a wide variability of clinical CXR acquisitions. This variation is seen in the CXR projections used, image annotations added and in the inspiratory effort and degree of rotation of clinical images. The image analysis community has attempted to ease the burden on overstretched radiology departments during the pandemic by developing automated COVID-19 diagnostic algorithms, the input for which has been CXR imaging. Large publicly available CXR datasets have been leveraged to improve deep learning algorithms for COVID-19 diagnosis. Yet the variable quality of clinically-acquired CXRs within publicly available datasets could have a profound effect on algorithm performance. COVID-19 diagnosis may be inferred by an algorithm from non-anatomical features on an image such as image labels. These imaging shortcuts may be dataset-specific and limit the generalisability of AI systems. Understanding and correcting key potential biases in CXR images is therefore an essential first step prior to CXR image analysis. In this study, we propose a simple and effective step-wise approach to pre-processing a COVID-19 chest X-ray dataset to remove undesired biases. We perform ablation studies to show the impact of each individual step. The results suggest that using our proposed pipeline could increase accuracy of the baseline COVID-19 detection algorithm by up to 13%.
Enhancing Cancer Prediction in Challenging Screen-Detected Incident Lung Nodules Using Time-Series Deep Learning
Mar 30, 2022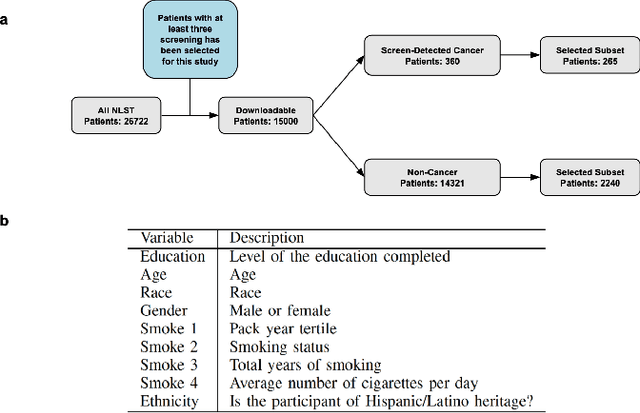
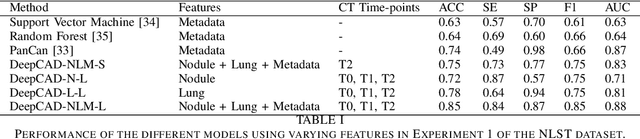
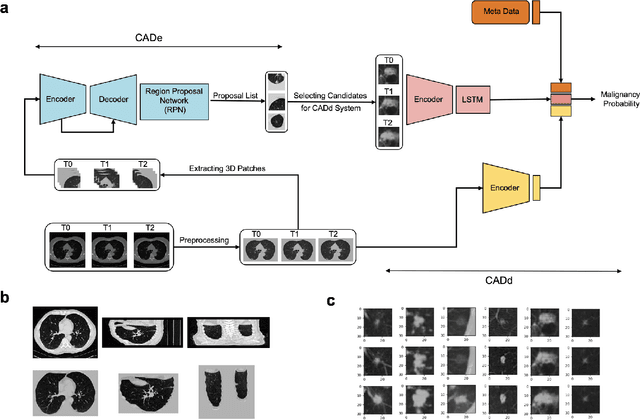
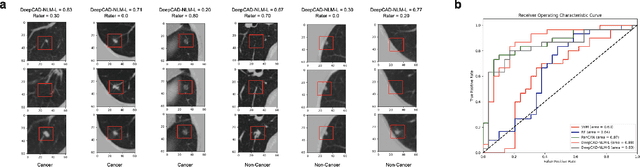

Abstract:Lung cancer is the leading cause of cancer-related mortality worldwide. Lung cancer screening (LCS) using annual low-dose computed tomography (CT) scanning has been proven to significantly reduce lung cancer mortality by detecting cancerous lung nodules at an earlier stage. Improving risk stratification of malignancy risk in lung nodules can be enhanced using machine/deep learning algorithms. However most existing algorithms: a) have primarily assessed single time-point CT data alone thereby failing to utilize the inherent advantages contained within longitudinal imaging datasets; b) have not integrated into computer models pertinent clinical data that might inform risk prediction; c) have not assessed algorithm performance on the spectrum of nodules that are most challenging for radiologists to interpret and where assistance from analytic tools would be most beneficial. Here we show the performance of our time-series deep learning model (DeepCAD-NLM-L) which integrates multi-model information across three longitudinal data domains: nodule-specific, lung-specific, and clinical demographic data. We compared our time-series deep learning model to a) radiologist performance on CTs from the National Lung Screening Trial enriched with the most challenging nodules for diagnosis; b) a nodule management algorithm from a North London LCS study (SUMMIT). Our model demonstrated comparable and complementary performance to radiologists when interpreting challenging lung nodules and showed improved performance (AUC=88\%) against models utilizing single time-point data only. The results emphasise the importance of time-series, multi-modal analysis when interpreting malignancy risk in LCS.
Scanner Invariant Multiple Sclerosis Lesion Segmentation from MRI
Oct 22, 2019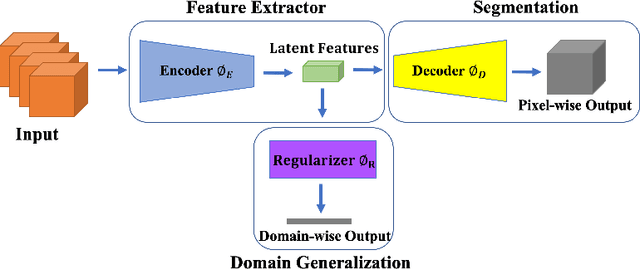
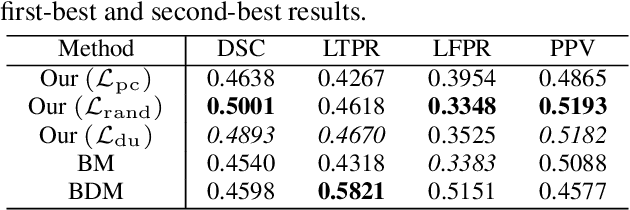
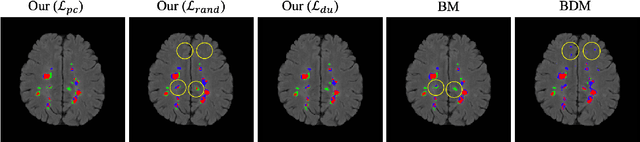
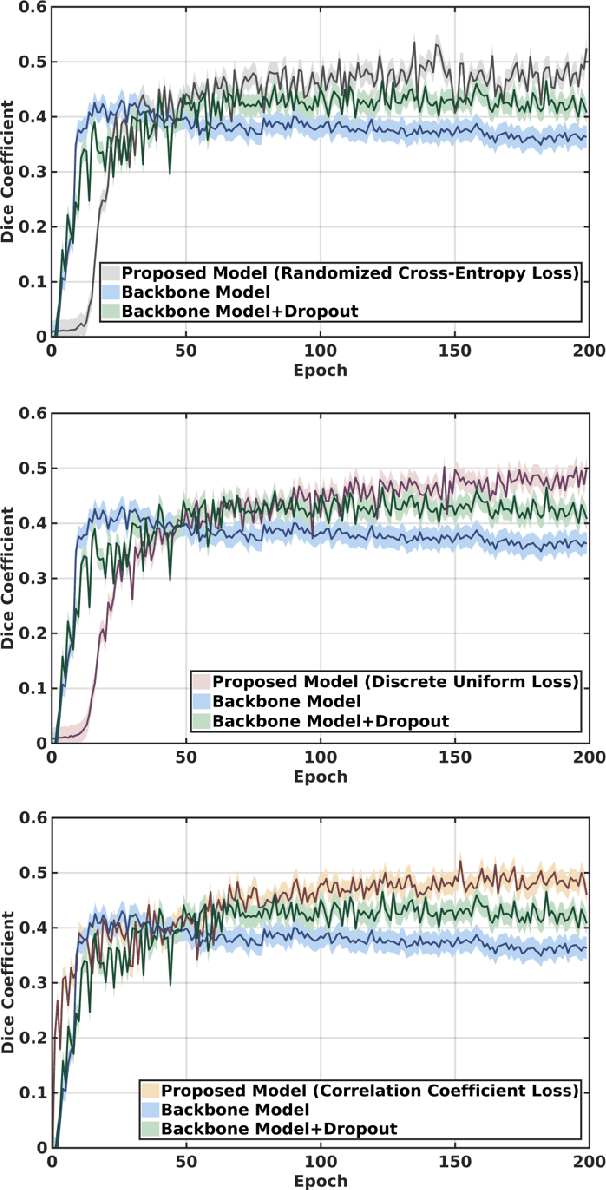
Abstract:This paper presents a simple and effective generalization method for magnetic resonance imaging (MRI) segmentation when data is collected from multiple MRI scanning sites and as a consequence is affected by (site-)domain shifts. We propose to integrate a traditional encoder-decoder network with a regularization network. This added network includes an auxiliary loss term which is responsible for the reduction of the domain shift problem and for the resulting improved generalization. The proposed method was evaluated on multiple sclerosis lesion segmentation from MRI data. We tested the proposed model on an in-house clinical dataset including 117 patients from 56 different scanning sites. In the experiments, our method showed better generalization performance than other baseline networks.
Multi-branch Convolutional Neural Network for Multiple Sclerosis Lesion Segmentation
Nov 16, 2018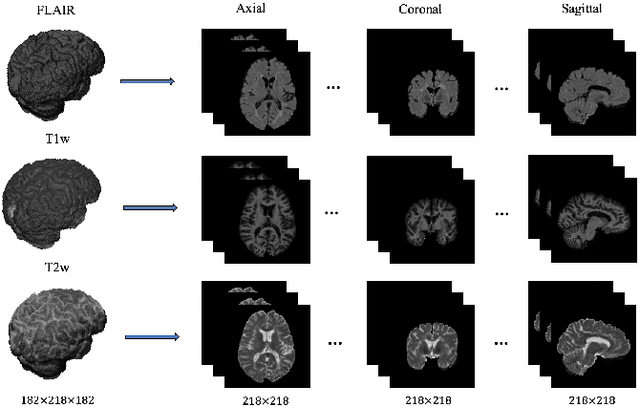
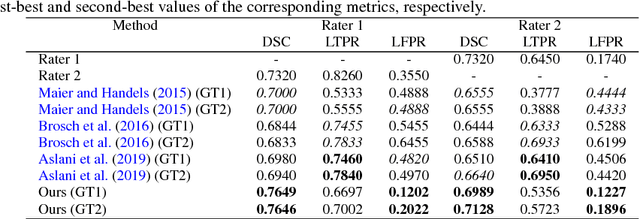
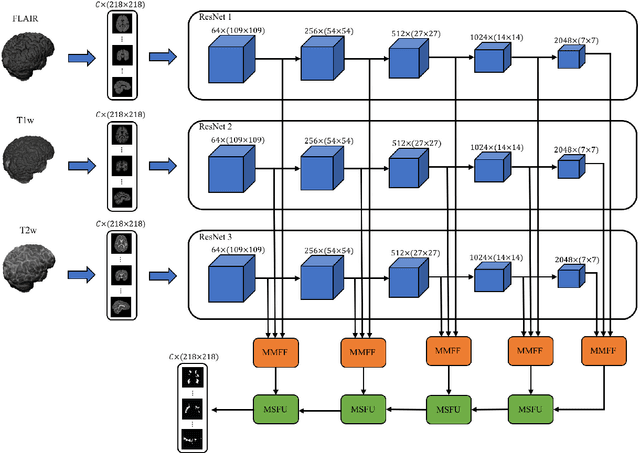
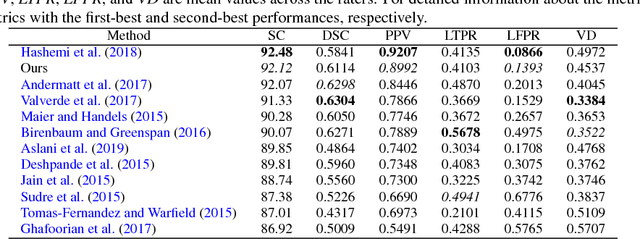
Abstract:In this paper, we present an automated approach for segmenting multiple sclerosis (MS) lesions from multi-modal brain magnetic resonance images. Our method is based on a deep end-to-end 2D convolutional neural network (CNN) for slice-based segmentation of 3D volumetric data. The proposed CNN includes a multi-branch downsampling path, which enables the network to encode slices from multiple modalities separately. Multi-scale feature fusion blocks are proposed to combine feature maps from different modalities at different stages of the network. Then, multi-scale feature upsampling blocks are introduced to upsize combined feature maps with different resolutions to leverage information from the lesion shape and location. We trained and tested our model using orthogonal plane orientations of each 3D modality to exploit the contextual information in all directions. The proposed pipeline is evaluated on two different datasets: a private dataset including 37 MS patients and a publicly available dataset known as the ISBI 2015 longitudinal MS lesion segmentation challenge dataset, consisting of 14 MS patients. Considering the ISBI challenge, at the time of submission, our method was amongst the top performing solutions. On the private dataset, using the same array of performance metrics as in the ISBI challenge, the proposed approach shows high improvements in MS lesion segmentation comparing with other publicly available tools.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge