Meike Vernooij
Reproducible White Matter Tract Segmentation Using 3D U-Net on a Large-scale DTI Dataset
Aug 26, 2019
Abstract:Tract-specific diffusion measures, as derived from brain diffusion MRI, have been linked to white matter tract structural integrity and neurodegeneration. As a consequence, there is a large interest in the automatic segmentation of white matter tract in diffusion tensor MRI data. Methods based on the tractography are popular for white matter tract segmentation. However, because of the limited consistency and long processing time, such methods may not be suitable for clinical practice. We therefore developed a novel convolutional neural network based method to directly segment white matter tract trained on a low-resolution dataset of 9149 DTI images. The method is optimized on input, loss function and network architecture selections. We evaluated both segmentation accuracy and reproducibility, and reproducibility of determining tract-specific diffusion measures. The reproducibility of the method is higher than that of the reference standard and the determined diffusion measures are consistent. Therefore, we expect our method to be applicable in clinical practice and in longitudinal analysis of white matter microstructure.
A hybrid deep learning framework for integrated segmentation and registration: evaluation on longitudinal white matter tract changes
Aug 26, 2019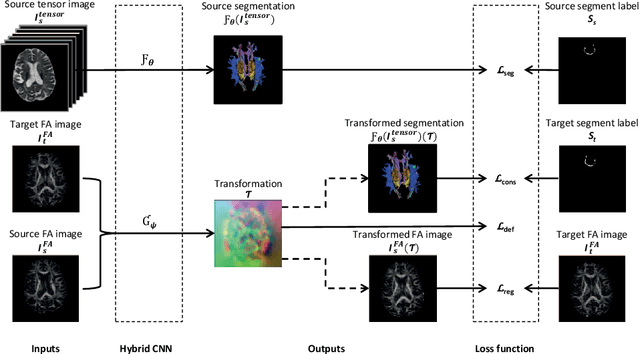
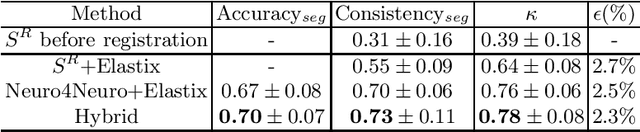

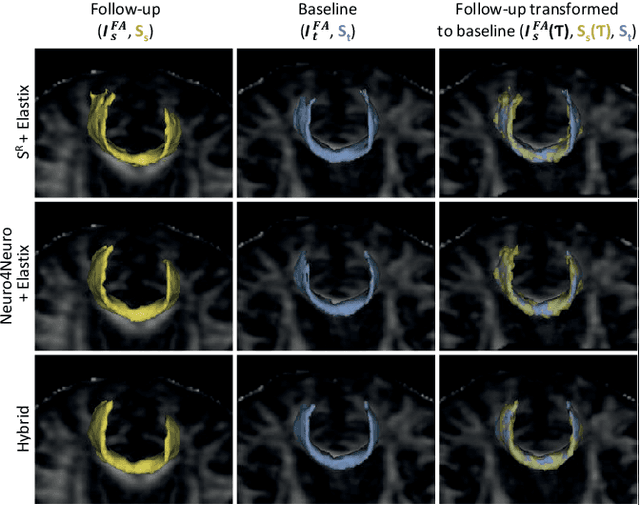
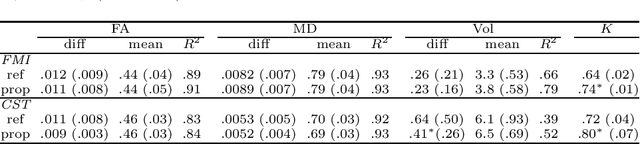
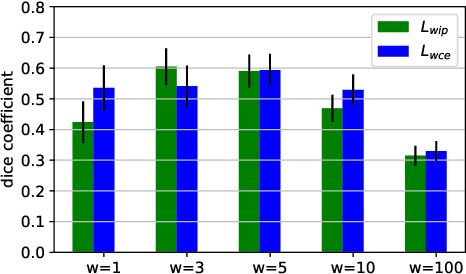
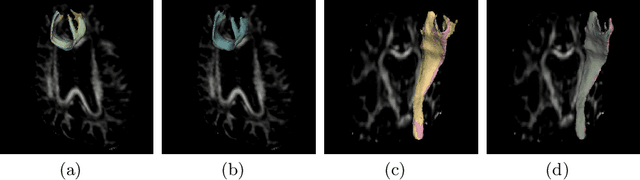
Abstract:To accurately analyze changes of anatomical structures in longitudinal imaging studies, consistent segmentation across multiple time-points is required. Existing solutions often involve independent registration and segmentation components. Registration between time-points is used either as a prior for segmentation in a subsequent time point or to perform segmentation in a common space. In this work, we propose a novel hybrid convolutional neural network (CNN) that integrates segmentation and registration into a single procedure. We hypothesize that the joint optimization leads to increased performance on both tasks. The hybrid CNN is trained by minimizing an integrated loss function composed of four different terms, measuring segmentation accuracy, similarity between registered images, deformation field smoothness, and segmentation consistency. We applied this method to the segmentation of white matter tracts, describing functionally grouped axonal fibers, using N=8045 longitudinal brain MRI data of 3249 individuals. The proposed method was compared with two multistage pipelines using two existing segmentation methods combined with a conventional deformable registration algorithm. In addition, we assessed the added value of the joint optimization for segmentation and registration separately. The hybrid CNN yielded significantly higher accuracy, consistency and reproducibility of segmentation than the multistage pipelines, and was orders of magnitude faster. Therefore, we expect it can serve as a novel tool to support clinical and epidemiological analyses on understanding microstructural brain changes over time.
Automated Image Registration Quality Assessment Utilizing Deep-learning based Ventricle Extraction in Clinical Data
Jul 01, 2019

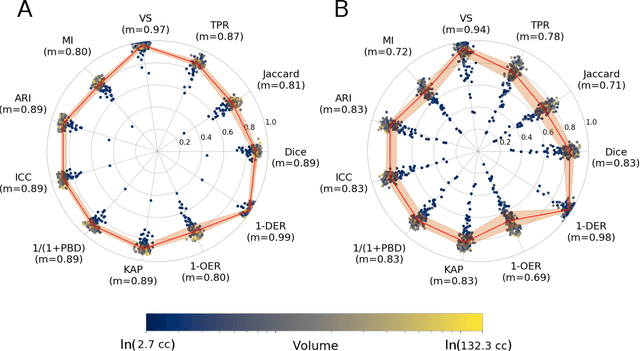
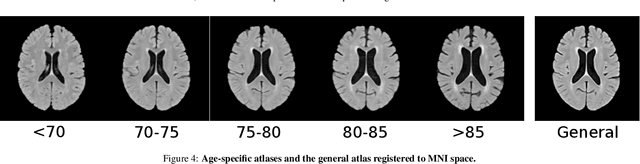
Abstract:Registration is a core component of many imaging pipelines. In case of clinical scans, with lower resolution and sometimes substantial motion artifacts, registration can produce poor results. Visual assessment of registration quality in large clinical datasets is inefficient. In this work, we propose to automatically assess the quality of registration to an atlas in clinical FLAIR MRI scans of the brain. The method consists of automatically segmenting the ventricles of a given scan using a neural network, and comparing the segmentation to the atlas' ventricles propagated to image space. We used the proposed method to improve clinical image registration to a general atlas by computing multiple registrations and then selecting the registration that yielded the highest ventricle overlap. Methods were evaluated in a single-site dataset of more than 1000 scans, as well as a multi-center dataset comprising 142 clinical scans from 12 sites. The automated ventricle segmentation reached a Dice coefficient with manual annotations of 0.89 in the single-site dataset, and 0.83 in the multi-center dataset. Registration via age-specific atlases could improve ventricle overlap compared to a direct registration to the general atlas (Dice similarity coefficient increase up to 0.15). Experiments also showed that selecting scans with the registration quality assessment method could improve the quality of average maps of white matter hyperintensity burden, instead of using all scans for the computation of the white matter hyperintensity map. In this work, we demonstrated the utility of an automated tool for assessing image registration quality in clinical scans. This image quality assessment step could ultimately assist in the translation of automated neuroimaging pipelines to the clinic.
Weakly Supervised Object Detection with 2D and 3D Regression Neural Networks
Jun 14, 2019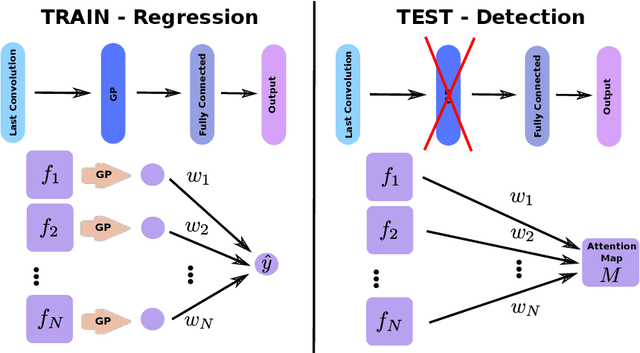
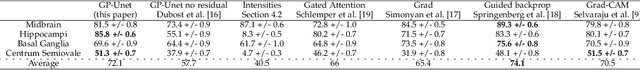
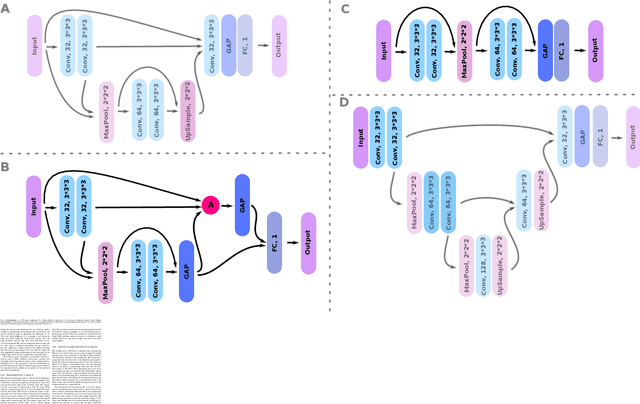
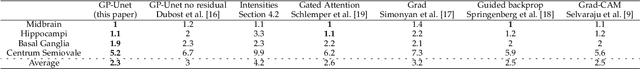
Abstract:Weakly supervised detection methods can infer the location of target objects in an image without requiring location or appearance information during training. We propose a weakly supervised deep learning method for the detection of objects that appear at multiple locations in an image. The method computes attention maps using the last feature maps of an encoder-decoder network optimized only with global labels: the number of occurrences of the target object in an image. In contrast with previous approaches, attention maps are generated at full input resolution thanks to the decoder part. The proposed approach is compared to multiple state-of-the-art methods in two tasks: the detection of digits in MNIST-based datasets, and the real life application of detection of enlarged perivascular spaces -- a type of brain lesion -- in four brain regions in a dataset of 2202 3D brain MRI scans. In MNIST-based datasets, the proposed method outperforms the other methods. In the brain dataset, several weakly supervised detection methods come close to the human intrarater agreement in each region. The proposed method reaches the lowest number of false positive detections in all brain regions at the operating point, while its average sensitivity is similar to that of the other best methods.
3D Regression Neural Network for the Quantification of Enlarged Perivascular Spaces in Brain MRI
Oct 28, 2018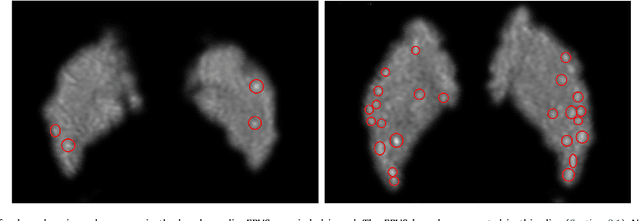
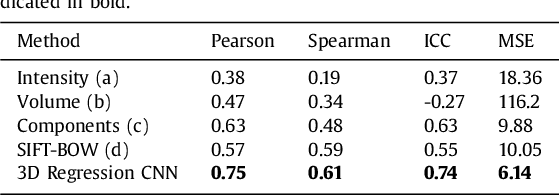
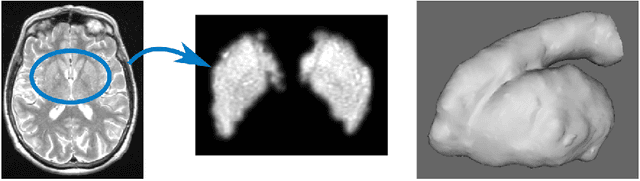
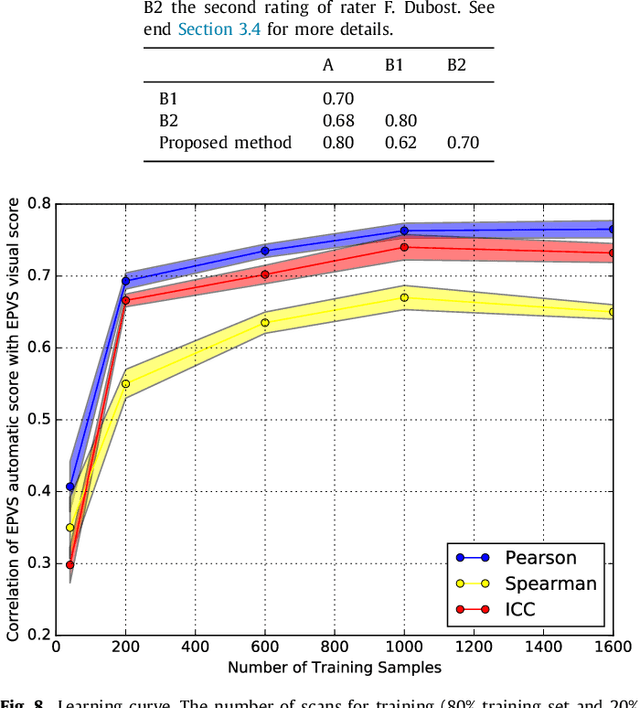
Abstract:Enlarged perivascular spaces (EPVS) in the brain are an emerging imaging marker for cerebral small vessel disease, and have been shown to be related to increased risk of various neurological diseases, including stroke and dementia. Automatic quantification of EPVS would greatly help to advance research into its etiology and its potential as a risk indicator of disease. We propose a convolutional network regression method to quantify the extent of EPVS in the basal ganglia from 3D brain MRI. We first segment the basal ganglia and subsequently apply a 3D convolutional regression network designed for small object detection within this region of interest. The network takes an image as input, and outputs a quantification score of EPVS. The network has significantly more convolution operations than pooling ones and no final activation, allowing it to span the space of real numbers. We validated our approach using a dataset of 2000 brain MRI scans scored visually. Experiments with varying sizes of training and test sets showed that a good performance can be achieved with a training set of only 200 scans. With a training set of 1000 scans, the intraclass correlation coefficient (ICC) between our scoring method and the expert's visual score was 0.74. Our method outperforms by a large margin - more than 0.10 - four more conventional automated approaches based on intensities, scale-invariant feature transform, and random forest. We show that the network learns the structures of interest and investigate the influence of hyper-parameters on the performance. We also evaluate the reproducibility of our network using a set of 60 subjects scanned twice (scan-rescan reproducibility). On this set our network achieves an ICC of 0.93, while the intrarater agreement reaches 0.80. Furthermore, the automatic EPVS scoring correlates similarly to age as visual scoring.
Hydranet: Data Augmentation for Regression Neural Networks
Jul 12, 2018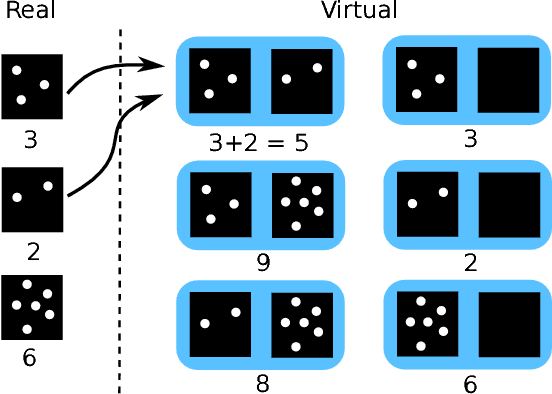
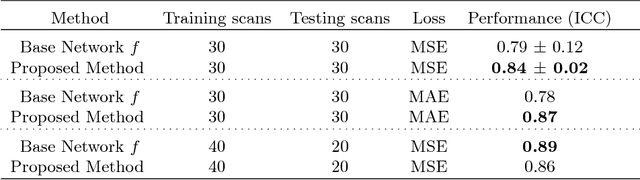
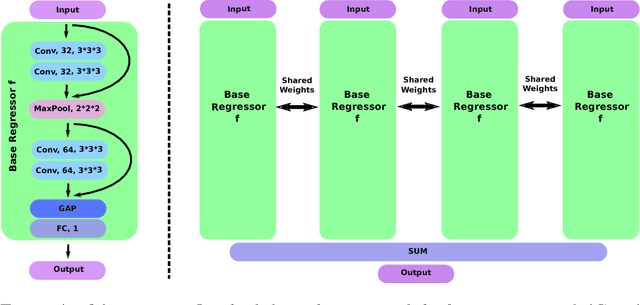
Abstract:Despite recent efforts, deep learning techniques remain often heavily dependent on a large quantity of labeled data. This problem is even more challenging in medical image analysis where the annotator expertise is often scarce. In this paper we propose a novel data-augmentation method to regularize neural network regressors, learning from a single global label per image. The principle of the method is to create new samples by recombining existing ones. We demonstrate the performance of our algorithm on two tasks: the regression of number of enlarged perivascular spaces in the basal ganglia; and the regression of white matter hyperintensities volume. We show that the proposed method improves the performance even when more basic data augmentation is used. Furthermore we reached an intraclass correlation coefficient between ground truth and network predictions of 0.73 on the first task and 0.86 on the second task, only using between 25 and 30 scans with a single global label per scan for training. To achieve a similar correlation on the first task, state-of-the-art methods needed more than 1000 training scans.
GP-Unet: Lesion Detection from Weak Labels with a 3D Regression Network
Oct 30, 2017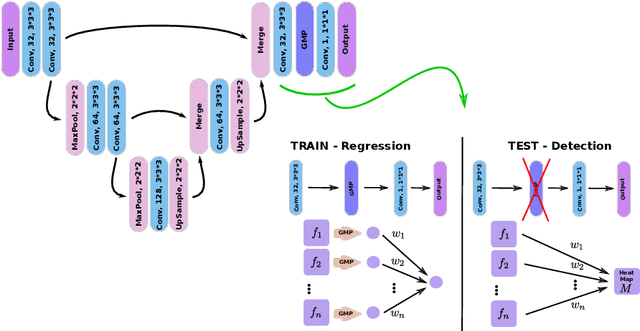
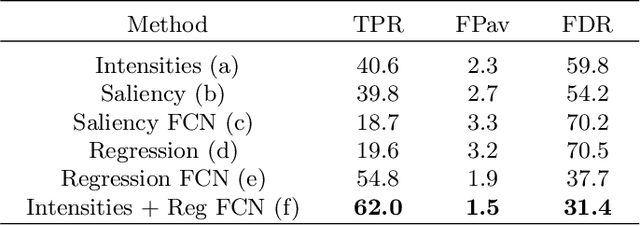
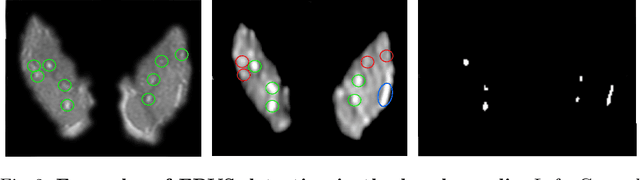
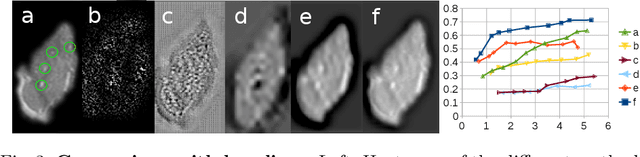
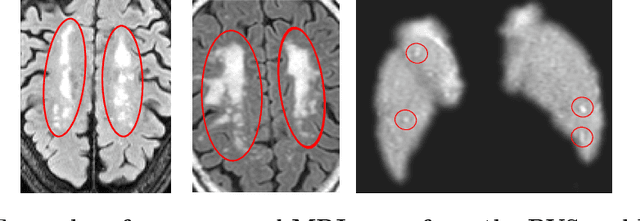
Abstract:We propose a novel convolutional neural network for lesion detection from weak labels. Only a single, global label per image - the lesion count - is needed for training. We train a regression network with a fully convolutional architecture combined with a global pooling layer to aggregate the 3D output into a scalar indicating the lesion count. When testing on unseen images, we first run the network to estimate the number of lesions. Then we remove the global pooling layer to compute localization maps of the size of the input image. We evaluate the proposed network on the detection of enlarged perivascular spaces in the basal ganglia in MRI. Our method achieves a sensitivity of 62% with on average 1.5 false positives per image. Compared with four other approaches based on intensity thresholding, saliency and class maps, our method has a 20% higher sensitivity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge