Julia M. H. Noothout
Generative Models for Reproducible Coronary Calcium Scoring
May 24, 2022
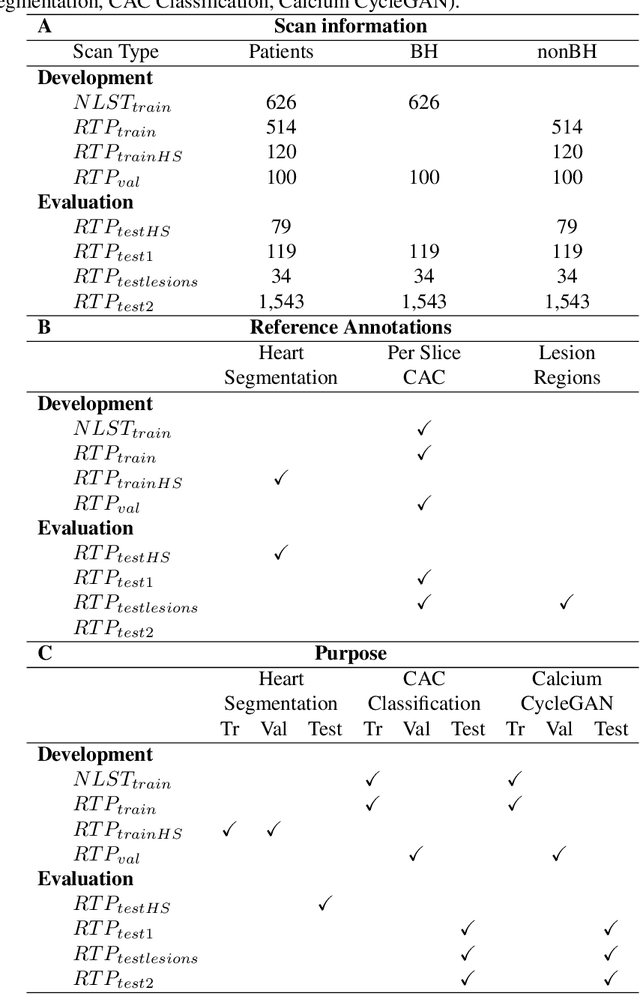


Abstract:Purpose: Coronary artery calcium (CAC) score, i.e. the amount of CAC quantified in CT, is a strong and independent predictor of coronary heart disease (CHD) events. However, CAC scoring suffers from limited interscan reproducibility, which is mainly due to the clinical definition requiring application of a fixed intensity level threshold for segmentation of calcifications. This limitation is especially pronounced in non-ECG-synchronized CT where lesions are more impacted by cardiac motion and partial volume effects. Therefore, we propose a CAC quantification method that does not require a threshold for segmentation of CAC. Approach: Our method utilizes a generative adversarial network where a CT with CAC is decomposed into an image without CAC and an image showing only CAC. The method, using a CycleGAN, was trained using 626 low-dose chest CTs and 514 radiotherapy treatment planning CTs. Interscan reproducibility was compared to clinical calcium scoring in radiotherapy treatment planning CTs of 1,662 patients, each having two scans. Results: A lower relative interscan difference in CAC mass was achieved by the proposed method: 47% compared to 89% manual clinical calcium scoring. The intraclass correlation coefficient of Agatston scores was 0.96 for the proposed method compared to 0.91 for automatic clinical calcium scoring. Conclusions: The increased interscan reproducibility achieved by our method may lead to increased reliability of CHD risk categorization and improved accuracy of CHD event prediction.
* In press
Deep Learning-Based Regression and Classification for Automatic Landmark Localization in Medical Images
Jul 10, 2020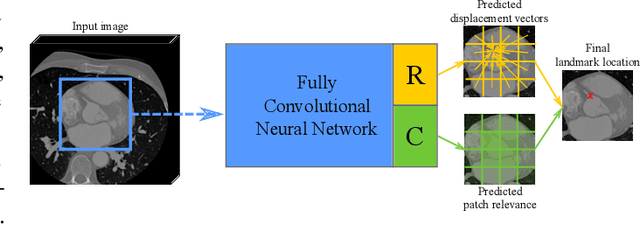
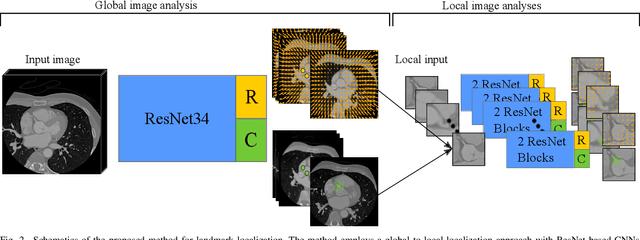
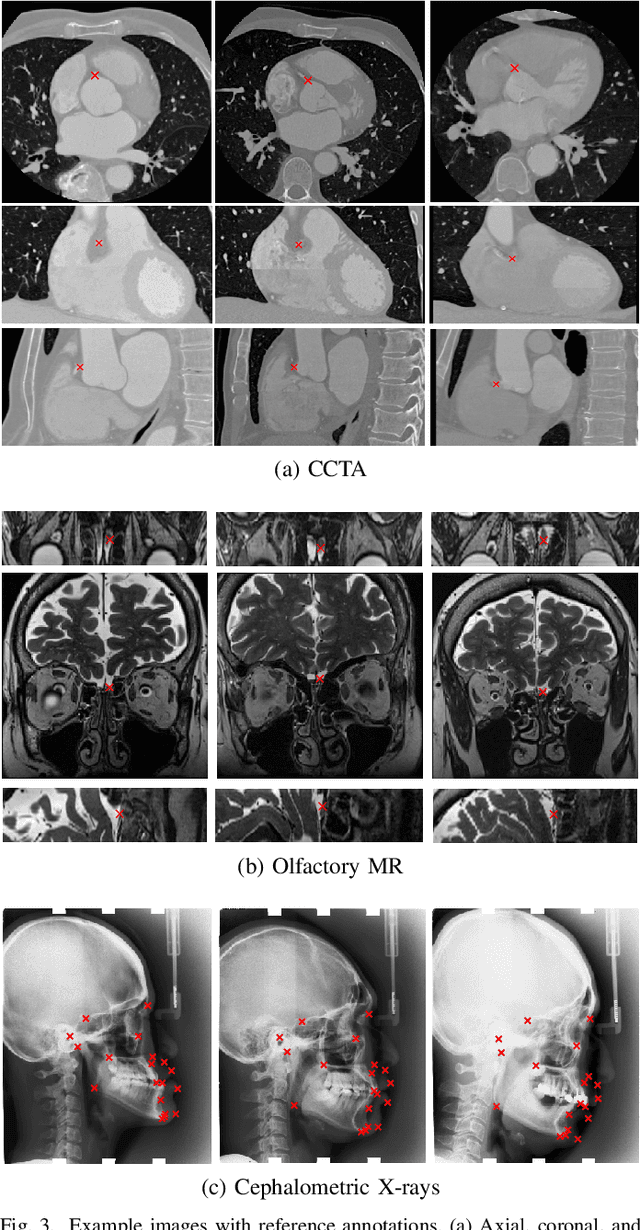
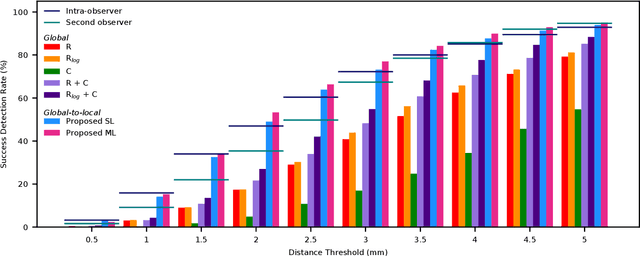
Abstract:In this study, we propose a fast and accurate method to automatically localize anatomical landmarks in medical images. We employ a global-to-local localization approach using fully convolutional neural networks (FCNNs). First, a global FCNN localizes multiple landmarks through the analysis of image patches, performing regression and classification simultaneously. In regression, displacement vectors pointing from the center of image patches towards landmark locations are determined. In classification, presence of landmarks of interest in the patch is established. Global landmark locations are obtained by averaging the predicted displacement vectors, where the contribution of each displacement vector is weighted by the posterior classification probability of the patch that it is pointing from. Subsequently, for each landmark localized with global localization, local analysis is performed. Specialized FCNNs refine the global landmark locations by analyzing local sub-images in a similar manner, i.e. by performing regression and classification simultaneously and combining the results. Evaluation was performed through localization of 8 anatomical landmarks in CCTA scans, 2 landmarks in olfactory MR scans, and 19 landmarks in cephalometric X-rays. We demonstrate that the method performs similarly to a second observer and is able to localize landmarks in a diverse set of medical images, differing in image modality, image dimensionality, and anatomical coverage.
Automatic Segmentation of Thoracic Aorta Segments in Low-Dose Chest CT
Oct 09, 2018Abstract:Morphological analysis and identification of pathologies in the aorta are important for cardiovascular diagnosis and risk assessment in patients. Manual annotation is time-consuming and cumbersome in CT scans acquired without contrast enhancement and with low radiation dose. Hence, we propose an automatic method to segment the ascending aorta, the aortic arch and the thoracic descending aorta in low-dose chest CT without contrast enhancement. Segmentation was performed using a dilated convolutional neural network (CNN), with a receptive field of 131X131 voxels, that classified voxels in axial, coronal and sagittal image slices. To obtain a final segmentation, the obtained probabilities of the three planes were averaged per class, and voxels were subsequently assigned to the class with the highest class probability. Two-fold cross-validation experiments were performed where ten scans were used to train the network and another ten to evaluate the performance. Dice coefficients of 0.83, 0.86 and 0.88, and Average Symmetrical Surface Distances (ASSDs) of 2.44, 1.56 and 1.87 mm were obtained for the ascending aorta, the aortic arch, and the descending aorta, respectively. The results indicate that the proposed method could be used in large-scale studies analyzing the anatomical location of pathology and morphology of the thoracic aorta.
CNN-based Landmark Detection in Cardiac CTA Scans
Apr 13, 2018



Abstract:Fast and accurate anatomical landmark detection can benefit many medical image analysis methods. Here, we propose a method to automatically detect anatomical landmarks in medical images. Automatic landmark detection is performed with a patch-based fully convolutional neural network (FCNN) that combines regression and classification. For any given image patch, regression is used to predict the 3D displacement vector from the image patch to the landmark. Simultaneously, classification is used to identify patches that contain the landmark. Under the assumption that patches close to a landmark can determine the landmark location more precisely than patches farther from it, only those patches that contain the landmark according to classification are used to determine the landmark location. The landmark location is obtained by calculating the average landmark location using the computed 3D displacement vectors. The method is evaluated using detection of six clinically relevant landmarks in coronary CT angiography (CCTA) scans: the right and left ostium, the bifurcation of the left main coronary artery (LM) into the left anterior descending and the left circumflex artery, and the origin of the right, non-coronary, and left aortic valve commissure. The proposed method achieved an average Euclidean distance error of 2.19 mm and 2.88 mm for the right and left ostium respectively, 3.78 mm for the bifurcation of the LM, and 1.82 mm, 2.10 mm and 1.89 mm for the origin of the right, non-coronary, and left aortic valve commissure respectively, demonstrating accurate performance. The proposed combination of regression and classification can be used to accurately detect landmarks in CCTA scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge