Eric Deutsch
Université Paris-Saclay, Institut Gustave Roussy, Inserm, Radiothérapie Moléculaire et Innovation Thérapeutique
GuidedRec: Guiding Ill-Posed Unsupervised Volumetric Recovery
May 20, 2024



Abstract:We introduce a novel unsupervised approach to reconstructing a 3D volume from only two planar projections that exploits a previous\-ly-captured 3D volume of the patient. Such volume is readily available in many important medical procedures and previous methods already used such a volume. Earlier methods that work by deforming this volume to match the projections typically fail when the number of projections is very low as the alignment becomes underconstrained. We show how to use a generative model of the volume structures to constrain the deformation and obtain a correct estimate. Moreover, our method is not bounded to a specific sensor calibration and can be applied to new calibrations without retraining. We evaluate our approach on a challenging dataset and show it outperforms state-of-the-art methods. As a result, our method could be used in treatment scenarios such as surgery and radiotherapy while drastically reducing patient radiation exposure.
Region-guided CycleGANs for Stain Transfer in Whole Slide Images
Aug 26, 2022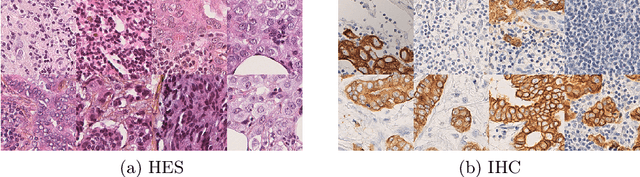
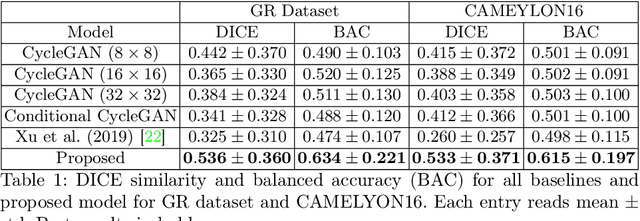
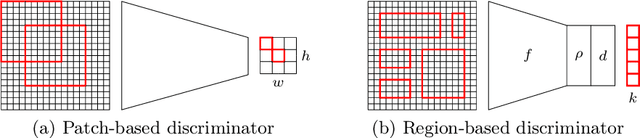
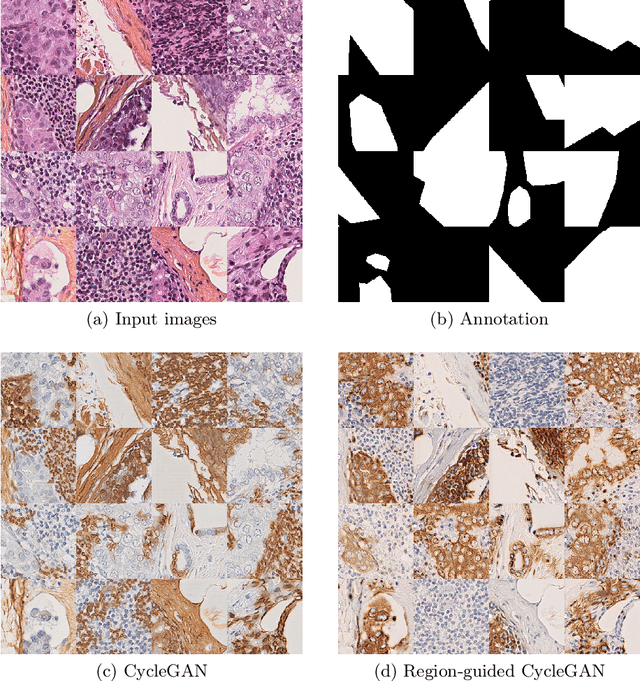
Abstract:In whole slide imaging, commonly used staining techniques based on hematoxylin and eosin (H&E) and immunohistochemistry (IHC) stains accentuate different aspects of the tissue landscape. In the case of detecting metastases, IHC provides a distinct readout that is readily interpretable by pathologists. IHC, however, is a more expensive approach and not available at all medical centers. Virtually generating IHC images from H&E using deep neural networks thus becomes an attractive alternative. Deep generative models such as CycleGANs learn a semantically-consistent mapping between two image domains, while emulating the textural properties of each domain. They are therefore a suitable choice for stain transfer applications. However, they remain fully unsupervised, and possess no mechanism for enforcing biological consistency in stain transfer. In this paper, we propose an extension to CycleGANs in the form of a region of interest discriminator. This allows the CycleGAN to learn from unpaired datasets where, in addition, there is a partial annotation of objects for which one wishes to enforce consistency. We present a use case on whole slide images, where an IHC stain provides an experimentally generated signal for metastatic cells. We demonstrate the superiority of our approach over prior art in stain transfer on histopathology tiles over two datasets. Our code and model are available at https://github.com/jcboyd/miccai2022-roigan.
MICS : Multi-steps, Inverse Consistency and Symmetric deep learning registration network
Nov 23, 2021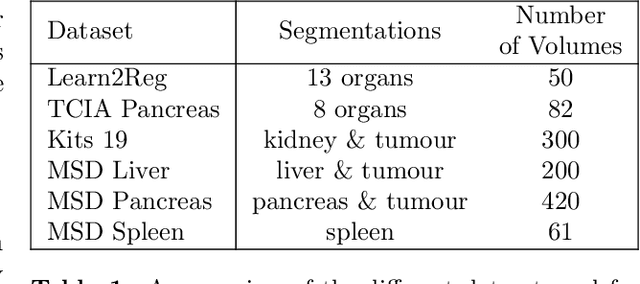
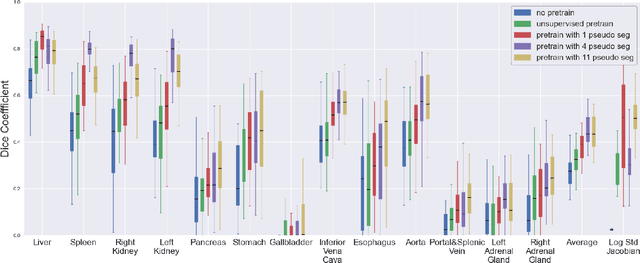
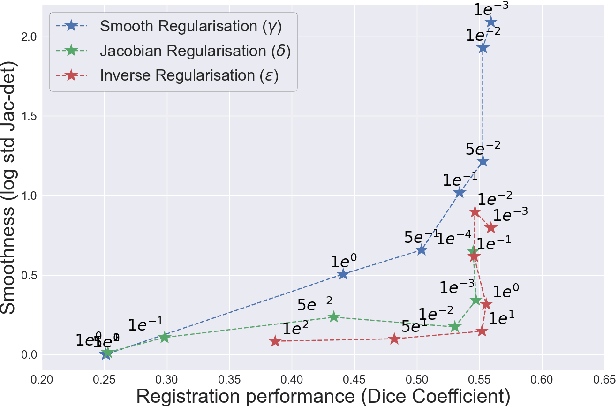

Abstract:Deformable registration consists of finding the best dense correspondence between two different images. Many algorithms have been published, but the clinical application was made difficult by the high calculation time needed to solve the optimisation problem. Deep learning overtook this limitation by taking advantage of GPU calculation and the learning process. However, many deep learning methods do not take into account desirable properties respected by classical algorithms. In this paper, we present MICS, a novel deep learning algorithm for medical imaging registration. As registration is an ill-posed problem, we focused our algorithm on the respect of different properties: inverse consistency, symmetry and orientation conservation. We also combined our algorithm with a multi-step strategy to refine and improve the deformation grid. While many approaches applied registration to brain MRI, we explored a more challenging body localisation: abdominal CT. Finally, we evaluated our method on a dataset used during the Learn2Reg challenge, allowing a fair comparison with published methods.
Self-Supervised Representation Learning using Visual Field Expansion on Digital Pathology
Sep 07, 2021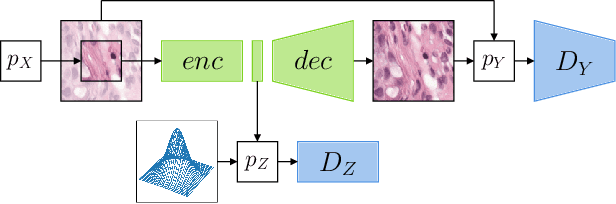
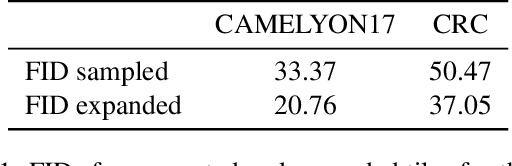
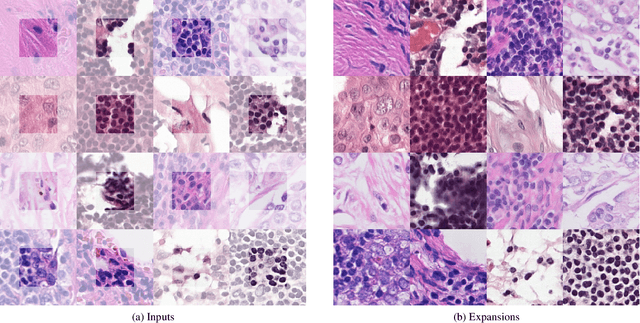
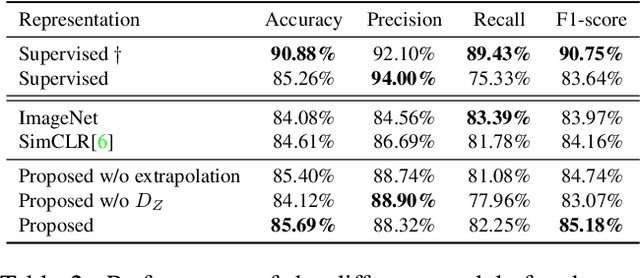
Abstract:The examination of histopathology images is considered to be the gold standard for the diagnosis and stratification of cancer patients. A key challenge in the analysis of such images is their size, which can run into the gigapixels and can require tedious screening by clinicians. With the recent advances in computational medicine, automatic tools have been proposed to assist clinicians in their everyday practice. Such tools typically process these large images by slicing them into tiles that can then be encoded and utilized for different clinical models. In this study, we propose a novel generative framework that can learn powerful representations for such tiles by learning to plausibly expand their visual field. In particular, we developed a progressively grown generative model with the objective of visual field expansion. Thus trained, our model learns to generate different tissue types with fine details, while simultaneously learning powerful representations that can be used for different clinical endpoints, all in a self-supervised way. To evaluate the performance of our model, we conducted classification experiments on CAMELYON17 and CRC benchmark datasets, comparing favorably to other self-supervised and pre-trained strategies that are commonly used in digital pathology. Our code is available at https://github.com/jcboyd/cdpath21-gan.
Exploring Deep Registration Latent Spaces
Jul 23, 2021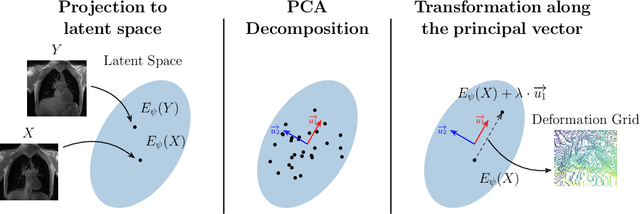
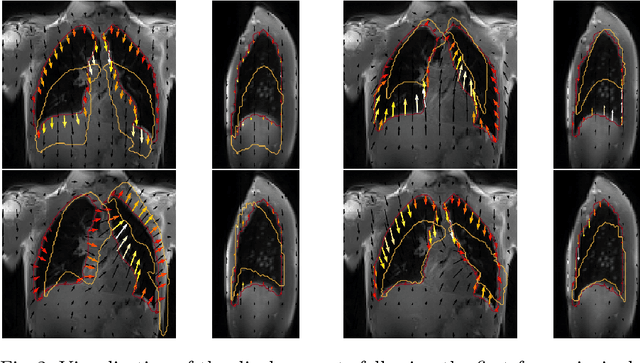
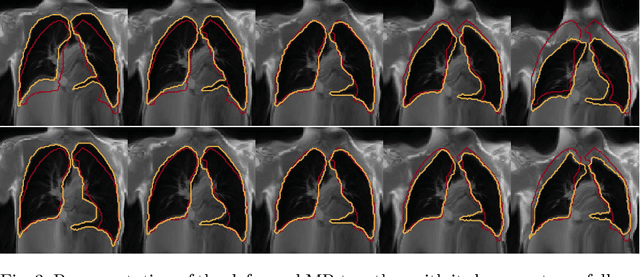
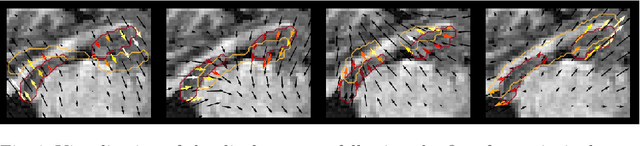
Abstract:Explainability of deep neural networks is one of the most challenging and interesting problems in the field. In this study, we investigate the topic focusing on the interpretability of deep learning-based registration methods. In particular, with the appropriate model architecture and using a simple linear projection, we decompose the encoding space, generating a new basis, and we empirically show that this basis captures various decomposed anatomically aware geometrical transformations. We perform experiments using two different datasets focusing on lungs and hippocampus MRI. We show that such an approach can decompose the highly convoluted latent spaces of registration pipelines in an orthogonal space with several interesting properties. We hope that this work could shed some light on a better understanding of deep learning-based registration methods.
Weakly supervised pan-cancer segmentation tool
May 10, 2021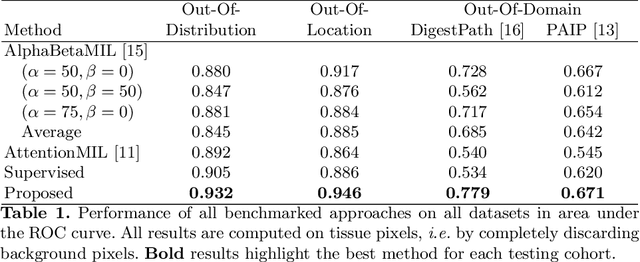
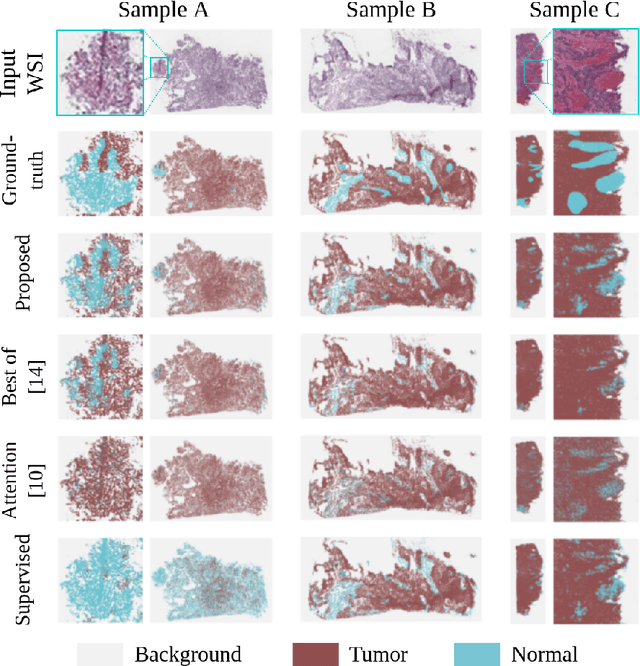
Abstract:The vast majority of semantic segmentation approaches rely on pixel-level annotations that are tedious and time consuming to obtain and suffer from significant inter and intra-expert variability. To address these issues, recent approaches have leveraged categorical annotations at the slide-level, that in general suffer from robustness and generalization. In this paper, we propose a novel weakly supervised multi-instance learning approach that deciphers quantitative slide-level annotations which are fast to obtain and regularly present in clinical routine. The extreme potentials of the proposed approach are demonstrated for tumor segmentation of solid cancer subtypes. The proposed approach achieves superior performance in out-of-distribution, out-of-location, and out-of-domain testing sets.
Sparse convolutional context-aware multiple instance learning for whole slide image classification
May 06, 2021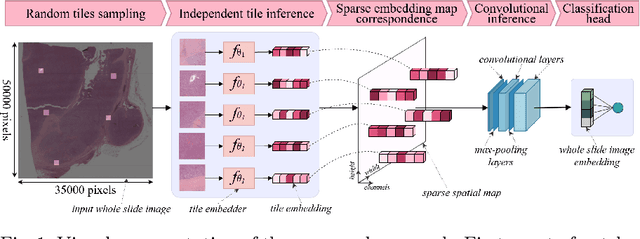
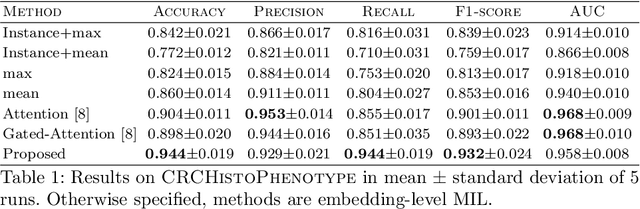
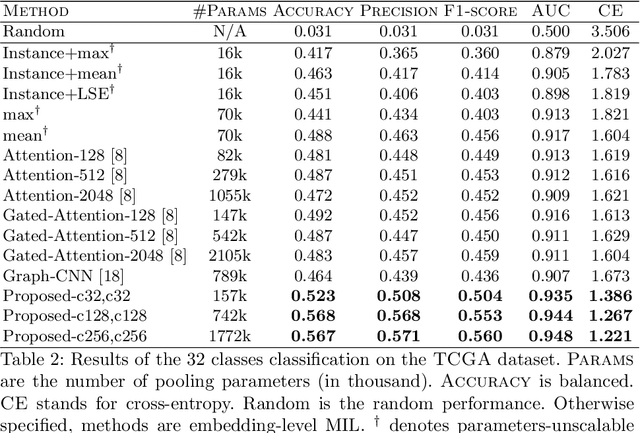
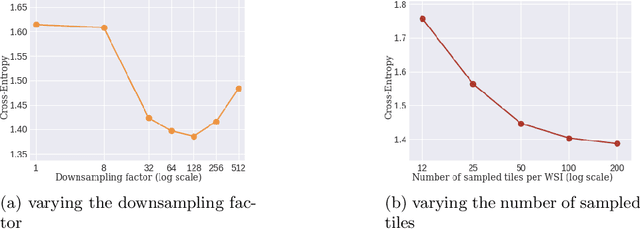
Abstract:Whole slide microscopic slides display many cues about the underlying tissue guiding diagnostic and the choice of therapy for many diseases. However, their enormous size often in gigapixels hampers the use of traditional neural network architectures. To tackle this issue, multiple instance learning (MIL) classifies bags of patches instead of whole slide images. Most MIL strategies consider that patches are independent and identically distributed. Our approach presents a paradigm shift through the integration of spatial information of patches with a sparse-input convolutional-based MIL strategy. The formulated framework is generic, flexible, scalable and is the first to introduce contextual dependencies between decisions taken at the patch level. It achieved state-of-the-art performance in pan-cancer subtype classification. The code of this work will be made available.
Cancer Gene Profiling through Unsupervised Discovery
Feb 11, 2021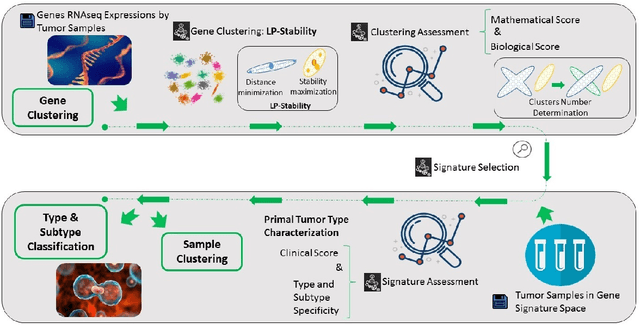
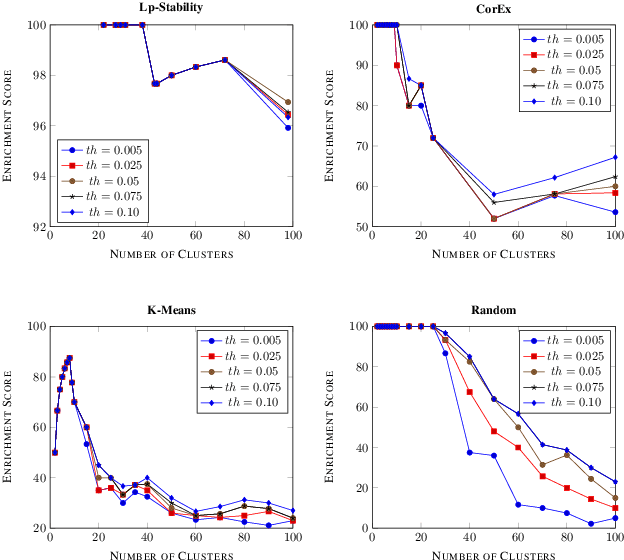
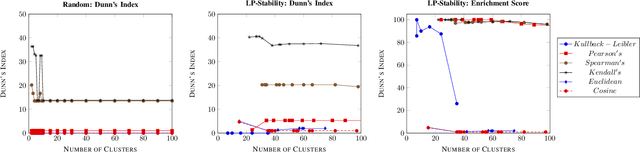
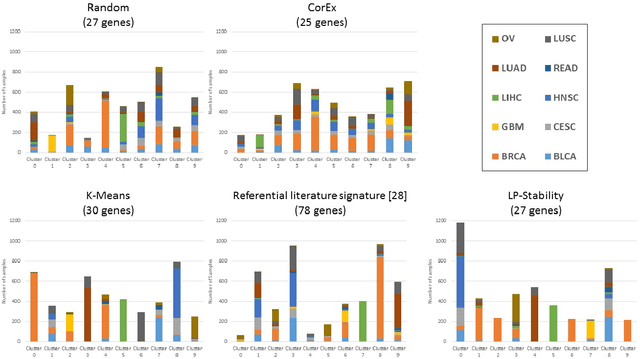
Abstract:Precision medicine is a paradigm shift in healthcare relying heavily on genomics data. However, the complexity of biological interactions, the large number of genes as well as the lack of comparisons on the analysis of data, remain a tremendous bottleneck regarding clinical adoption. In this paper, we introduce a novel, automatic and unsupervised framework to discover low-dimensional gene biomarkers. Our method is based on the LP-Stability algorithm, a high dimensional center-based unsupervised clustering algorithm, that offers modularity as concerns metric functions and scalability, while being able to automatically determine the best number of clusters. Our evaluation includes both mathematical and biological criteria. The recovered signature is applied to a variety of biological tasks, including screening of biological pathways and functions, and characterization relevance on tumor types and subtypes. Quantitative comparisons among different distance metrics, commonly used clustering methods and a referential gene signature used in the literature, confirm state of the art performance of our approach. In particular, our signature, that is based on 27 genes, reports at least $30$ times better mathematical significance (average Dunn's Index) and 25% better biological significance (average Enrichment in Protein-Protein Interaction) than those produced by other referential clustering methods. Finally, our signature reports promising results on distinguishing immune inflammatory and immune desert tumors, while reporting a high balanced accuracy of 92% on tumor types classification and averaged balanced accuracy of 68% on tumor subtypes classification, which represents, respectively 7% and 9% higher performance compared to the referential signature.
Top 10 BraTS 2020 challenge solution: Brain tumor segmentation with self-ensembled, deeply-supervised 3D-Unet like neural networks
Oct 30, 2020

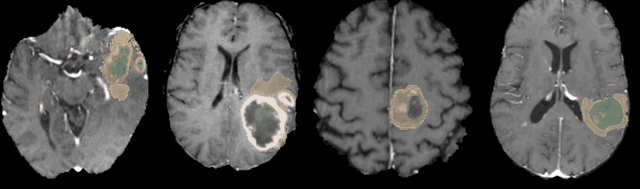

Abstract:Brain tumor segmentation is a critical task for patient's disease management. To this end, we trained multiple U-net like neural networks, mainly with deep supervision and stochastic weight averaging, on the Multimodal Brain Tumor Segmentation Challenge (BraTS) 2020 training dataset, in a cross-validated fashion. Final brain tumor segmentations were produced by first averaging independently two sets of models, and then custom merging the labelmaps to account for individual performance of each set. Our performance on the online validation dataset with test time augmentation were as follows: Dice of 0.81, 0.91 and 0.85; Hausdorff (95%) of 20.6, 4,3, 5.7 mm for the enhancing tumor, whole tumor and tumor core, respectively. Similarly, our ensemble achieved a Dice of 0.79, 0.89 and 0.84, as well as Hausdorff (95%) of 20.4, 6.7 and 19.5mm on the final test dataset. More complicated training schemes and neural network architectures were investigated, without significant performance gain, at the cost of greatly increased training time. While relatively straightforward, our approach yielded good and balanced performance for each tumor subregions. Our solution is open sourced at https://github.com/lescientifik/xxxxx.
Deep learning based registration using spatial gradients and noisy segmentation labels
Oct 21, 2020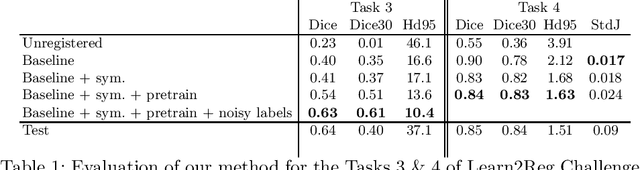
Abstract:Image registration is one of the most challenging problems in medical image analysis. In the recent years, deep learning based approaches became quite popular, providing fast and performing registration strategies. In this short paper, we summarise our work presented on Learn2Reg challenge 2020. The main contributions of our work rely on (i) a symmetric formulation, predicting the transformations from source to target and from target to source simultaneously, enforcing the trained representations to be similar and (ii) integration of variety of publicly available datasets used both for pretraining and for augmenting segmentation labels. Our method reports a mean dice of $0.64$ for task 3 and $0.85$ for task 4 on the test sets, taking third place on the challenge. Our code and models are publicly available at https://github.com/TheoEst/abdominal_registration and \https://github.com/TheoEst/hippocampus_registration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge