Ona Wu
Diffusion based Zero-shot Medical Image-to-Image Translation for Cross Modality Segmentation
Apr 09, 2024

Abstract:Cross-modality image segmentation aims to segment the target modalities using a method designed in the source modality. Deep generative models can translate the target modality images into the source modality, thus enabling cross-modality segmentation. However, a vast body of existing cross-modality image translation methods relies on supervised learning. In this work, we aim to address the challenge of zero-shot learning-based image translation tasks (extreme scenarios in the target modality is unseen in the training phase). To leverage generative learning for zero-shot cross-modality image segmentation, we propose a novel unsupervised image translation method. The framework learns to translate the unseen source image to the target modality for image segmentation by leveraging the inherent statistical consistency between different modalities for diffusion guidance. Our framework captures identical cross-modality features in the statistical domain, offering diffusion guidance without relying on direct mappings between the source and target domains. This advantage allows our method to adapt to changing source domains without the need for retraining, making it highly practical when sufficient labeled source domain data is not available. The proposed framework is validated in zero-shot cross-modality image segmentation tasks through empirical comparisons with influential generative models, including adversarial-based and diffusion-based models.
Zero-shot-Learning Cross-Modality Data Translation Through Mutual Information Guided Stochastic Diffusion
Jan 31, 2023
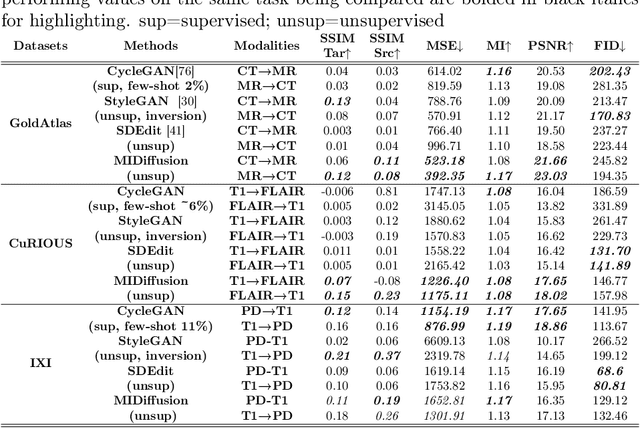
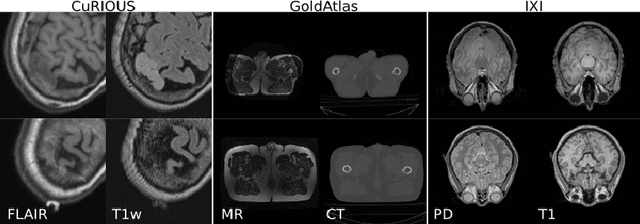
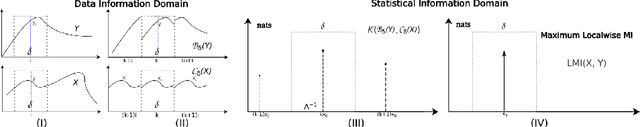
Abstract:Cross-modality data translation has attracted great interest in image computing. Deep generative models (\textit{e.g.}, GANs) show performance improvement in tackling those problems. Nevertheless, as a fundamental challenge in image translation, the problem of Zero-shot-Learning Cross-Modality Data Translation with fidelity remains unanswered. This paper proposes a new unsupervised zero-shot-learning method named Mutual Information guided Diffusion cross-modality data translation Model (MIDiffusion), which learns to translate the unseen source data to the target domain. The MIDiffusion leverages a score-matching-based generative model, which learns the prior knowledge in the target domain. We propose a differentiable local-wise-MI-Layer ($LMI$) for conditioning the iterative denoising sampling. The $LMI$ captures the identical cross-modality features in the statistical domain for the diffusion guidance; thus, our method does not require retraining when the source domain is changed, as it does not rely on any direct mapping between the source and target domains. This advantage is critical for applying cross-modality data translation methods in practice, as a reasonable amount of source domain dataset is not always available for supervised training. We empirically show the advanced performance of MIDiffusion in comparison with an influential group of generative models, including adversarial-based and other score-matching-based models.
Automated Image Registration Quality Assessment Utilizing Deep-learning based Ventricle Extraction in Clinical Data
Jul 01, 2019

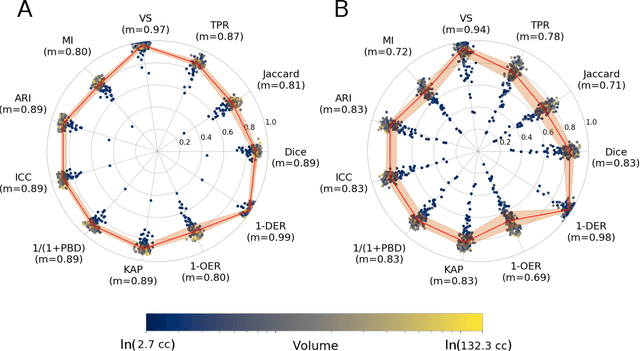
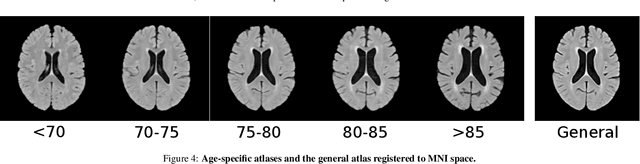
Abstract:Registration is a core component of many imaging pipelines. In case of clinical scans, with lower resolution and sometimes substantial motion artifacts, registration can produce poor results. Visual assessment of registration quality in large clinical datasets is inefficient. In this work, we propose to automatically assess the quality of registration to an atlas in clinical FLAIR MRI scans of the brain. The method consists of automatically segmenting the ventricles of a given scan using a neural network, and comparing the segmentation to the atlas' ventricles propagated to image space. We used the proposed method to improve clinical image registration to a general atlas by computing multiple registrations and then selecting the registration that yielded the highest ventricle overlap. Methods were evaluated in a single-site dataset of more than 1000 scans, as well as a multi-center dataset comprising 142 clinical scans from 12 sites. The automated ventricle segmentation reached a Dice coefficient with manual annotations of 0.89 in the single-site dataset, and 0.83 in the multi-center dataset. Registration via age-specific atlases could improve ventricle overlap compared to a direct registration to the general atlas (Dice similarity coefficient increase up to 0.15). Experiments also showed that selecting scans with the registration quality assessment method could improve the quality of average maps of white matter hyperintensity burden, instead of using all scans for the computation of the white matter hyperintensity map. In this work, we demonstrated the utility of an automated tool for assessing image registration quality in clinical scans. This image quality assessment step could ultimately assist in the translation of automated neuroimaging pipelines to the clinic.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge