R. Nils Planken
Fighting MRI Anisotropy: Learning Multiple Cardiac Shapes From a Single Implicit Neural Representation
Feb 11, 2026Abstract:The anisotropic nature of short-axis (SAX) cardiovascular magnetic resonance imaging (CMRI) limits cardiac shape analysis. To address this, we propose to leverage near-isotropic, higher resolution computed tomography angiography (CTA) data of the heart. We use this data to train a single neural implicit function to jointly represent cardiac shapes from CMRI at any resolution. We evaluate the method for the reconstruction of right ventricle (RV) and myocardium (MYO), where MYO simultaneously models endocardial and epicardial left-ventricle surfaces. Since high-resolution SAX reference segmentations are unavailable, we evaluate performance by extracting a 4-chamber (4CH) slice of RV and MYO from their reconstructed shapes. When compared with the reference 4CH segmentation masks from CMRI, our method achieved a Dice similarity coefficient of 0.91 $\pm$ 0.07 and 0.75 $\pm$ 0.13, and a Hausdorff distance of 6.21 $\pm$ 3.97 mm and 7.53 $\pm$ 5.13 mm for RV and MYO, respectively. Quantitative and qualitative assessment demonstrate the model's ability to reconstruct accurate, smooth and anatomically plausible shapes, supporting improvements in cardiac shape analysis.
Neural Implicit 3D Cardiac Shape Reconstruction from Sparse CT Angiography Slices Mimicking 2D Transthoracic Echocardiography Views
Feb 05, 2026Abstract:Accurate 3D representations of cardiac structures allow quantitative analysis of anatomy and function. In this work, we propose a method for reconstructing complete 3D cardiac shapes from segmentations of sparse planes in CT angiography (CTA) for application in 2D transthoracic echocardiography (TTE). Our method uses a neural implicit function to reconstruct the 3D shape of the cardiac chambers and left-ventricle myocardium from sparse CTA planes. To investigate the feasibility of achieving 3D reconstruction from 2D TTE, we select planes that mimic the standard apical 2D TTE views. During training, a multi-layer perceptron learns shape priors from 3D segmentations of the target structures in CTA. At test time, the network reconstructs 3D cardiac shapes from segmentations of TTE-mimicking CTA planes by jointly optimizing the latent code and the rigid transforms that map the observed planes into 3D space. For each heart, we simulate four realistic apical views, and we compare reconstructed multi-class volumes with the reference CTA volumes. On a held-out set of CTA segmentations, our approach achieves an average Dice coefficient of 0.86 $\pm$ 0.04 across all structures. Our method also achieves markedly lower volume errors than the clinical standard, Simpson's biplane rule: 4.88 $\pm$ 4.26 mL vs. 8.14 $\pm$ 6.04 mL, respectively, for the left ventricle; and 6.40 $\pm$ 7.37 mL vs. 37.76 $\pm$ 22.96 mL, respectively, for the left atrium. This suggests that our approach offers a viable route to more accurate 3D chamber quantification in 2D transthoracic echocardiography.
Classifier-guided registration of coronary CT angiography and intravascular ultrasound
Dec 22, 2024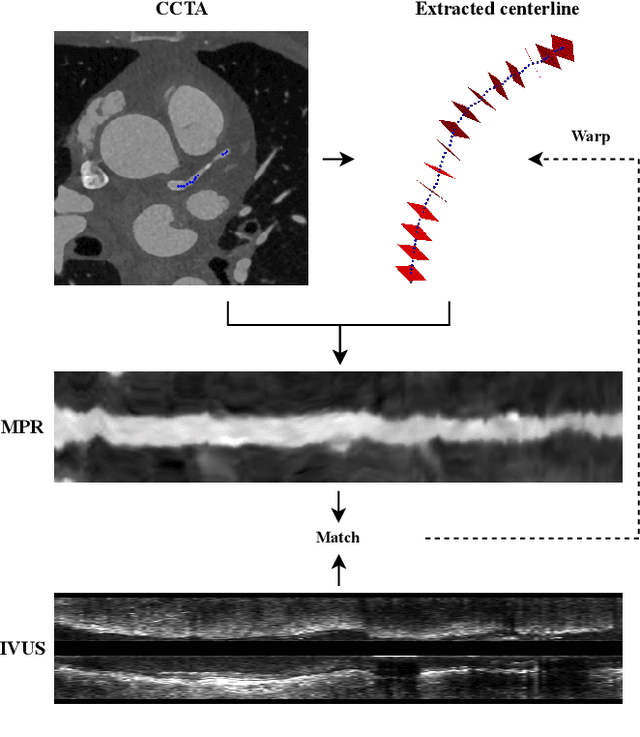
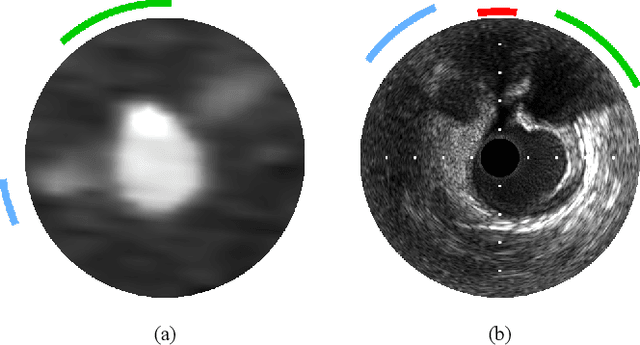
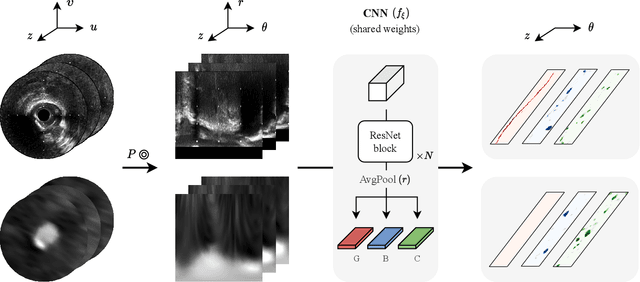
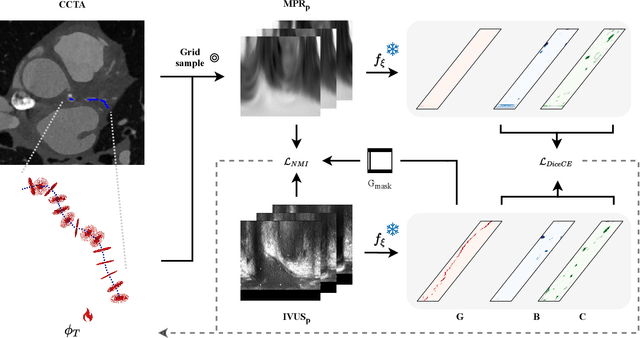
Abstract:Coronary CT angiography (CCTA) and intravascular ultrasound (IVUS) provide complementary information for coronary artery disease assessment, making their registration valuable for comprehensive analysis. However, existing registration methods require manual interaction or extensive segmentations, limiting their practical application. In this work, we present a fully automatic framework for CCTA-IVUS registration using deep learning-based feature detection and a differentiable image registration module. Our approach leverages a convolutional neural network trained to identify key anatomical features from polar-transformed multiplanar reformatted CCTA or IVUS data. These detected anatomical featuers subsequently guide a differentiable registration module to optimize transformation parameters of an automatically extracted coronary artery centerline. The method does not require landmark selection or segmentations as input, while accounting for the presence of IVUS guidewire artifacts. Evaluated on 48 clinical cases with reference CCTA centerlines corresponding to IVUS pullback, our method achieved successful registration in 83.3\% of cases, with a median centerline overlap F$_1$-score of 0.982 and median cosine similarities of 0.940 and 0.944 for cross-sectional plane orientation. Our results demonstrate that automatically detected anatomical features can be leveraged for accurate registration. The fully automatic nature of the approach represents a significant step toward streamlined multimodal coronary analysis, potentially facilitating large-scale studies of coronary plaque characteristics across modalities.
World of Forms: Deformable Geometric Templates for One-Shot Surface Meshing in Coronary CT Angiography
Sep 18, 2024Abstract:Deep learning-based medical image segmentation and surface mesh generation typically involve a sequential pipeline from image to segmentation to meshes, often requiring large training datasets while making limited use of prior geometric knowledge. This may lead to topological inconsistencies and suboptimal performance in low-data regimes. To address these challenges, we propose a data-efficient deep learning method for direct 3D anatomical object surface meshing using geometric priors. Our approach employs a multi-resolution graph neural network that operates on a prior geometric template which is deformed to fit object boundaries of interest. We show how different templates may be used for the different surface meshing targets, and introduce a novel masked autoencoder pretraining strategy for 3D spherical data. The proposed method outperforms nnUNet in a one-shot setting for segmentation of the pericardium, left ventricle (LV) cavity and the LV myocardium. Similarly, the method outperforms other lumen segmentation operating on multi-planar reformatted images. Results further indicate that mesh quality is on par with or improves upon marching cubes post-processing of voxel mask predictions, while remaining flexible in the choice of mesh triangulation prior, thus paving the way for more accurate and topologically consistent 3D medical object surface meshing.
Automatic Coronary Artery Plaque Quantification and CAD-RADS Prediction using Mesh Priors
Oct 17, 2023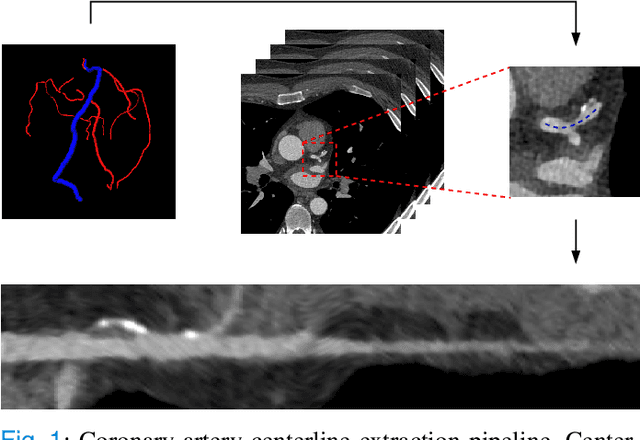
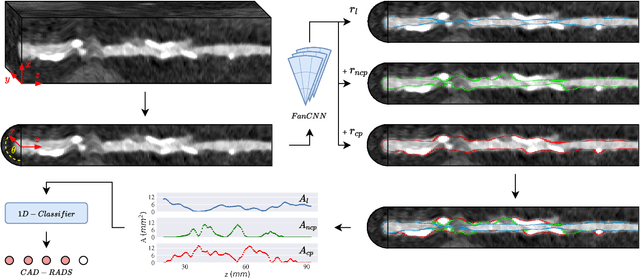
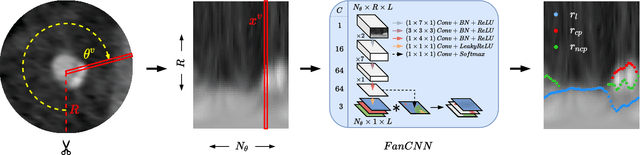
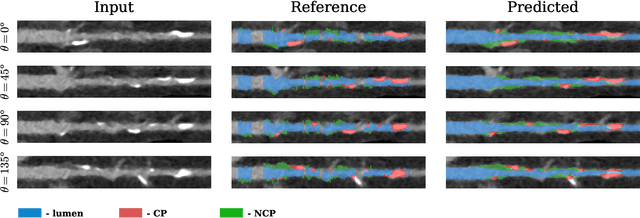
Abstract:Coronary artery disease (CAD) remains the leading cause of death worldwide. Patients with suspected CAD undergo coronary CT angiography (CCTA) to evaluate the risk of cardiovascular events and determine the treatment. Clinical analysis of coronary arteries in CCTA comprises the identification of atherosclerotic plaque, as well as the grading of any coronary artery stenosis typically obtained through the CAD-Reporting and Data System (CAD-RADS). This requires analysis of the coronary lumen and plaque. While voxel-wise segmentation is a commonly used approach in various segmentation tasks, it does not guarantee topologically plausible shapes. To address this, in this work, we propose to directly infer surface meshes for coronary artery lumen and plaque based on a centerline prior and use it in the downstream task of CAD-RADS scoring. The method is developed and evaluated using a total of 2407 CCTA scans. Our method achieved lesion-wise volume intraclass correlation coefficients of 0.98, 0.79, and 0.85 for calcified, non-calcified, and total plaque volume respectively. Patient-level CAD-RADS categorization was evaluated on a representative hold-out test set of 300 scans, for which the achieved linearly weighted kappa ($\kappa$) was 0.75. CAD-RADS categorization on the set of 658 scans from another hospital and scanner led to a $\kappa$ of 0.71. The results demonstrate that direct inference of coronary artery meshes for lumen and plaque is feasible, and allows for the automated prediction of routinely performed CAD-RADS categorization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge