Charles Brown
A Petri Dish for Histopathology Image Analysis
Jan 29, 2021
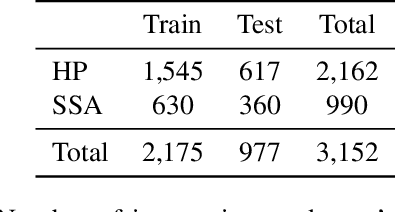
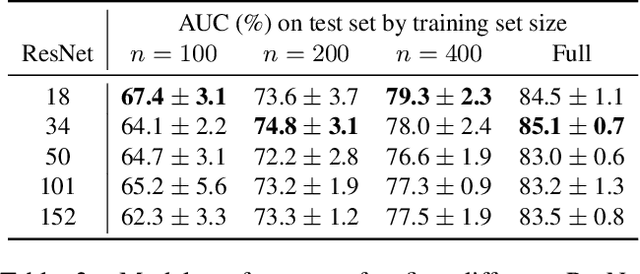
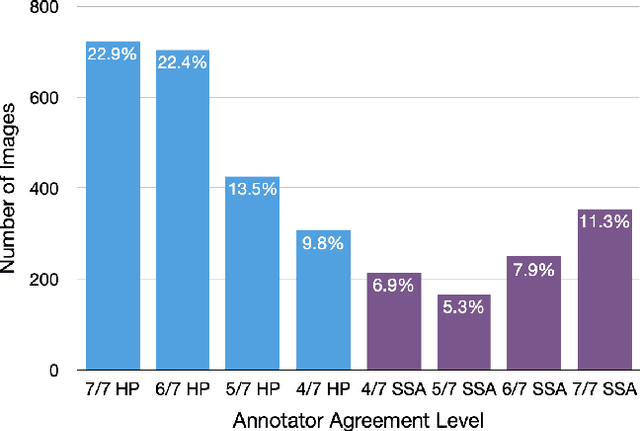
Abstract:With the rise of deep learning, there has been increased interest in using neural networks for histopathology image analysis, a field that investigates the properties of biopsy or resected specimens that are traditionally manually examined under a microscope by pathologists. In histopathology image analysis, however, challenges such as limited data, costly annotation, and processing high-resolution and variable-size images create a high barrier of entry and make it difficult to quickly iterate over model designs. Throughout scientific history, many significant research directions have leveraged small-scale experimental setups as petri dishes to efficiently evaluate exploratory ideas, which are then validated in large-scale applications. For instance, the Drosophila fruit fly in genetics and MNIST in computer vision are well-known petri dishes. In this paper, we introduce a minimalist histopathology image analysis dataset (MHIST), an analogous petri dish for histopathology image analysis. MHIST is a binary classification dataset of 3,152 fixed-size images of colorectal polyps, each with a gold-standard label determined by the majority vote of seven board-certified gastrointestinal pathologists and annotator agreement level. MHIST occupies less than 400 MB of disk space, and a ResNet-18 baseline can be trained to convergence on MHIST in just 6 minutes using 3.5 GB of memory on a NVIDIA RTX 3090. As example use cases, we use MHIST to study natural questions such as how dataset size, network depth, transfer learning, and high-disagreement examples affect model performance. By introducing MHIST, we hope to not only help facilitate the work of current histopathology imaging researchers, but also make histopathology image analysis more accessible to the general computer vision community. Our dataset is available at https://bmirds.github.io/MHIST.
Learn like a Pathologist: Curriculum Learning by Annotator Agreement for Histopathology Image Classification
Sep 29, 2020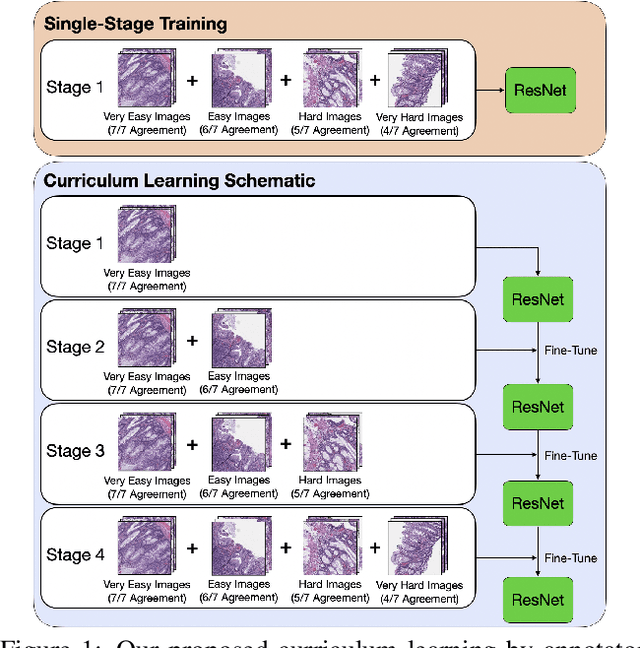
Abstract:Applying curriculum learning requires both a range of difficulty in data and a method for determining the difficulty of examples. In many tasks, however, satisfying these requirements can be a formidable challenge. In this paper, we contend that histopathology image classification is a compelling use case for curriculum learning. Based on the nature of histopathology images, a range of difficulty inherently exists among examples, and, since medical datasets are often labeled by multiple annotators, annotator agreement can be used as a natural proxy for the difficulty of a given example. Hence, we propose a simple curriculum learning method that trains on progressively-harder images as determined by annotator agreement. We evaluate our hypothesis on the challenging and clinically-important task of colorectal polyp classification. Whereas vanilla training achieves an AUC of 83.7% for this task, a model trained with our proposed curriculum learning approach achieves an AUC of 88.2%, an improvement of 4.5%. Our work aims to inspire researchers to think more creatively and rigorously when choosing contexts for applying curriculum learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge