Zhihao Zhan
DC-VLAQ: Query-Residual Aggregation for Robust Visual Place Recognition
Jan 19, 2026Abstract:One of the central challenges in visual place recognition (VPR) is learning a robust global representation that remains discriminative under large viewpoint changes, illumination variations, and severe domain shifts. While visual foundation models (VFMs) provide strong local features, most existing methods rely on a single model, overlooking the complementary cues offered by different VFMs. However, exploiting such complementary information inevitably alters token distributions, which challenges the stability of existing query-based global aggregation schemes. To address these challenges, we propose DC-VLAQ, a representation-centric framework that integrates the fusion of complementary VFMs and robust global aggregation. Specifically, we first introduce a lightweight residual-guided complementary fusion that anchors representations in the DINOv2 feature space while injecting complementary semantics from CLIP through a learned residual correction. In addition, we propose the Vector of Local Aggregated Queries (VLAQ), a query--residual global aggregation scheme that encodes local tokens by their residual responses to learnable queries, resulting in improved stability and the preservation of fine-grained discriminative cues. Extensive experiments on standard VPR benchmarks, including Pitts30k, Tokyo24/7, MSLS, Nordland, SPED, and AmsterTime, demonstrate that DC-VLAQ consistently outperforms strong baselines and achieves state-of-the-art performance, particularly under challenging domain shifts and long-term appearance changes.
Stable Language Guidance for Vision-Language-Action Models
Jan 07, 2026Abstract:Vision-Language-Action (VLA) models have demonstrated impressive capabilities in generalized robotic control; however, they remain notoriously brittle to linguistic perturbations. We identify a critical ``modality collapse'' phenomenon where strong visual priors overwhelm sparse linguistic signals, causing agents to overfit to specific instruction phrasings while ignoring the underlying semantic intent. To address this, we propose \textbf{Residual Semantic Steering (RSS)}, a probabilistic framework that disentangles physical affordance from semantic execution. RSS introduces two theoretical innovations: (1) \textbf{Monte Carlo Syntactic Integration}, which approximates the true semantic posterior via dense, LLM-driven distributional expansion, and (2) \textbf{Residual Affordance Steering}, a dual-stream decoding mechanism that explicitly isolates the causal influence of language by subtracting the visual affordance prior. Theoretical analysis suggests that RSS effectively maximizes the mutual information between action and intent while suppressing visual distractors. Empirical results across diverse manipulation benchmarks demonstrate that RSS achieves state-of-the-art robustness, maintaining performance even under adversarial linguistic perturbations.
High-Quality Spatial Reconstruction and Orthoimage Generation Using Efficient 2D Gaussian Splatting
Mar 25, 2025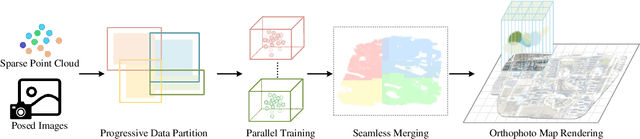
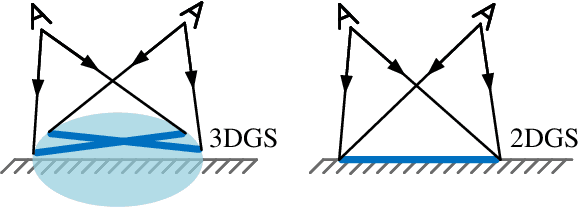
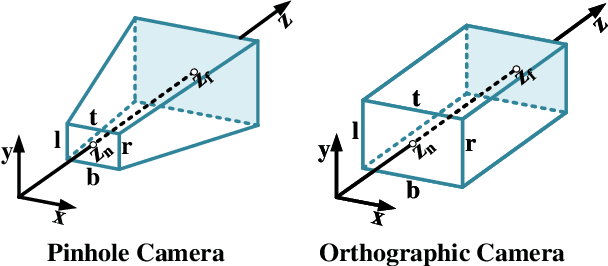
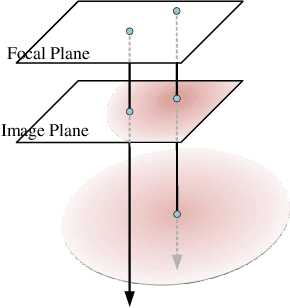
Abstract:Highly accurate geometric precision and dense image features characterize True Digital Orthophoto Maps (TDOMs), which are in great demand for applications such as urban planning, infrastructure management, and environmental monitoring. Traditional TDOM generation methods need sophisticated processes, such as Digital Surface Models (DSM) and occlusion detection, which are computationally expensive and prone to errors. This work presents an alternative technique rooted in 2D Gaussian Splatting (2DGS), free of explicit DSM and occlusion detection. With depth map generation, spatial information for every pixel within the TDOM is retrieved and can reconstruct the scene with high precision. Divide-and-conquer strategy achieves excellent GS training and rendering with high-resolution TDOMs at a lower resource cost, which preserves higher quality of rendering on complex terrain and thin structure without a decrease in efficiency. Experimental results demonstrate the efficiency of large-scale scene reconstruction and high-precision terrain modeling. This approach provides accurate spatial data, which assists users in better planning and decision-making based on maps.
A Multi-Sensor Fusion Approach for Rapid Orthoimage Generation in Large-Scale UAV Mapping
Mar 05, 2025Abstract:Rapid generation of large-scale orthoimages from Unmanned Aerial Vehicles (UAVs) has been a long-standing focus of research in the field of aerial mapping. A multi-sensor UAV system, integrating the Global Positioning System (GPS), Inertial Measurement Unit (IMU), 4D millimeter-wave radar and camera, can provide an effective solution to this problem. In this paper, we utilize multi-sensor data to overcome the limitations of conventional orthoimage generation methods in terms of temporal performance, system robustness, and geographic reference accuracy. A prior-pose-optimized feature matching method is introduced to enhance matching speed and accuracy, reducing the number of required features and providing precise references for the Structure from Motion (SfM) process. The proposed method exhibits robustness in low-texture scenes like farmlands, where feature matching is difficult. Experiments show that our approach achieves accurate feature matching orthoimage generation in a short time. The proposed drone system effectively aids in farmland detection and management.
Video Super-Resolution: All You Need is a Video Diffusion Model
Mar 05, 2025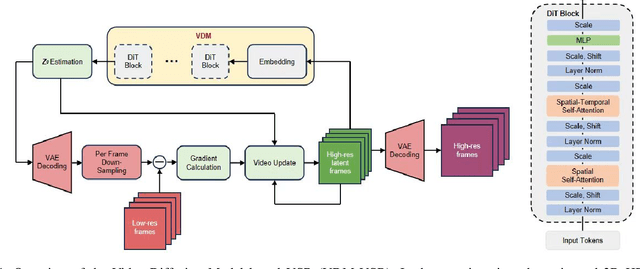
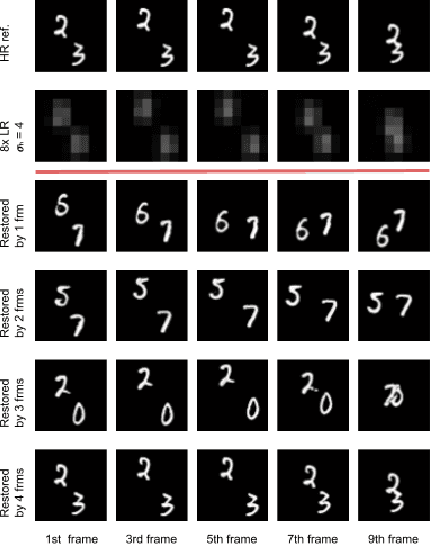

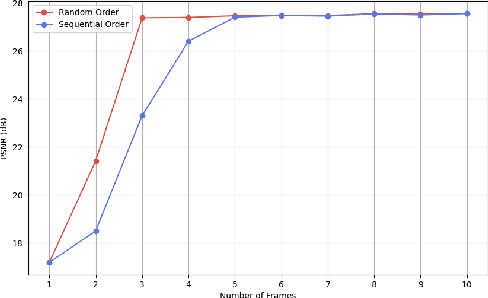

Abstract:We present a generic video super-resolution algorithm in this paper, based on the Diffusion Posterior Sampling framework with an unconditional video generation model in latent space. The video generation model, a diffusion transformer, functions as a space-time model. We argue that a powerful model, which learns the physics of the real world, can easily handle various kinds of motion patterns as prior knowledge, thus eliminating the need for explicit estimation of optical flows or motion parameters for pixel alignment. Furthermore, a single instance of the proposed video diffusion transformer model can adapt to different sampling conditions without re-training. Due to limited computational resources and training data, our experiments provide empirical evidence of the algorithm's strong super-resolution capabilities using synthetic data.
Cell-ontology guided transcriptome foundation model
Aug 22, 2024



Abstract:Transcriptome foundation models TFMs hold great promises of deciphering the transcriptomic language that dictate diverse cell functions by self-supervised learning on large-scale single-cell gene expression data, and ultimately unraveling the complex mechanisms of human diseases. However, current TFMs treat cells as independent samples and ignore the taxonomic relationships between cell types, which are available in cell ontology graphs. We argue that effectively leveraging this ontology information during the TFM pre-training can improve learning biologically meaningful gene co-expression patterns while preserving TFM as a general purpose foundation model for downstream zero-shot and fine-tuning tasks. To this end, we present \textbf{s}ingle \textbf{c}ell, \textbf{Cell}-\textbf{o}ntology guided TFM scCello. We introduce cell-type coherence loss and ontology alignment loss, which are minimized along with the masked gene expression prediction loss during the pre-training. The novel loss component guide scCello to learn the cell-type-specific representation and the structural relation between cell types from the cell ontology graph, respectively. We pre-trained scCello on 22 million cells from CellxGene database leveraging their cell-type labels mapped to the cell ontology graph from Open Biological and Biomedical Ontology Foundry. Our TFM demonstrates competitive generalization and transferability performance over the existing TFMs on biologically important tasks including identifying novel cell types of unseen cells, prediction of cell-type-specific marker genes, and cancer drug responses.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge