René Hosch
SALT: Introducing a Framework for Hierarchical Segmentations in Medical Imaging using Softmax for Arbitrary Label Trees
Jul 11, 2024
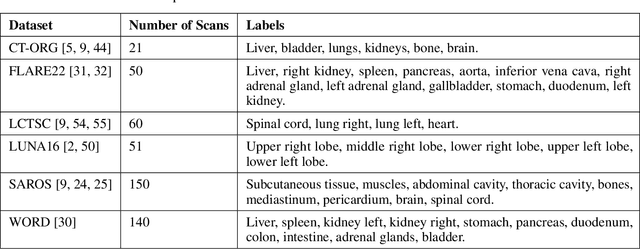
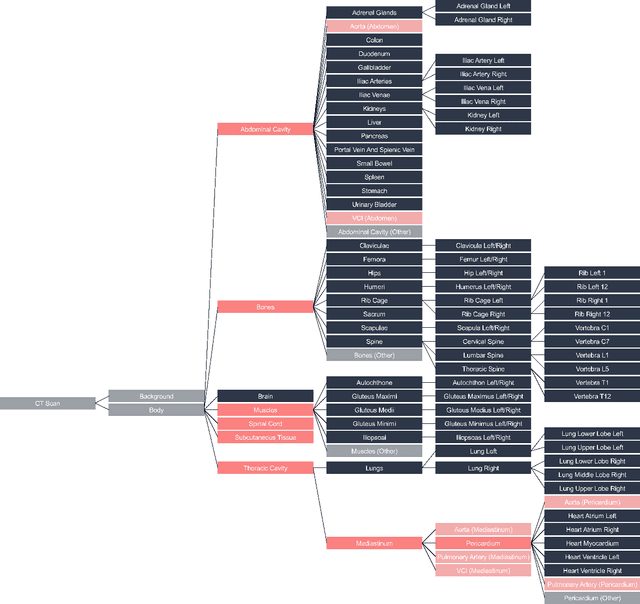
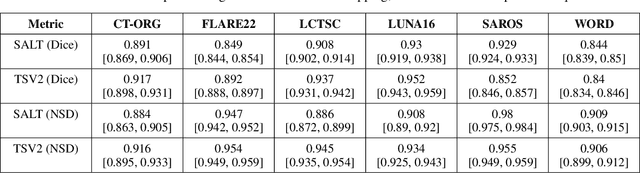
Abstract:Traditional segmentation networks approach anatomical structures as standalone elements, overlooking the intrinsic hierarchical connections among them. This study introduces Softmax for Arbitrary Label Trees (SALT), a novel approach designed to leverage the hierarchical relationships between labels, improving the efficiency and interpretability of the segmentations. This study introduces a novel segmentation technique for CT imaging, which leverages conditional probabilities to map the hierarchical structure of anatomical landmarks, such as the spine's division into lumbar, thoracic, and cervical regions and further into individual vertebrae. The model was developed using the SAROS dataset from The Cancer Imaging Archive (TCIA), comprising 900 body region segmentations from 883 patients. The dataset was further enhanced by generating additional segmentations with the TotalSegmentator, for a total of 113 labels. The model was trained on 600 scans, while validation and testing were conducted on 150 CT scans. Performance was assessed using the Dice score across various datasets, including SAROS, CT-ORG, FLARE22, LCTSC, LUNA16, and WORD. Among the evaluated datasets, SALT achieved its best results on the LUNA16 and SAROS datasets, with Dice scores of 0.93 and 0.929 respectively. The model demonstrated reliable accuracy across other datasets, scoring 0.891 on CT-ORG and 0.849 on FLARE22. The LCTSC dataset showed a score of 0.908 and the WORD dataset also showed good performance with a score of 0.844. SALT used the hierarchical structures inherent in the human body to achieve whole-body segmentations with an average of 35 seconds for 100 slices. This rapid processing underscores its potential for integration into clinical workflows, facilitating the automatic and efficient computation of full-body segmentations with each CT scan, thus enhancing diagnostic processes and patient care.
Why does my medical AI look at pictures of birds? Exploring the efficacy of transfer learning across domain boundaries
Jun 30, 2023Abstract:It is an open secret that ImageNet is treated as the panacea of pretraining. Particularly in medical machine learning, models not trained from scratch are often finetuned based on ImageNet-pretrained models. We posit that pretraining on data from the domain of the downstream task should almost always be preferred instead. We leverage RadNet-12M, a dataset containing more than 12 million computed tomography (CT) image slices, to explore the efficacy of self-supervised pretraining on medical and natural images. Our experiments cover intra- and cross-domain transfer scenarios, varying data scales, finetuning vs. linear evaluation, and feature space analysis. We observe that intra-domain transfer compares favorably to cross-domain transfer, achieving comparable or improved performance (0.44% - 2.07% performance increase using RadNet pretraining, depending on the experiment) and demonstrate the existence of a domain boundary-related generalization gap and domain-specific learned features.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge