Irina Grigorescu
School of Biomedical Engineering & Imaging Sciences, King's College London, London, United Kingdom
Automatic 8-tissue Segmentation for 6-month Infant Brains
Aug 27, 2024
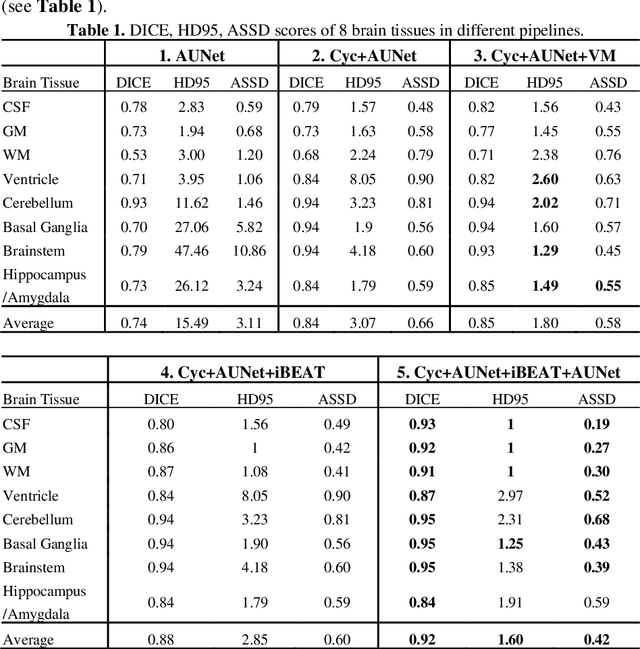
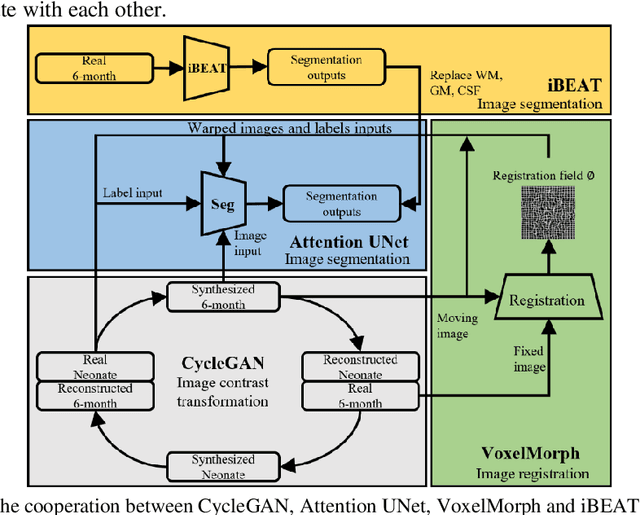
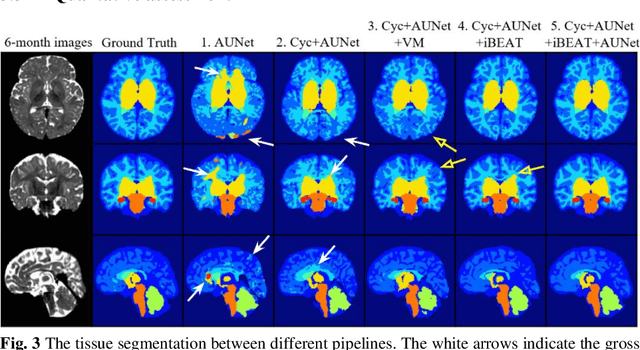
Abstract:Numerous studies have highlighted that atypical brain development, particularly during infancy and toddlerhood, is linked to an increased likelihood of being diagnosed with a neurodevelopmental condition, such as autism. Accurate brain tissue segmentations for morphological analysis are essential in numerous infant studies. However, due to ongoing white matter (WM) myelination changing tissue contrast in T1- and T2-weighted images, automatic tissue segmentation in 6-month infants is particularly difficult. On the other hand, manual labelling by experts is time-consuming and labor-intensive. In this study, we propose the first 8-tissue segmentation pipeline for six-month-old infant brains. This pipeline utilizes domain adaptation (DA) techniques to leverage our longitudinal data, including neonatal images segmented with the neonatal Developing Human Connectome Project structural pipeline. Our pipeline takes raw 6-month images as inputs and generates the 8-tissue segmentation as outputs, forming an end-to-end segmentation pipeline. The segmented tissues include WM, gray matter (GM), cerebrospinal fluid (CSF), ventricles, cerebellum, basal ganglia, brainstem, and hippocampus/amygdala. Cycle-Consistent Generative Adversarial Network (CycleGAN) and Attention U-Net were employed to achieve the image contrast transformation between neonatal and 6-month images and perform tissue segmentation on the synthesized 6-month images (neonatal images with 6-month intensity contrast), respectively. Moreover, we incorporated the segmentation outputs from Infant Brain Extraction and Analysis Toolbox (iBEAT) and another Attention U-Net to further enhance the performance and construct the end-to-end segmentation pipeline. Our evaluation with real 6-month images achieved a DICE score of 0.92, an HD95 of 1.6, and an ASSD of 0.42.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024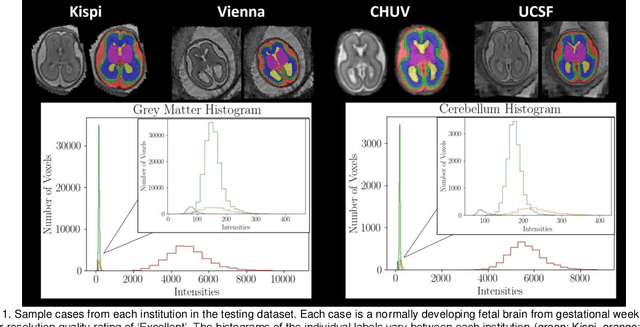
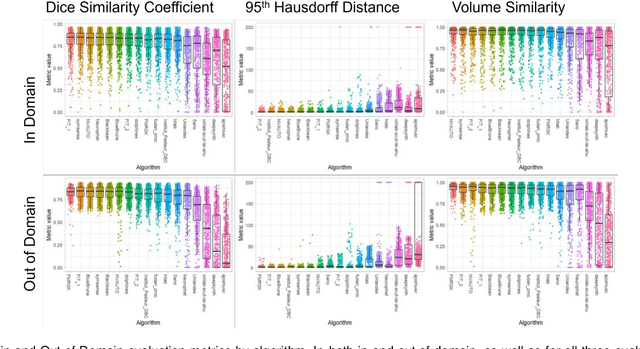
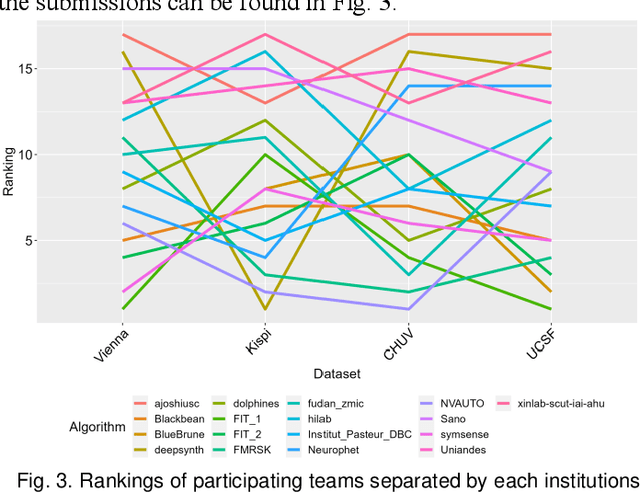
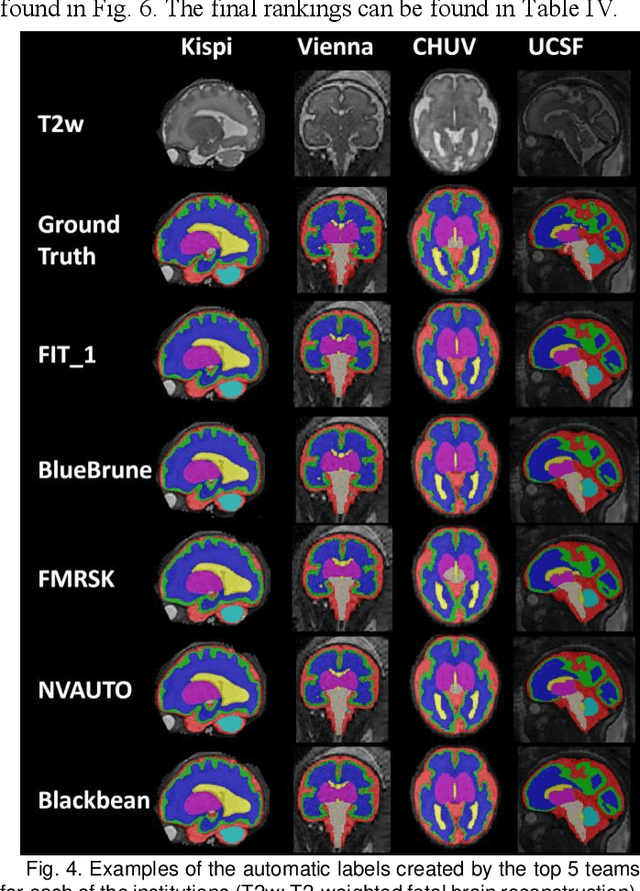
Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
Multi-task learning for joint weakly-supervised segmentation and aortic arch anomaly classification in fetal cardiac MRI
Nov 13, 2023



Abstract:Congenital Heart Disease (CHD) is a group of cardiac malformations present already during fetal life, representing the prevailing category of birth defects globally. Our aim in this study is to aid 3D fetal vessel topology visualisation in aortic arch anomalies, a group which encompasses a range of conditions with significant anatomical heterogeneity. We present a multi-task framework for automated multi-class fetal vessel segmentation from 3D black blood T2w MRI and anomaly classification. Our training data consists of binary manual segmentation masks of the cardiac vessels' region in individual subjects and fully-labelled anomaly-specific population atlases. Our framework combines deep learning label propagation using VoxelMorph with 3D Attention U-Net segmentation and DenseNet121 anomaly classification. We target 11 cardiac vessels and three distinct aortic arch anomalies, including double aortic arch, right aortic arch, and suspected coarctation of the aorta. We incorporate an anomaly classifier into our segmentation pipeline, delivering a multi-task framework with the primary motivation of correcting topological inaccuracies of the segmentation. The hypothesis is that the multi-task approach will encourage the segmenter network to learn anomaly-specific features. As a secondary motivation, an automated diagnosis tool may have the potential to enhance diagnostic confidence in a decision support setting. Our results showcase that our proposed training strategy significantly outperforms label propagation and a network trained exclusively on propagated labels. Our classifier outperforms a classifier trained exclusively on T2w volume images, with an average balanced accuracy of 0.99 (0.01) after joint training. Adding a classifier improves the anatomical and topological accuracy of all correctly classified double aortic arch subjects.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2023:015
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge