B. Milan Horacek
Fast Posterior Estimation of Cardiac Electrophysiological Model Parameters via Bayesian Active Learning
Oct 13, 2021



Abstract:Probabilistic estimation of cardiac electrophysiological model parameters serves an important step towards model personalization and uncertain quantification. The expensive computation associated with these model simulations, however, makes direct Markov Chain Monte Carlo (MCMC) sampling of the posterior probability density function (pdf) of model parameters computationally intensive. Approximated posterior pdfs resulting from replacing the simulation model with a computationally efficient surrogate, on the other hand, have seen limited accuracy. In this paper, we present a Bayesian active learning method to directly approximate the posterior pdf function of cardiac model parameters, in which we intelligently select training points to query the simulation model in order to learn the posterior pdf using a small number of samples. We integrate a generative model into Bayesian active learning to allow approximating posterior pdf of high-dimensional model parameters at the resolution of the cardiac mesh. We further introduce new acquisition functions to focus the selection of training points on better approximating the shape rather than the modes of the posterior pdf of interest. We evaluated the presented method in estimating tissue excitability in a 3D cardiac electrophysiological model in a range of synthetic and real-data experiments. We demonstrated its improved accuracy in approximating the posterior pdf compared to Bayesian active learning using regular acquisition functions, and substantially reduced computational cost in comparison to existing standard or accelerated MCMC sampling.
Improving Disentangled Representation Learning with the Beta Bernoulli Process
Sep 03, 2019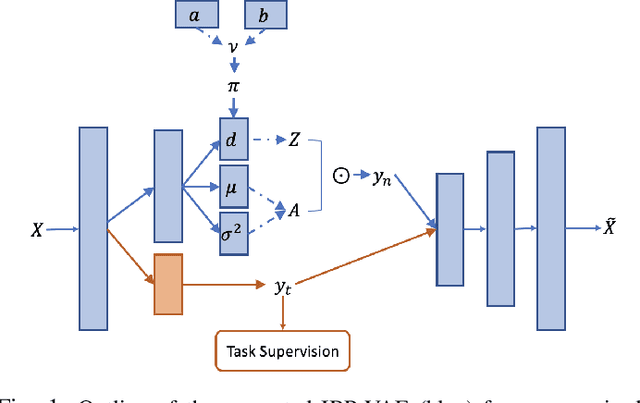
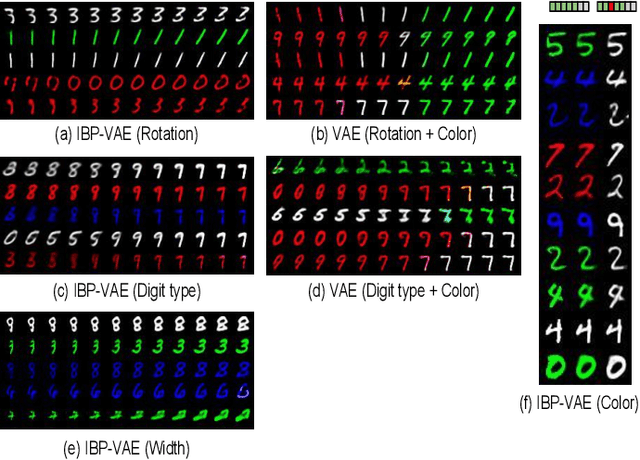
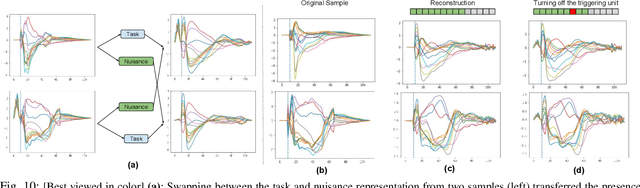


Abstract:To improve the ability of VAE to disentangle in the latent space, existing works mostly focus on enforcing independence among the learned latent factors. However, the ability of these models to disentangle often decreases as the complexity of the generative factors increases. In this paper, we investigate the little-explored effect of the modeling capacity of a posterior density on the disentangling ability of the VAE. We note that the independence within and the complexity of the latent density are two different properties we constrain when regularizing the posterior density: while the former promotes the disentangling ability of VAE, the latter -- if overly limited -- creates an unnecessary competition with the data reconstruction objective in VAE. Therefore, if we preserve the independence but allow richer modeling capacity in the posterior density, we will lift this competition and thereby allow improved independence and data reconstruction at the same time. We investigate this theoretical intuition with a VAE that utilizes a non-parametric latent factor model, the Indian Buffet Process (IBP), as a latent density that is able to grow with the complexity of the data. Across three widely-used benchmark data sets and two clinical data sets little explored for disentangled learning, we qualitatively and quantitatively demonstrated the improved disentangling performance of IBP-VAE over the state of the art. In the latter two clinical data sets riddled with complex factors of variations, we further demonstrated that unsupervised disentangling of nuisance factors via IBP-VAE -- when combined with a supervised objective -- can not only improve task accuracy in comparison to relevant supervised deep architectures but also facilitate knowledge discovery related to task decision-making. A shorter version of this work will appear in the ICDM 2019 conference proceedings.
Bayesian Optimization on Large Graphs via a Graph Convolutional Generative Model: Application in Cardiac Model Personalization
Jul 01, 2019
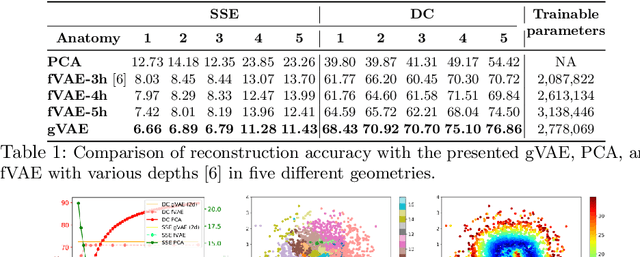

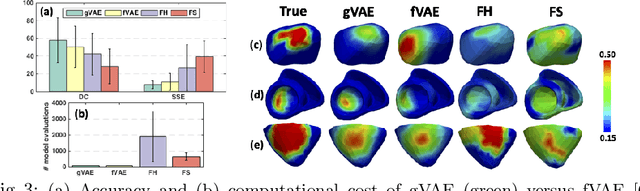
Abstract:Personalization of cardiac models involves the optimization of organ tissue properties that vary spatially over the non-Euclidean geometry model of the heart. To represent the high-dimensional (HD) unknown of tissue properties, most existing works rely on a low-dimensional (LD) partitioning of the geometrical model. While this exploits the geometry of the heart, it is of limited expressiveness to allow partitioning that is small enough for effective optimization. Recently, a variational auto-encoder (VAE) was utilized as a more expressive generative model to embed the HD optimization into the LD latent space. Its Euclidean nature, however, neglects the rich geometrical information in the heart. In this paper, we present a novel graph convolutional VAE to allow generative modeling of non-Euclidean data, and utilize it to embed Bayesian optimization of large graphs into a small latent space. This approach bridges the gap of previous works by introducing an expressive generative model that is able to incorporate the knowledge of spatial proximity and hierarchical compositionality of the underlying geometry. It further allows transferring of the learned features across different geometries, which was not possible with a regular VAE. We demonstrate these benefits of the presented method in synthetic and real data experiments of estimating tissue excitability in a cardiac electrophysiological model.
Generative Modeling and Inverse Imaging of Cardiac Transmembrane Potential
May 12, 2019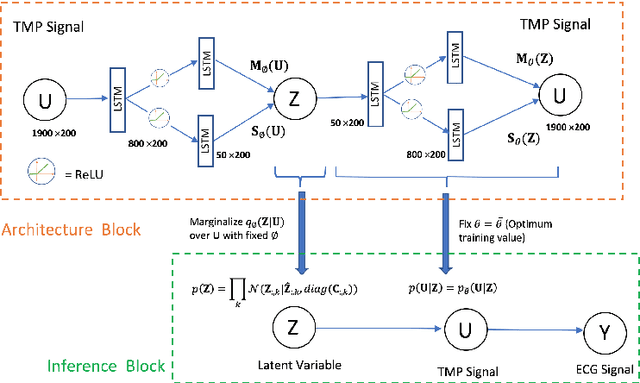

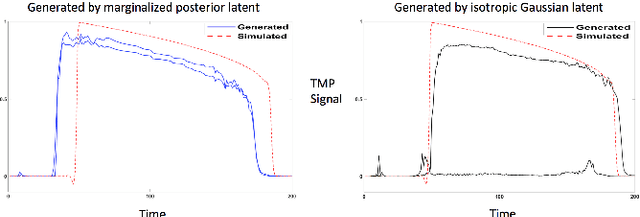
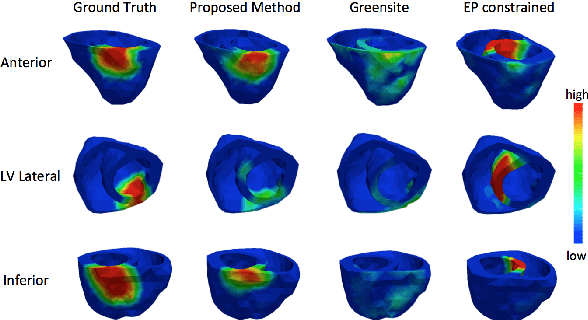
Abstract:Noninvasive reconstruction of cardiac transmembrane potential (TMP) from surface electrocardiograms (ECG) involves an ill-posed inverse problem. Model-constrained regularization is powerful for incorporating rich physiological knowledge about spatiotemporal TMP dynamics. These models are controlled by high-dimensional physical parameters which, if fixed, can introduce model errors and reduce the accuracy of TMP reconstruction. Simultaneous adaptation of these parameters during TMP reconstruction, however, is difficult due to their high dimensionality. We introduce a novel model-constrained inference framework that replaces conventional physiological models with a deep generative model trained to generate TMP sequences from low-dimensional generative factors. Using a variational auto-encoder (VAE) with long short-term memory (LSTM) networks, we train the VAE decoder to learn the conditional likelihood of TMP, while the encoder learns the prior distribution of generative factors. These two components allow us to develop an efficient algorithm to simultaneously infer the generative factors and TMP signals from ECG data. Synthetic and real-data experiments demonstrate that the presented method significantly improve the accuracy of TMP reconstruction compared with methods constrained by conventional physiological models or without physiological constraints.
Deep Generative Model with Beta Bernoulli Process for Modeling and Learning Confounding Factors
Oct 31, 2018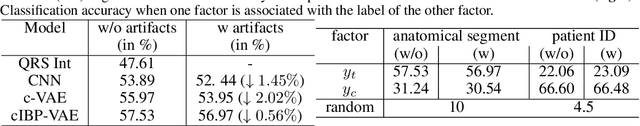
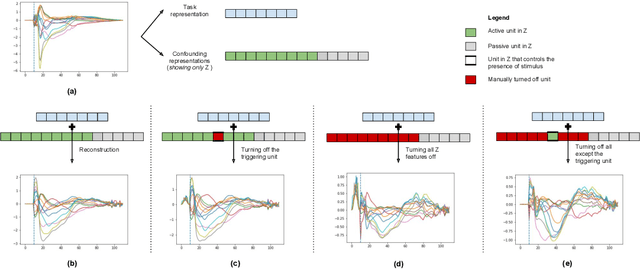
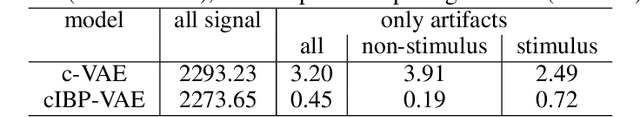

Abstract:While deep representation learning has become increasingly capable of separating task-relevant representations from other confounding factors in the data, two significant challenges remain. First, there is often an unknown and potentially infinite number of confounding factors coinciding in the data. Second, not all of these factors are readily observable. In this paper, we present a deep conditional generative model that learns to disentangle a task-relevant representation from an unknown number of confounding factors that may grow infinitely. This is achieved by marrying the representational power of deep generative models with Bayesian non-parametric factor models, where a supervised deterministic encoder learns task-related representation and a probabilistic encoder with an Indian Buffet Process (IBP) learns the unknown number of unobservable confounding factors. We tested the presented model in two datasets: a handwritten digit dataset (MNIST) augmented with colored digits and a clinical ECG dataset with significant inter-subject variations and augmented with signal artifacts. These diverse data sets highlighted the ability of the presented model to grow with the complexity of the data and identify the absence or presence of unobserved confounding factors.
Learning disentangled representation from 12-lead electrograms: application in localizing the origin of Ventricular Tachycardia
Aug 04, 2018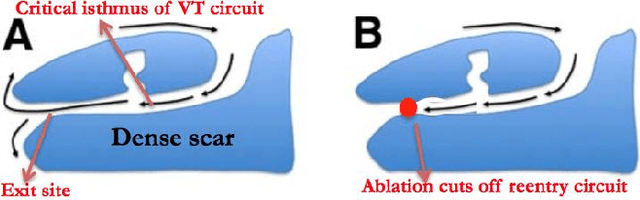
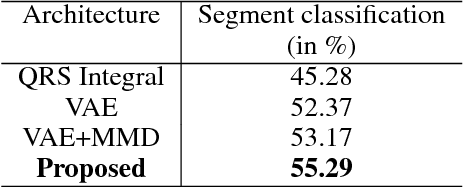
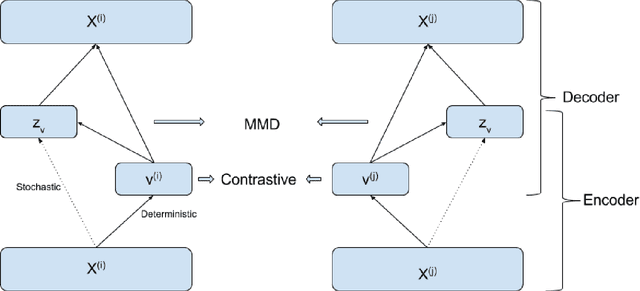
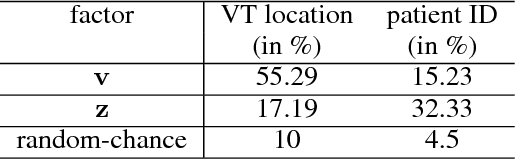
Abstract:The increasing availability of electrocardiogram (ECG) data has motivated the use of data-driven models for automating various clinical tasks based on ECG data. The development of subject-specific models are limited by the cost and difficulty of obtaining sufficient training data for each individual. The alternative of population model, however, faces challenges caused by the significant inter-subject variations within the ECG data. We address this challenge by investigating for the first time the problem of learning representations for clinically-informative variables while disentangling other factors of variations within the ECG data. In this work, we present a conditional variational autoencoder (VAE) to extract the subject-specific adjustment to the ECG data, conditioned on task-specific representations learned from a deterministic encoder. To encourage the representation for inter-subject variations to be independent from the task-specific representation, maximum mean discrepancy is used to match all the moments between the distributions learned by the VAE conditioning on the code from the deterministic encoder. The learning of the task-specific representation is regularized by a weak supervision in the form of contrastive regularization. We apply the proposed method to a novel yet important clinical task of classifying the origin of ventricular tachycardia (VT) into pre-defined segments, demonstrating the efficacy of the proposed method against the standard VAE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge