Richard A. P. Takx
Automatic Coronary Artery Plaque Quantification and CAD-RADS Prediction using Mesh Priors
Oct 17, 2023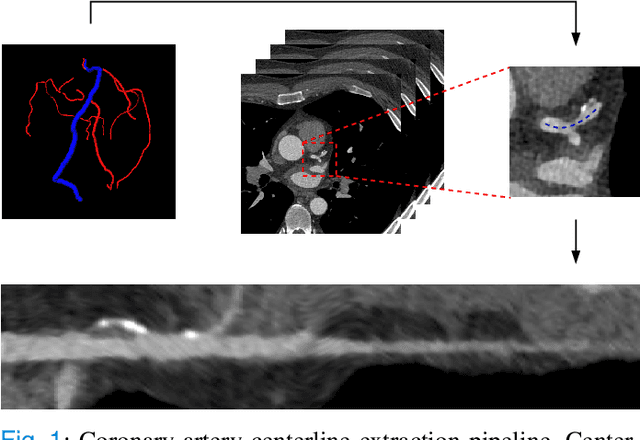
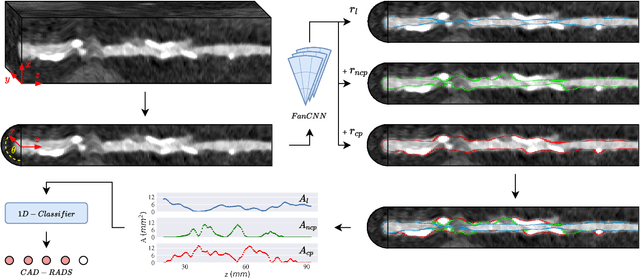
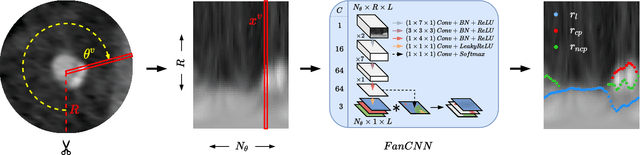
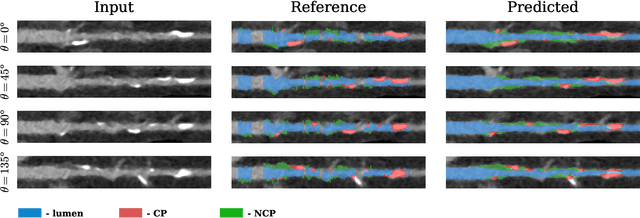
Abstract:Coronary artery disease (CAD) remains the leading cause of death worldwide. Patients with suspected CAD undergo coronary CT angiography (CCTA) to evaluate the risk of cardiovascular events and determine the treatment. Clinical analysis of coronary arteries in CCTA comprises the identification of atherosclerotic plaque, as well as the grading of any coronary artery stenosis typically obtained through the CAD-Reporting and Data System (CAD-RADS). This requires analysis of the coronary lumen and plaque. While voxel-wise segmentation is a commonly used approach in various segmentation tasks, it does not guarantee topologically plausible shapes. To address this, in this work, we propose to directly infer surface meshes for coronary artery lumen and plaque based on a centerline prior and use it in the downstream task of CAD-RADS scoring. The method is developed and evaluated using a total of 2407 CCTA scans. Our method achieved lesion-wise volume intraclass correlation coefficients of 0.98, 0.79, and 0.85 for calcified, non-calcified, and total plaque volume respectively. Patient-level CAD-RADS categorization was evaluated on a representative hold-out test set of 300 scans, for which the achieved linearly weighted kappa ($\kappa$) was 0.75. CAD-RADS categorization on the set of 658 scans from another hospital and scanner led to a $\kappa$ of 0.71. The results demonstrate that direct inference of coronary artery meshes for lumen and plaque is feasible, and allows for the automated prediction of routinely performed CAD-RADS categorization.
Deep Learning from Dual-Energy Information for Whole-Heart Segmentation in Dual-Energy and Single-Energy Non-Contrast-Enhanced Cardiac CT
Aug 10, 2020



Abstract:Deep learning-based whole-heart segmentation in coronary CT angiography (CCTA) allows the extraction of quantitative imaging measures for cardiovascular risk prediction. Automatic extraction of these measures in patients undergoing only non-contrast-enhanced CT (NCCT) scanning would be valuable. In this work, we leverage information provided by a dual-layer detector CT scanner to obtain a reference standard in virtual non-contrast (VNC) CT images mimicking NCCT images, and train a 3D convolutional neural network (CNN) for the segmentation of VNC as well as NCCT images. Contrast-enhanced acquisitions on a dual-layer detector CT scanner were reconstructed into a CCTA and a perfectly aligned VNC image. In each CCTA image, manual reference segmentations of the left ventricular (LV) myocardium, LV cavity, right ventricle, left atrium, right atrium, ascending aorta, and pulmonary artery trunk were obtained and propagated to the corresponding VNC image. These VNC images and reference segmentations were used to train 3D CNNs for automatic segmentation in either VNC images or NCCT images. Automatic segmentations in VNC images showed good agreement with reference segmentations, with an average Dice similarity coefficient of 0.897 \pm 0.034 and an average symmetric surface distance of 1.42 \pm 0.45 mm. Volume differences [95% confidence interval] between automatic NCCT and reference CCTA segmentations were -19 [-67; 30] mL for LV myocardium, -25 [-78; 29] mL for LV cavity, -29 [-73; 14] mL for right ventricle, -20 [-62; 21] mL for left atrium, and -19 [-73; 34] mL for right atrium, respectively. In 214 (74%) NCCT images from an independent multi-vendor multi-center set, two observers agreed that the automatic segmentation was mostly accurate or better. This method might enable quantification of additional cardiac measures from NCCT images for improved cardiovascular risk prediction.
Deep Learning-Based Regression and Classification for Automatic Landmark Localization in Medical Images
Jul 10, 2020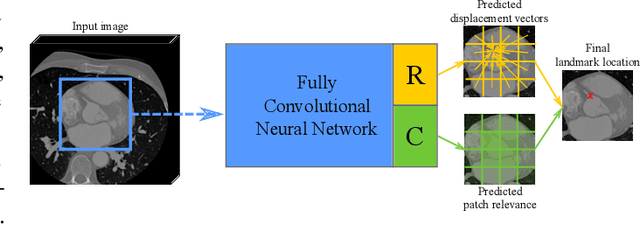
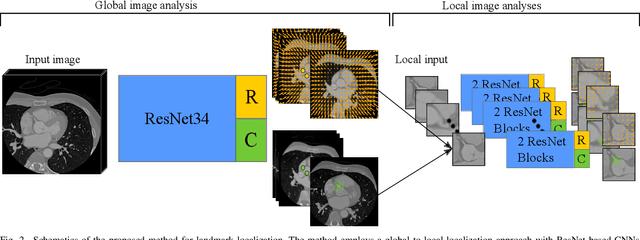
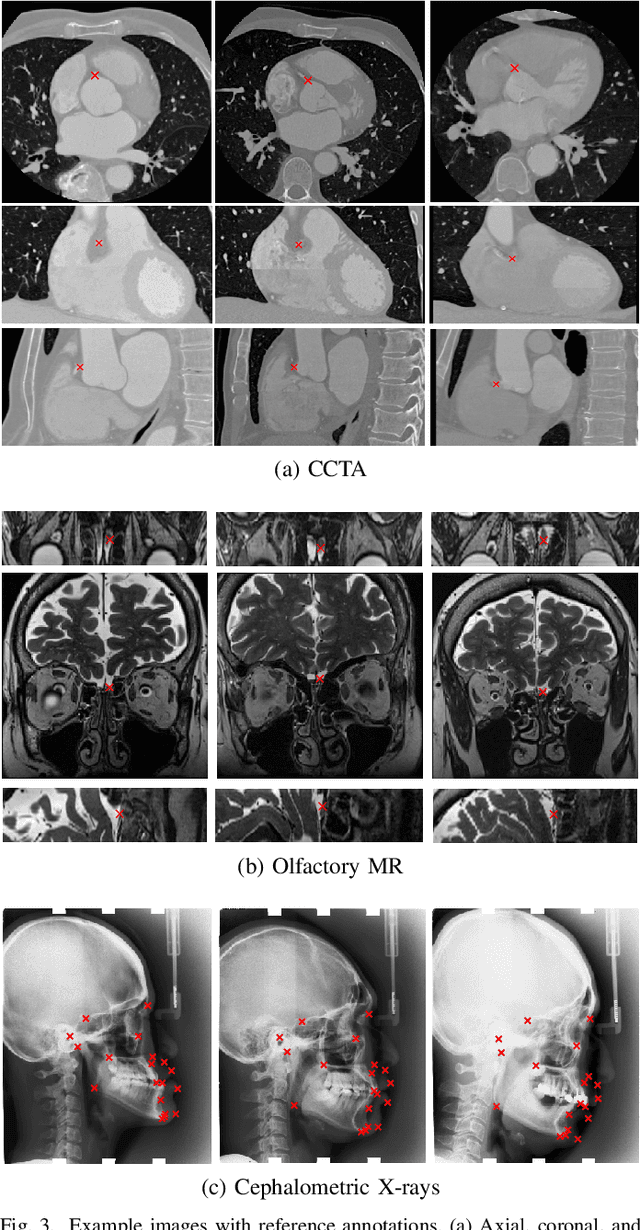
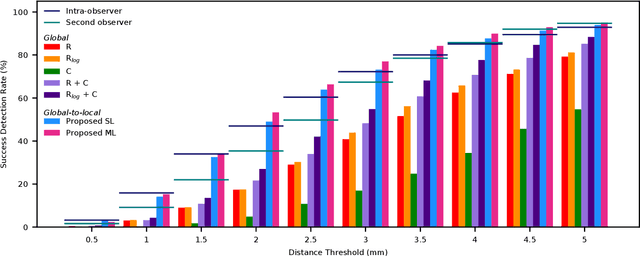
Abstract:In this study, we propose a fast and accurate method to automatically localize anatomical landmarks in medical images. We employ a global-to-local localization approach using fully convolutional neural networks (FCNNs). First, a global FCNN localizes multiple landmarks through the analysis of image patches, performing regression and classification simultaneously. In regression, displacement vectors pointing from the center of image patches towards landmark locations are determined. In classification, presence of landmarks of interest in the patch is established. Global landmark locations are obtained by averaging the predicted displacement vectors, where the contribution of each displacement vector is weighted by the posterior classification probability of the patch that it is pointing from. Subsequently, for each landmark localized with global localization, local analysis is performed. Specialized FCNNs refine the global landmark locations by analyzing local sub-images in a similar manner, i.e. by performing regression and classification simultaneously and combining the results. Evaluation was performed through localization of 8 anatomical landmarks in CCTA scans, 2 landmarks in olfactory MR scans, and 19 landmarks in cephalometric X-rays. We demonstrate that the method performs similarly to a second observer and is able to localize landmarks in a diverse set of medical images, differing in image modality, image dimensionality, and anatomical coverage.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge