Guy Cloutier
Channel-Selective Normalization for Label-Shift Robust Test-Time Adaptation
Feb 07, 2024



Abstract:Deep neural networks have useful applications in many different tasks, however their performance can be severely affected by changes in the data distribution. For example, in the biomedical field, their performance can be affected by changes in the data (different machines, populations) between training and test datasets. To ensure robustness and generalization to real-world scenarios, test-time adaptation has been recently studied as an approach to adjust models to a new data distribution during inference. Test-time batch normalization is a simple and popular method that achieved compelling performance on domain shift benchmarks. It is implemented by recalculating batch normalization statistics on test batches. Prior work has focused on analysis with test data that has the same label distribution as the training data. However, in many practical applications this technique is vulnerable to label distribution shifts, sometimes producing catastrophic failure. This presents a risk in applying test time adaptation methods in deployment. We propose to tackle this challenge by only selectively adapting channels in a deep network, minimizing drastic adaptation that is sensitive to label shifts. Our selection scheme is based on two principles that we empirically motivate: (1) later layers of networks are more sensitive to label shift (2) individual features can be sensitive to specific classes. We apply the proposed technique to three classification tasks, including CIFAR10-C, Imagenet-C, and diagnosis of fatty liver, where we explore both covariate and label distribution shifts. We find that our method allows to bring the benefits of TTA while significantly reducing the risk of failure common in other methods, while being robust to choice in hyperparameters.
Homodyned K-Distribution Parameter Estimation in Quantitative Ultrasound: Autoencoder and Bayesian Neural Network Approaches
Jan 19, 2024



Abstract:Quantitative ultrasound (QUS) analyzes the ultrasound backscattered data to find the properties of scatterers that correlate with the tissue microstructure. Statistics of the envelope of the backscattered radiofrequency (RF) data can be utilized to estimate several QUS parameters. Different distributions have been proposed to model envelope data. The homodyned K-distribution (HK distribution) is one of the most comprehensive distributions that can model ultrasound backscattered envelope data under diverse scattering conditions (varying scatterer number density and coherent scattering). The scatterer clustering parameter (alpha) and the ratio of the coherent to diffuse scattering power (k) are the parameters of this distribution that have been used extensively for tissue characterization in diagnostic ultrasound. The estimation of these two parameters (which we refer to as HK parameters) is done using optimization algorithms in which statistical features such as the envelope point-wise signalto-noise ratio (SNR), skewness, kurtosis, and the log-based moments have been utilized as input to such algorithms. The optimization methods minimize the difference between features and their theoretical value from the HK model. We propose that the true value of these statistical features is a hyperplane that covers a small portion of the feature space. In this paper, we follow two approaches to reduce the effect of sample features' error. We propose a model projection neural network based on denoising autoencoders to project the noisy features into this space based on this assumption. We also investigate if the noise distribution can be learned by the deep estimators. We compare the proposed methods with conventional methods using simulations, an experimental phantom, and data from an in vivo animal model of hepatic steatosis. A demo code are available online at http://code.sonography.ai
Deep Learning in Ultrasound Elastography Imaging
Oct 31, 2020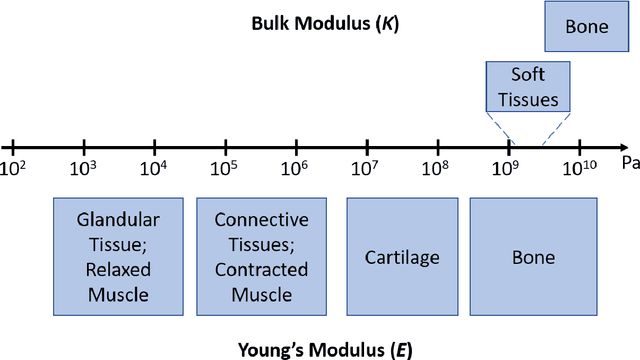

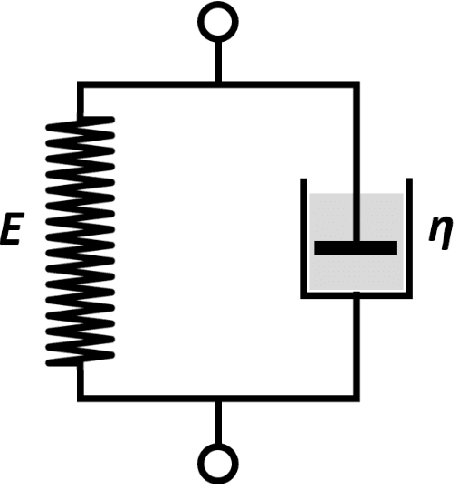
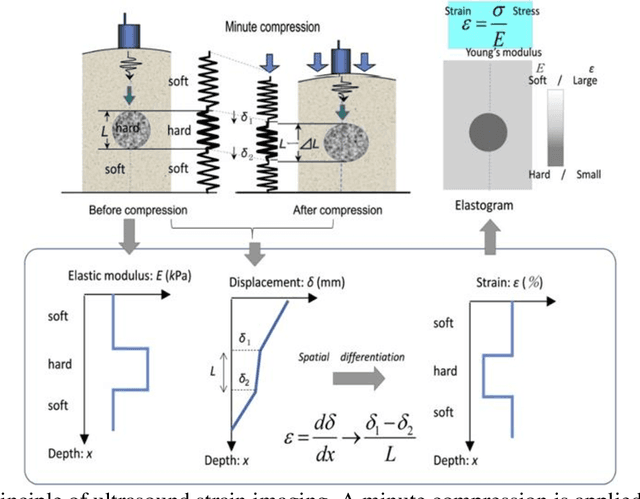
Abstract:It is known that changes in the mechanical properties of tissues are associated with the onset and progression of certain diseases. Ultrasound elastography is a technique to characterize tissue stiffness using ultrasound imaging either by measuring tissue strain using quasi-static elastography or natural organ pulsation elastography, or by tracing a propagated shear wave induced by a source or a natural vibration using dynamic elastography. In recent years, deep learning has begun to emerge in ultrasound elastography research. In this review, several common deep learning frameworks in the computer vision community, such as multilayer perceptron, convolutional neural network, and recurrent neural network are described. Then, recent advances in ultrasound elastography using such deep learning techniques are revisited in terms of algorithm development and clinical diagnosis. Finally, the current challenges and future developments of deep learning in ultrasound elastography are prospected.
Automatic Frame Selection using CNN in Ultrasound Elastography
Feb 17, 2020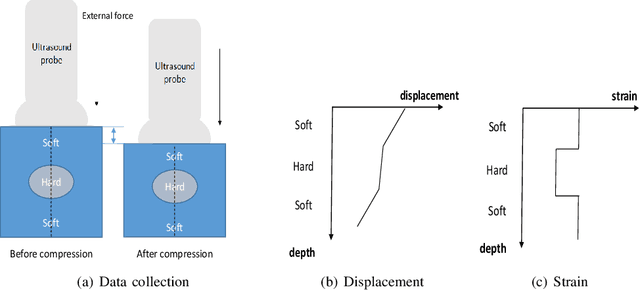
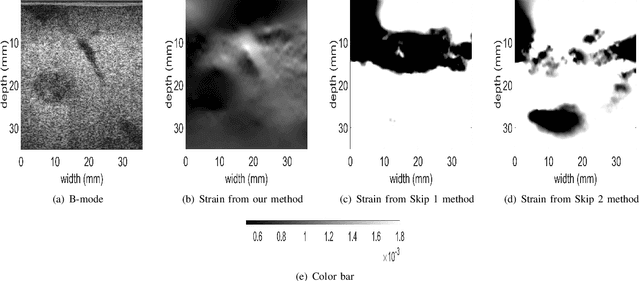
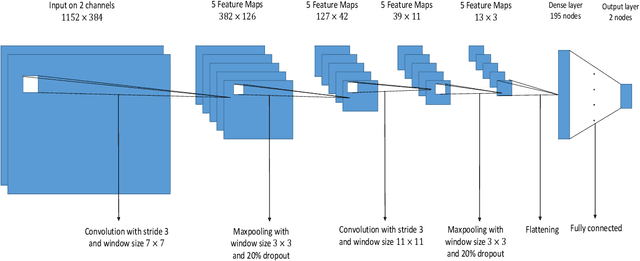

Abstract:Ultrasound elastography is used to estimate the mechanical properties of the tissue by monitoring its response to an internal or external force. Different levels of deformation are obtained from different tissue types depending on their mechanical properties, where stiffer tissues deform less. Given two radio frequency (RF) frames collected before and after some deformation, we estimate displacement and strain images by comparing the RF frames. The quality of the strain image is dependent on the type of motion that occurs during deformation. In-plane axial motion results in high-quality strain images, whereas out-of-plane motion results in low-quality strain images. In this paper, we introduce a new method using a convolutional neural network (CNN) to determine the suitability of a pair of RF frames for elastography in only 5.4 ms. Our method could also be used to automatically choose the best pair of RF frames, yielding a high-quality strain image. The CNN was trained on 3,818 pairs of RF frames, while testing was done on 986 new unseen pairs, achieving an accuracy of more than 91%. The RF frames were collected from both phantom and in vivo data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge