Alexandre Carré
Université Paris-Saclay, Institut Gustave Roussy, Inserm, Radiothérapie Moléculaire et Innovation Thérapeutique
Cancer Gene Profiling through Unsupervised Discovery
Feb 11, 2021
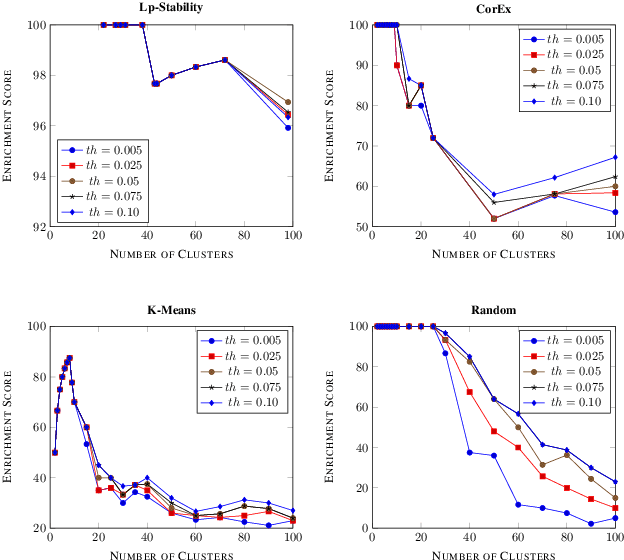
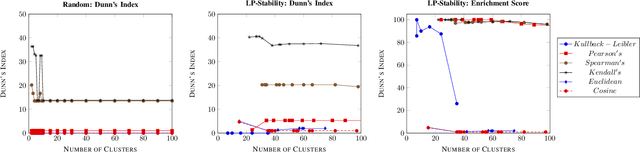
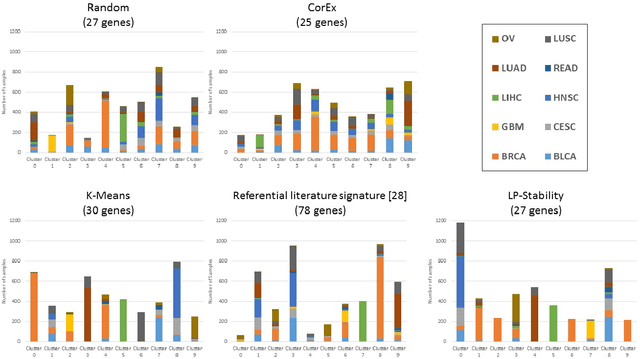
Abstract:Precision medicine is a paradigm shift in healthcare relying heavily on genomics data. However, the complexity of biological interactions, the large number of genes as well as the lack of comparisons on the analysis of data, remain a tremendous bottleneck regarding clinical adoption. In this paper, we introduce a novel, automatic and unsupervised framework to discover low-dimensional gene biomarkers. Our method is based on the LP-Stability algorithm, a high dimensional center-based unsupervised clustering algorithm, that offers modularity as concerns metric functions and scalability, while being able to automatically determine the best number of clusters. Our evaluation includes both mathematical and biological criteria. The recovered signature is applied to a variety of biological tasks, including screening of biological pathways and functions, and characterization relevance on tumor types and subtypes. Quantitative comparisons among different distance metrics, commonly used clustering methods and a referential gene signature used in the literature, confirm state of the art performance of our approach. In particular, our signature, that is based on 27 genes, reports at least $30$ times better mathematical significance (average Dunn's Index) and 25% better biological significance (average Enrichment in Protein-Protein Interaction) than those produced by other referential clustering methods. Finally, our signature reports promising results on distinguishing immune inflammatory and immune desert tumors, while reporting a high balanced accuracy of 92% on tumor types classification and averaged balanced accuracy of 68% on tumor subtypes classification, which represents, respectively 7% and 9% higher performance compared to the referential signature.
Deep learning based registration using spatial gradients and noisy segmentation labels
Oct 21, 2020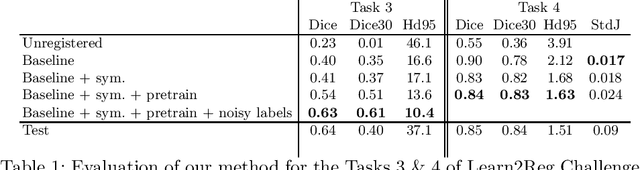
Abstract:Image registration is one of the most challenging problems in medical image analysis. In the recent years, deep learning based approaches became quite popular, providing fast and performing registration strategies. In this short paper, we summarise our work presented on Learn2Reg challenge 2020. The main contributions of our work rely on (i) a symmetric formulation, predicting the transformations from source to target and from target to source simultaneously, enforcing the trained representations to be similar and (ii) integration of variety of publicly available datasets used both for pretraining and for augmenting segmentation labels. Our method reports a mean dice of $0.64$ for task 3 and $0.85$ for task 4 on the test sets, taking third place on the challenge. Our code and models are publicly available at https://github.com/TheoEst/abdominal_registration and \https://github.com/TheoEst/hippocampus_registration.
Weakly supervised multiple instance learning histopathological tumor segmentation
Apr 21, 2020
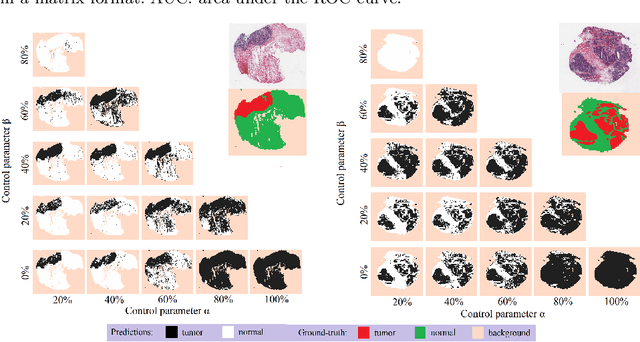
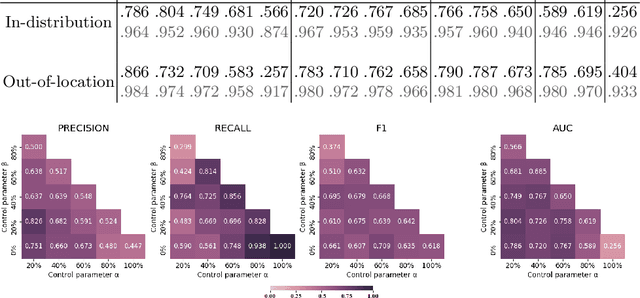

Abstract:Histopathological image segmentation is a challenging and important topic in medical imaging with tremendous potential impact in clinical practice. State of the art methods relying on hand-crafted annotations that reduce the scope of the solutions since digital histology suffers from standardization and samples differ significantly between cancer phenotypes. To this end, in this paper, we propose a weakly supervised framework relying on weak standard clinical practice annotations, available in most medical centers. In particular, we exploit a multiple instance learning scheme providing a label for each instance, establishing a detailed segmentation of whole slide images. The potential of the framework is assessed with multi-centric data experiments using The Cancer Genome Atlas repository and the publicly available PatchCamelyon dataset. Promising results when compared with experts' annotations demonstrate the potentials of our approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge