Naofumi Tomita
A Petri Dish for Histopathology Image Analysis
Jan 29, 2021
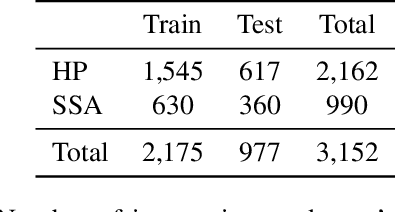
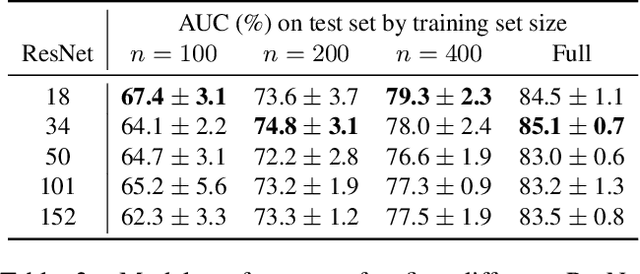
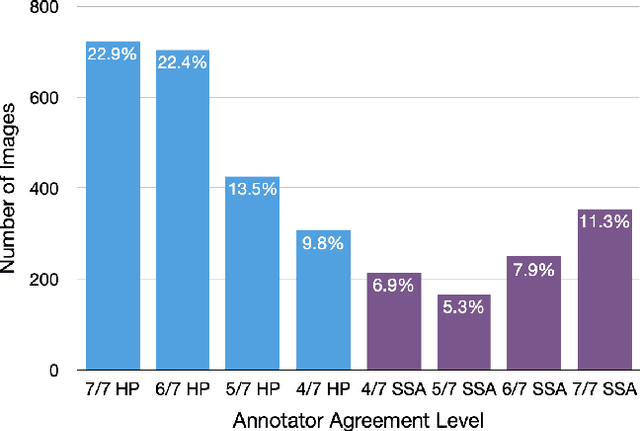
Abstract:With the rise of deep learning, there has been increased interest in using neural networks for histopathology image analysis, a field that investigates the properties of biopsy or resected specimens that are traditionally manually examined under a microscope by pathologists. In histopathology image analysis, however, challenges such as limited data, costly annotation, and processing high-resolution and variable-size images create a high barrier of entry and make it difficult to quickly iterate over model designs. Throughout scientific history, many significant research directions have leveraged small-scale experimental setups as petri dishes to efficiently evaluate exploratory ideas, which are then validated in large-scale applications. For instance, the Drosophila fruit fly in genetics and MNIST in computer vision are well-known petri dishes. In this paper, we introduce a minimalist histopathology image analysis dataset (MHIST), an analogous petri dish for histopathology image analysis. MHIST is a binary classification dataset of 3,152 fixed-size images of colorectal polyps, each with a gold-standard label determined by the majority vote of seven board-certified gastrointestinal pathologists and annotator agreement level. MHIST occupies less than 400 MB of disk space, and a ResNet-18 baseline can be trained to convergence on MHIST in just 6 minutes using 3.5 GB of memory on a NVIDIA RTX 3090. As example use cases, we use MHIST to study natural questions such as how dataset size, network depth, transfer learning, and high-disagreement examples affect model performance. By introducing MHIST, we hope to not only help facilitate the work of current histopathology imaging researchers, but also make histopathology image analysis more accessible to the general computer vision community. Our dataset is available at https://bmirds.github.io/MHIST.
Development and Evaluation of a Deep Neural Network for Histologic Classification of Renal Cell Carcinoma on Biopsy and Surgical Resection Slides
Oct 30, 2020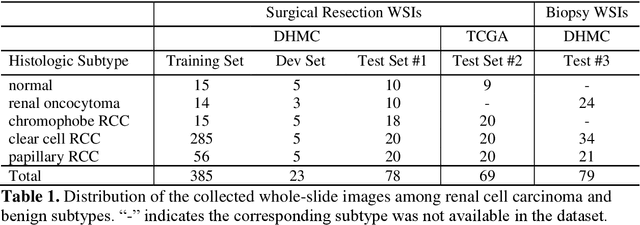
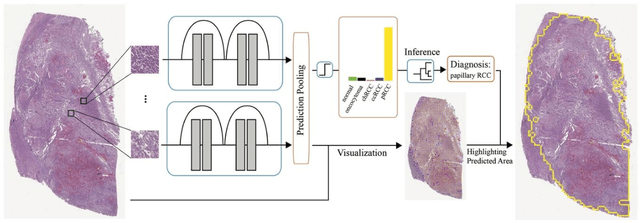

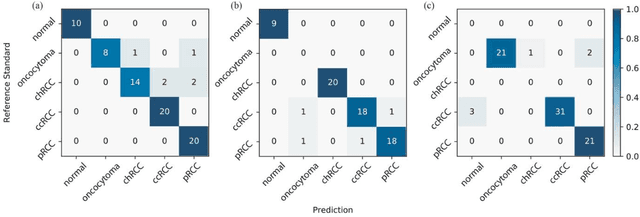
Abstract:Renal cell carcinoma (RCC) is the most common renal cancer in adults. The histopathologic classification of RCC is essential for diagnosis, prognosis, and management of patients. Reorganization and classification of complex histologic patterns of RCC on biopsy and surgical resection slides under a microscope remains a heavily specialized, error-prone, and time-consuming task for pathologists. In this study, we developed a deep neural network model that can accurately classify digitized surgical resection slides and biopsy slides into five related classes: clear cell RCC, papillary RCC, chromophobe RCC, renal oncocytoma, and normal. In addition to the whole-slide classification pipeline, we visualized the identified indicative regions and features on slides for classification by reprocessing patch-level classification results to ensure the explainability of our diagnostic model. We evaluated our model on independent test sets of 78 surgical resection whole slides and 79 biopsy slides from our tertiary medical institution, and 69 randomly selected surgical resection slides from The Cancer Genome Atlas (TCGA) database. The average area under the curve (AUC) of our classifier on the internal resection slides, internal biopsy slides, and external TCGA slides is 0.98, 0.98 and 0.99, respectively. Our results suggest that the high generalizability of our approach across different data sources and specimen types. More importantly, our model has the potential to assist pathologists by (1) automatically pre-screening slides to reduce false-negative cases, (2) highlighting regions of importance on digitized slides to accelerate diagnosis, and (3) providing objective and accurate diagnosis as the second opinion.
Learn like a Pathologist: Curriculum Learning by Annotator Agreement for Histopathology Image Classification
Sep 29, 2020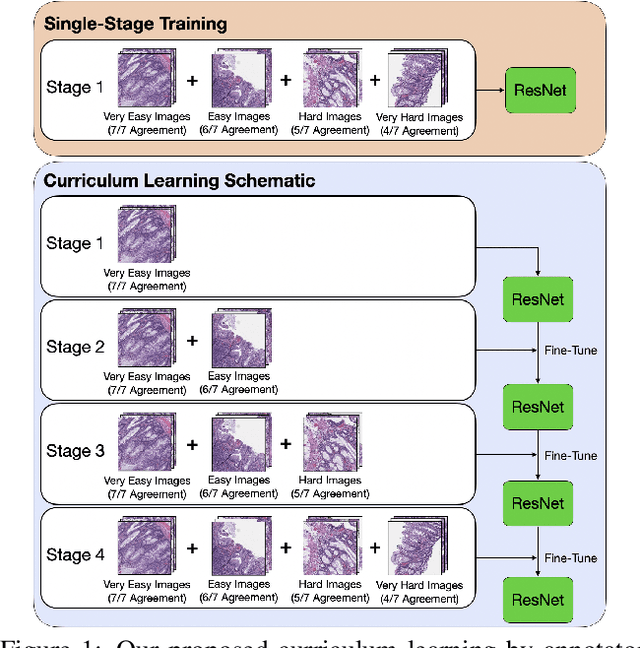
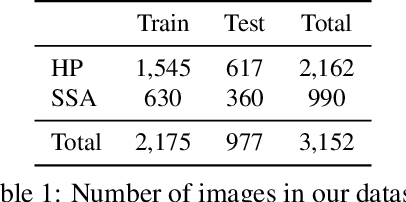
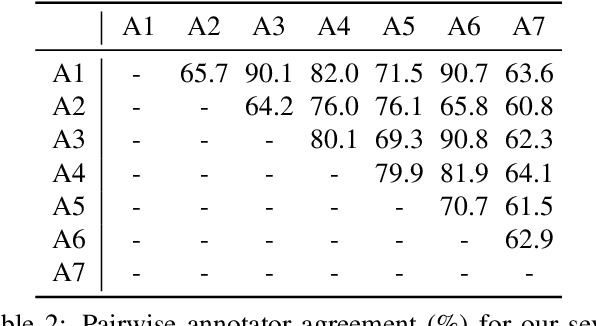
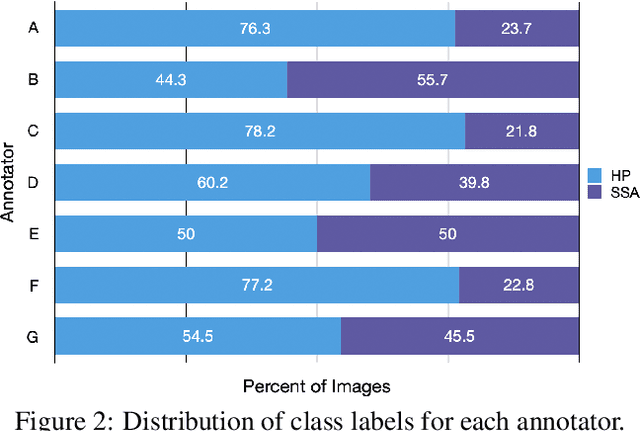
Abstract:Applying curriculum learning requires both a range of difficulty in data and a method for determining the difficulty of examples. In many tasks, however, satisfying these requirements can be a formidable challenge. In this paper, we contend that histopathology image classification is a compelling use case for curriculum learning. Based on the nature of histopathology images, a range of difficulty inherently exists among examples, and, since medical datasets are often labeled by multiple annotators, annotator agreement can be used as a natural proxy for the difficulty of a given example. Hence, we propose a simple curriculum learning method that trains on progressively-harder images as determined by annotator agreement. We evaluate our hypothesis on the challenging and clinically-important task of colorectal polyp classification. Whereas vanilla training achieves an AUC of 83.7% for this task, a model trained with our proposed curriculum learning approach achieves an AUC of 88.2%, an improvement of 4.5%. Our work aims to inspire researchers to think more creatively and rigorously when choosing contexts for applying curriculum learning.
Difficulty Translation in Histopathology Images
Apr 27, 2020

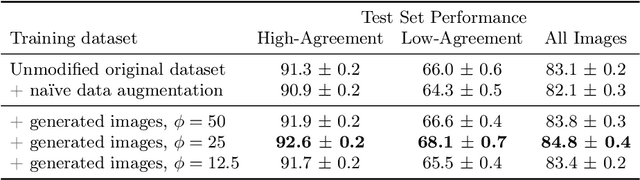
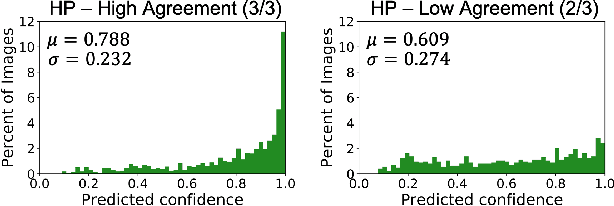
Abstract:The unique nature of histopathology images opens the door to domain-specific formulations of image translation models. We propose a difficulty translation model that modifies colorectal histopathology images to become more challenging to classify. Our model comprises a scorer, which provides an output confidence to measure the difficulty of images, and an image translator, which learns to translate images from easy-to-classify to hard-to-classify using a training set defined by the scorer. We present three findings. First, generated images were indeed harder to classify for both human pathologists and machine learning classifiers than their corresponding source images. Second, image classifiers trained with generated images as augmented data performed better on both easy and hard images from an independent test set. Finally, human annotator agreement and our model's measure of difficulty correlated strongly, implying that for future work requiring human annotator agreement, the confidence score of a machine learning classifier could be used instead as a proxy.
Multi-Ontology Refined Embeddings (MORE): A Hybrid Multi-Ontology and Corpus-based Semantic Representation for Biomedical Concepts
Apr 14, 2020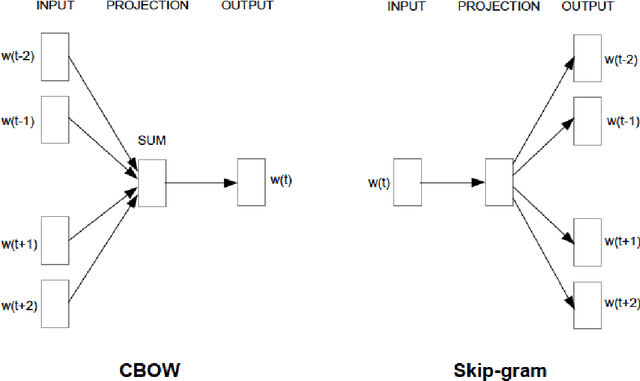
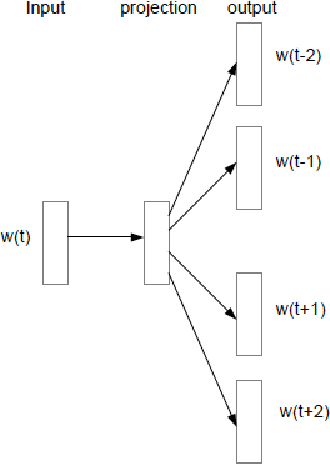
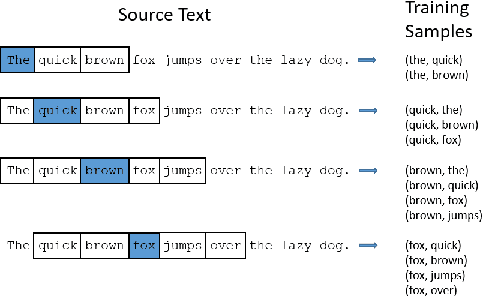
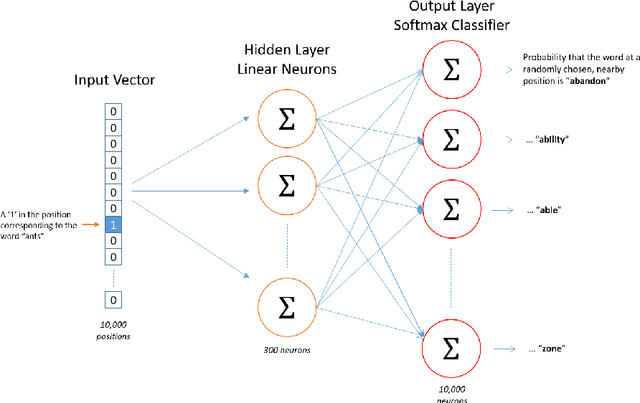
Abstract:Objective: Currently, a major limitation for natural language processing (NLP) analyses in clinical applications is that a concept can be referenced in various forms across different texts. This paper introduces Multi-Ontology Refined Embeddings (MORE), a novel hybrid framework for incorporating domain knowledge from multiple ontologies into a distributional semantic model, learned from a corpus of clinical text. Materials and Methods: We use the RadCore and MIMIC-III free-text datasets for the corpus-based component of MORE. For the ontology-based part, we use the Medical Subject Headings (MeSH) ontology and three state-of-the-art ontology-based similarity measures. In our approach, we propose a new learning objective, modified from the Sigmoid cross-entropy objective function. Results and Discussion: We evaluate the quality of the generated word embeddings using two established datasets of semantic similarities among biomedical concept pairs. On the first dataset with 29 concept pairs, with the similarity scores established by physicians and medical coders, MORE's similarity scores have the highest combined correlation (0.633), which is 5.0% higher than that of the baseline model and 12.4% higher than that of the best ontology-based similarity measure.On the second dataset with 449 concept pairs, MORE's similarity scores have a correlation of 0.481, with the average of four medical residents' similarity ratings, and that outperforms the skip-gram model by 8.1% and the best ontology measure by 6.9%.
Automatic Post-Stroke Lesion Segmentation on MR Images using 3D Residual Convolutional Neural Network
Nov 25, 2019



Abstract:In this paper, we demonstrate the feasibility and performance of deep residual neural networks for volumetric segmentation of irreversibly damaged brain tissue lesions on T1-weighted MRI scans for chronic stroke patients. A total of 239 T1-weighted MRI scans of chronic ischemic stroke patients from a public dataset were retrospectively analyzed by 3D deep convolutional segmentation models with residual learning, using a novel zoom-in&out strategy. Dice similarity coefficient (DSC), Average symmetric surface distance (ASSD), and Hausdorff distance (HD) of the identified lesions were measured by using the manual tracing of lesions as the reference standard. Bootstrapping was employed for all metrics to estimate 95% confidence intervals. The models were assessed on the test set of 31 scans. The average DSC was 0.64 (0.51-0.76) with a median of 0.78. ASSD and HD were 3.6 mm (1.7-6.2 mm) and 20.4 mm (10.0-33.3 mm), respectively. To the best of our knowledge, this performance is the highest achieved on this public dataset. The latest deep learning architecture and techniques were applied for 3D segmentation on MRI scans and demonstrated to be effective for volumetric segmentation of chronic ischemic stroke lesions.
Deep neural networks for automated classification of colorectal polyps on histopathology slides: A multi-institutional evaluation
Sep 27, 2019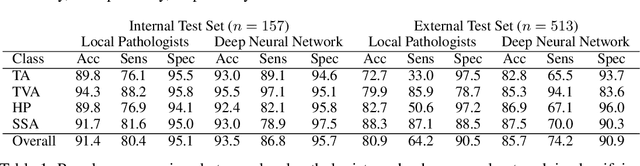
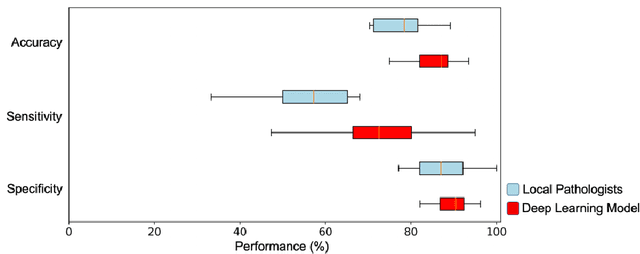
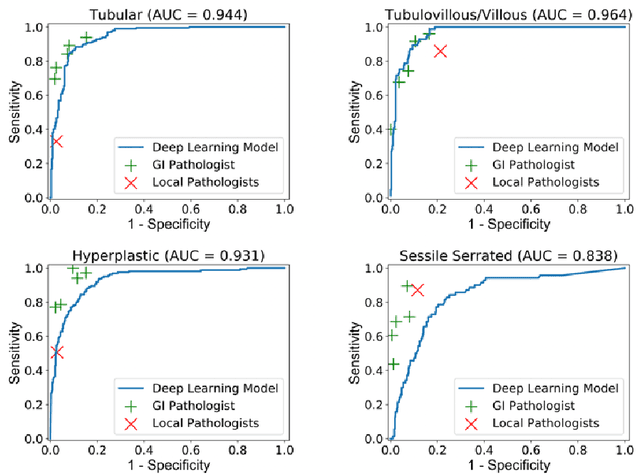
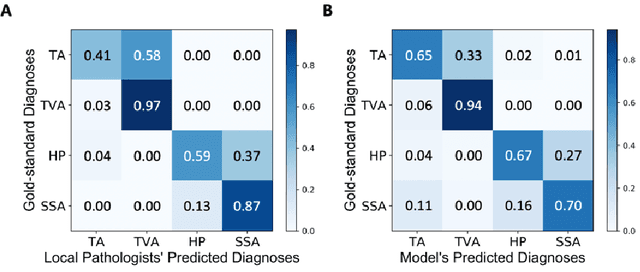
Abstract:Histological classification of colorectal polyps plays a critical role in both screening for colorectal cancer and care of affected patients. In this study, we developed a deep neural network for classification of four major colorectal polyp types on digitized histopathology slides and compared its performance to local pathologists' diagnoses at the point-of-care retrieved from corresponding pathology labs. We evaluated the deep neural network on an internal dataset of 157 histopathology slides from the Dartmouth-Hitchcock Medical Center (DHMC) in New Hampshire, as well as an external dataset of 513 histopathology slides from 24 different institutions spanning 13 states in the United States. For the internal evaluation, the deep neural network had a mean accuracy of 93.5% (95% CI 89.6%-97.4%), compared with local pathologists' accuracy of 91.4% (95% CI 87.0%-95.8%). On the external test set, the deep neural network achieved an accuracy of 85.7% (95% CI 82.7%-88.7%), significantly outperforming the accuracy of local pathologists at 80.9% (95% CI 77.5%-84.3%, p<0.05) at the point-of-care. If confirmed in clinical settings, our model could assist pathologists by improving the diagnostic efficiency, reproducibility, and accuracy of colorectal cancer screenings.
Pathologist-level classification of histologic patterns on resected lung adenocarcinoma slides with deep neural networks
Jan 31, 2019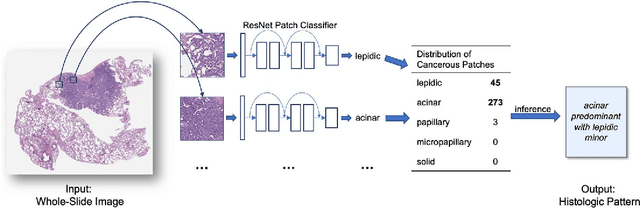
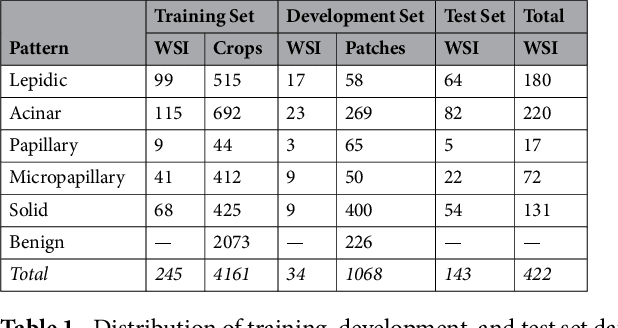
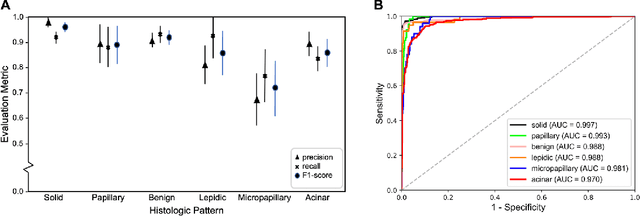
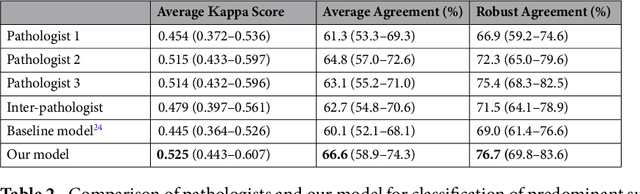
Abstract:Classification of histologic patterns in lung adenocarcinoma is critical for determining tumor grade and treatment for patients. However, this task is often challenging due to the heterogeneous nature of lung adenocarcinoma and the subjective criteria for evaluation. In this study, we propose a deep learning model that automatically classifies the histologic patterns of lung adenocarcinoma on surgical resection slides. Our model uses a convolutional neural network to identify regions of neoplastic cells, then aggregates those classifications to infer predominant and minor histologic patterns for any given whole-slide image. We evaluated our model on an independent set of 143 whole-slide images. It achieved a kappa score of 0.525 and an agreement of 66.6% with three pathologists for classifying the predominant patterns, slightly higher than the inter-pathologist kappa score of 0.485 and agreement of 62.7% on this test set. All evaluation metrics for our model and the three pathologists were within 95% confidence intervals of agreement. If confirmed in clinical practice, our model can assist pathologists in improving classification of lung adenocarcinoma patterns by automatically pre-screening and highlighting cancerous regions prior to review. Our approach can be generalized to any whole-slide image classification task, and code is made publicly available at https://github.com/BMIRDS/deepslide.
Finding a Needle in the Haystack: Attention-Based Classification of High Resolution Microscopy Images
Nov 20, 2018
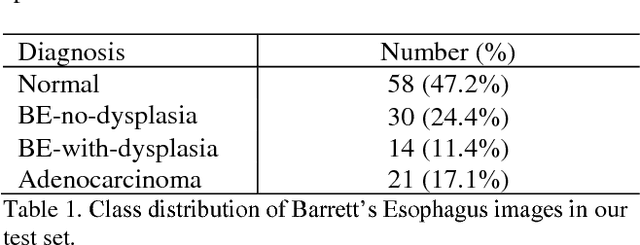

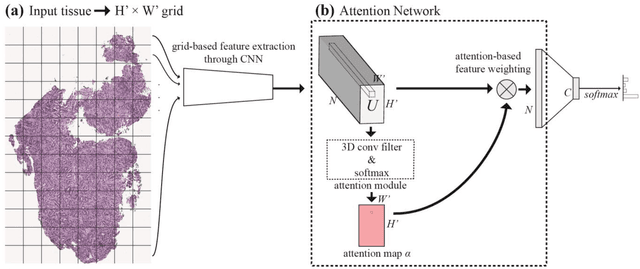
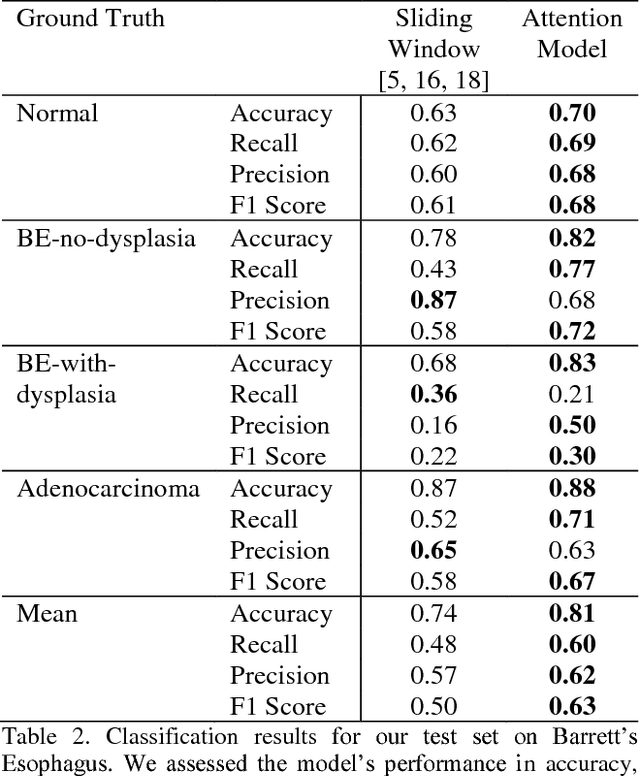
Abstract:Deep learning for classification of microscopy images is challenging because whole-slide images are high resolution. Due to the large size of these images, they cannot be transferred into GPU memory, so there are currently no end-to-end deep learning architectures for their analysis. Existing work has used a sliding window for crop classification, followed by a heuristic to determine the label for the whole slide. This pipeline is not efficient or robust, however, because crops are analyzed independently of their neighbors and the decisive features for classifying a whole slide are only found in a few regions of interest. In this paper, we present an attention-based model for classification of high resolution microscopy images. Our model dynamically finds regions of interest from a wide-view, then identifies characteristic patterns in those regions for whole-slide classification. This approach is analogous to how pathologists examine slides under the microscope and is the first to generalize the attention mechanism to high resolution images. Furthermore, our model does not require bounding box annotations for the regions of interest and is trainable end-to-end with flexible input. We evaluated our model on a microscopy dataset of Barrett's Esophagus images, and the results showed that our approach outperforms the current state-of-the-art sliding window method by a large margin.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge