Juan E. Arco
Tiled sparse coding in eigenspaces for the COVID-19 diagnosis in chest X-ray images
Jun 28, 2021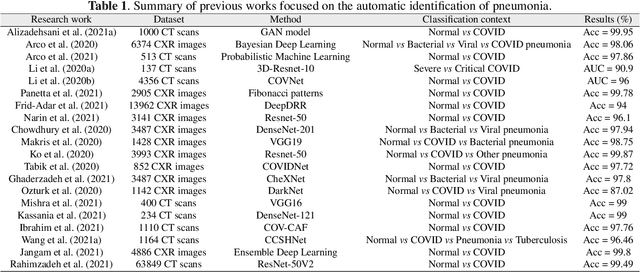
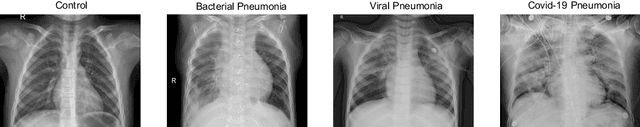
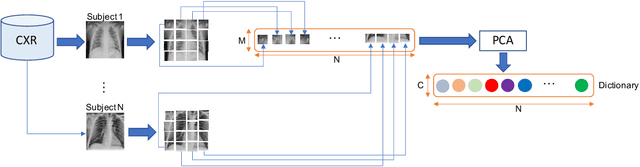
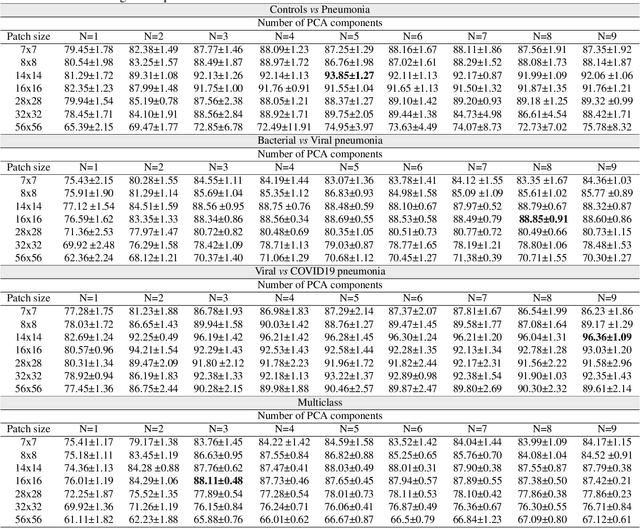
Abstract:The ongoing crisis of the COVID-19 (Coronavirus disease 2019) pandemic has changed the world. According to the World Health Organization (WHO), 4 million people have died due to this disease, whereas there have been more than 180 million confirmed cases of COVID-19. The collapse of the health system in many countries has demonstrated the need of developing tools to automatize the diagnosis of the disease from medical imaging. Previous studies have used deep learning for this purpose. However, the performance of this alternative highly depends on the size of the dataset employed for training the algorithm. In this work, we propose a classification framework based on sparse coding in order to identify the pneumonia patterns associated with different pathologies. Specifically, each chest X-ray (CXR) image is partitioned into different tiles. The most relevant features extracted from PCA are then used to build the dictionary within the sparse coding procedure. Once images are transformed and reconstructed from the elements of the dictionary, classification is performed from the reconstruction errors of individual patches associated with each image. Performance is evaluated in a real scenario where simultaneously differentiation between four different pathologies: control vs bacterial pneumonia vs viral pneumonia vs COVID-19. The accuracy when identifying the presence of pneumonia is 93.85%, whereas 88.11% is obtained in the 4-class classification context. The excellent results and the pioneering use of sparse coding in this scenario evidence the applicability of this approach as an aid for clinicians in a real-world environment.
Probabilistic combination of eigenlungs-based classifiers for COVID-19 diagnosis in chest CT images
Mar 04, 2021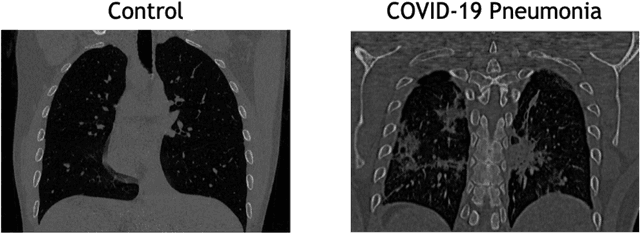
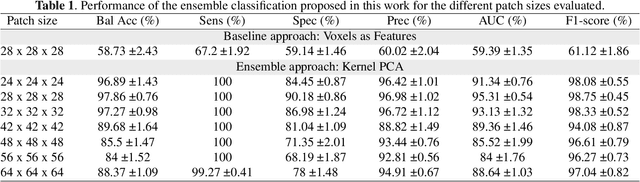
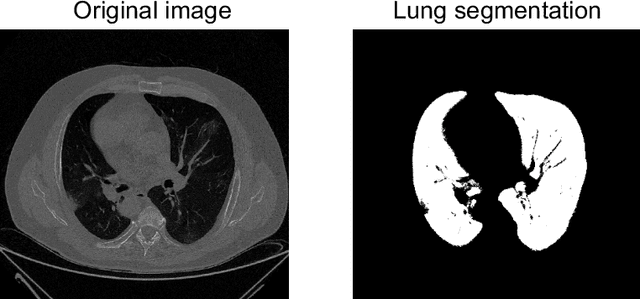
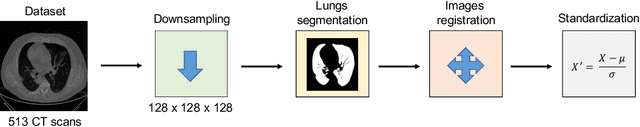
Abstract:The outbreak of the COVID-19 (Coronavirus disease 2019) pandemic has changed the world. According to the World Health Organization (WHO), there have been more than 100 million confirmed cases of COVID-19, including more than 2.4 million deaths. It is extremely important the early detection of the disease, and the use of medical imaging such as chest X-ray (CXR) and chest Computed Tomography (CCT) have proved to be an excellent solution. However, this process requires clinicians to do it within a manual and time-consuming task, which is not ideal when trying to speed up the diagnosis. In this work, we propose an ensemble classifier based on probabilistic Support Vector Machine (SVM) in order to identify pneumonia patterns while providing information about the reliability of the classification. Specifically, each CCT scan is divided into cubic patches and features contained in each one of them are extracted by applying kernel PCA. The use of base classifiers within an ensemble allows our system to identify the pneumonia patterns regardless of their size or location. Decisions of each individual patch are then combined into a global one according to the reliability of each individual classification: the lower the uncertainty, the higher the contribution. Performance is evaluated in a real scenario, yielding an accuracy of 97.86%. The large performance obtained and the simplicity of the system (use of deep learning in CCT images would result in a huge computational cost) evidence the applicability of our proposal in a real-world environment.
Uncertainty-Aware Semi-supervised Method using Large Unlabelled and Limited Labeled COVID-19 Data
Feb 12, 2021
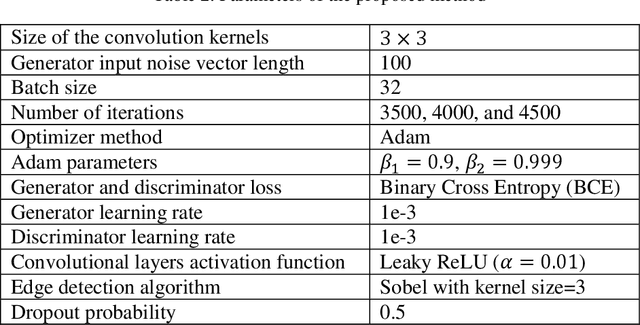
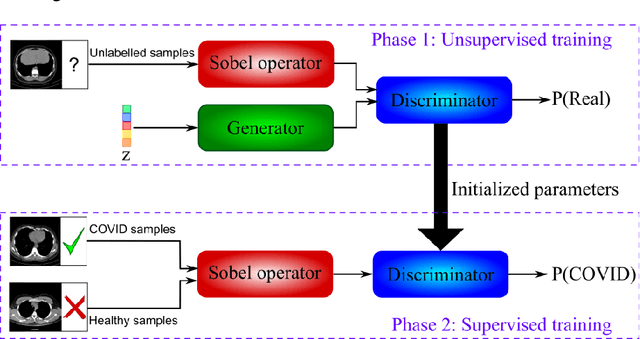

Abstract:The new coronavirus has caused more than 1 million deaths and continues to spread rapidly. This virus targets the lungs, causing respiratory distress which can be mild or severe. The X-ray or computed tomography (CT) images of lungs can reveal whether the patient is infected with COVID-19 or not. Many researchers are trying to improve COVID-19 detection using artificial intelligence. In this paper, relying on Generative Adversarial Networks (GAN), we propose a Semi-supervised Classification using Limited Labelled Data (SCLLD) for automated COVID-19 detection. Our motivation is to develop learning method which can cope with scenarios that preparing labelled data is time consuming or expensive. We further improved the detection accuracy of the proposed method by applying Sobel edge detection. The GAN discriminator output is a probability value which is used for classification in this work. The proposed system is trained using 10,000 CT scans collected from Omid hospital. Also, we validate our system using the public dataset. The proposed method is compared with other state of the art supervised methods such as Gaussian processes. To the best of our knowledge, this is the first time a COVID-19 semi-supervised detection method is presented. Our method is capable of learning from a mixture of limited labelled and unlabelled data where supervised learners fail due to lack of sufficient amount of labelled data. Our semi-supervised training method significantly outperforms the supervised training of Convolutional Neural Network (CNN) in case labelled training data is scarce. Our method has achieved an accuracy of 99.60%, sensitivity of 99.39%, and specificity of 99.80% where CNN (trained supervised) has achieved an accuracy of 69.87%, sensitivity of 94%, and specificity of 46.40%.
Uncertainty-driven ensembles of deep architectures for multiclass classification. Application to COVID-19 diagnosis in chest X-ray images
Nov 27, 2020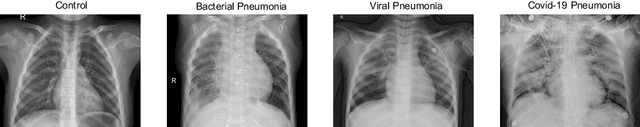
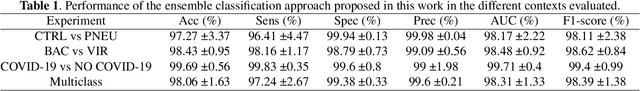
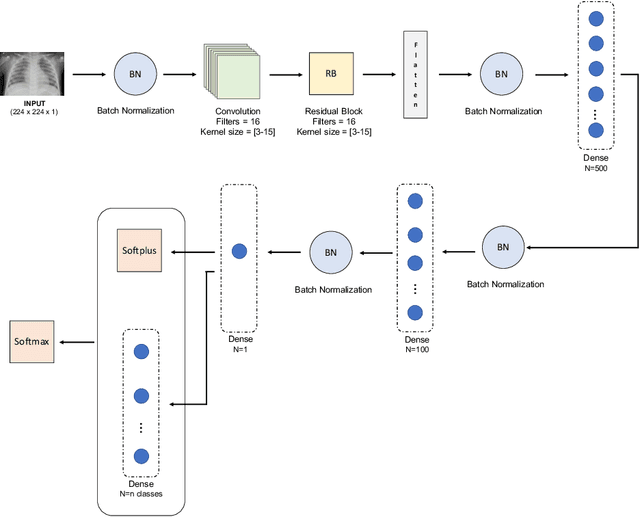
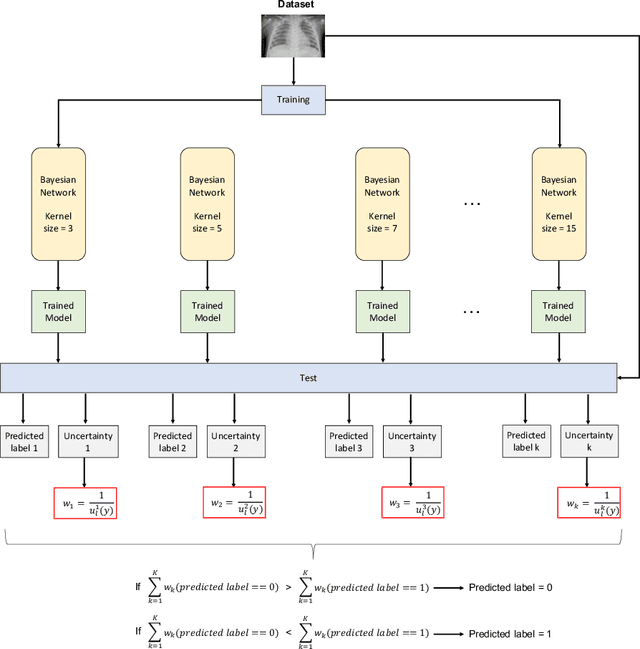
Abstract:Respiratory diseases kill million of people each year. Diagnosis of these pathologies is a manual, time-consuming process that has inter and intra-observer variability, delaying diagnosis and treatment. The recent COVID-19 pandemic has demonstrated the need of developing systems to automatize the diagnosis of pneumonia, whilst Convolutional Neural Network (CNNs) have proved to be an excellent option for the automatic classification of medical images. However, given the need of providing a confidence classification in this context it is crucial to quantify the reliability of the model's predictions. In this work, we propose a multi-level ensemble classification system based on a Bayesian Deep Learning approach in order to maximize performance while quantifying the uncertainty of each classification decision. This tool combines the information extracted from different architectures by weighting their results according to the uncertainty of their predictions. Performance of the Bayesian network is evaluated in a real scenario where simultaneously differentiating between four different pathologies: control vs bacterial pneumonia vs viral pneumonia vs COVID-19 pneumonia. A three-level decision tree is employed to divide the 4-class classification into three binary classifications, yielding an accuracy of 98.06% and overcoming the results obtained by recent literature. The reduced preprocessing needed for obtaining this high performance, in addition to the information provided about the reliability of the predictions evidence the applicability of the system to be used as an aid for clinicians.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge