Yufan Guo
See, Think, Act: Online Shopper Behavior Simulation with VLM Agents
Oct 22, 2025Abstract:LLMs have recently demonstrated strong potential in simulating online shopper behavior. Prior work has improved action prediction by applying SFT on action traces with LLM-generated rationales, and by leveraging RL to further enhance reasoning capabilities. Despite these advances, current approaches rely on text-based inputs and overlook the essential role of visual perception in shaping human decision-making during web GUI interactions. In this paper, we investigate the integration of visual information, specifically webpage screenshots, into behavior simulation via VLMs, leveraging OPeRA dataset. By grounding agent decision-making in both textual and visual modalities, we aim to narrow the gap between synthetic agents and real-world users, thereby enabling more cognitively aligned simulations of online shopping behavior. Specifically, we employ SFT for joint action prediction and rationale generation, conditioning on the full interaction context, which comprises action history, past HTML observations, and the current webpage screenshot. To further enhance reasoning capabilities, we integrate RL with a hierarchical reward structure, scaled by a difficulty-aware factor that prioritizes challenging decision points. Empirically, our studies show that incorporating visual grounding yields substantial gains: the combination of text and image inputs improves exact match accuracy by more than 6% over text-only inputs. These results indicate that multi-modal grounding not only boosts predictive accuracy but also enhances simulation fidelity in visually complex environments, which captures nuances of human attention and decision-making that text-only agents often miss. Finally, we revisit the design space of behavior simulation frameworks, identify key methodological limitations, and propose future research directions toward building efficient and effective human behavior simulators.
Shop-R1: Rewarding LLMs to Simulate Human Behavior in Online Shopping via Reinforcement Learning
Jul 23, 2025

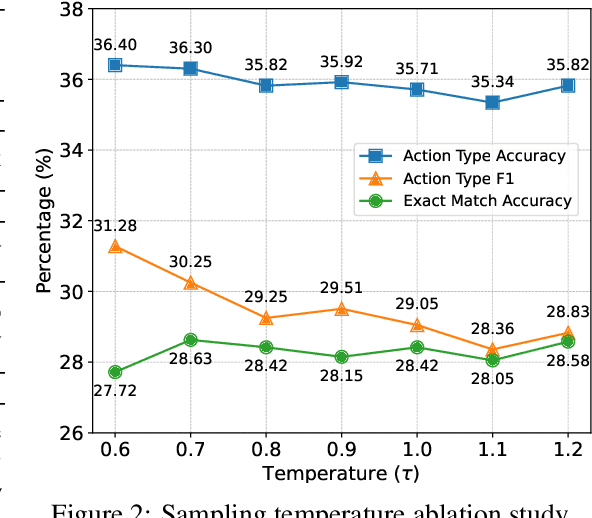

Abstract:Large Language Models (LLMs) have recently demonstrated strong potential in generating 'believable human-like' behavior in web environments. Prior work has explored augmenting training data with LLM-synthesized rationales and applying supervised fine-tuning (SFT) to enhance reasoning ability, which in turn can improve downstream action prediction. However, the performance of such approaches remains inherently bounded by the reasoning capabilities of the model used to generate the rationales. In this paper, we introduce Shop-R1, a novel reinforcement learning (RL) framework aimed at enhancing the reasoning ability of LLMs for simulation of real human behavior in online shopping environments Specifically, Shop-R1 decomposes the human behavior simulation task into two stages: rationale generation and action prediction, each guided by distinct reward signals. For rationale generation, we leverage internal model signals (e.g., logit distributions) to guide the reasoning process in a self-supervised manner. For action prediction, we propose a hierarchical reward structure with difficulty-aware scaling to prevent reward hacking and enable fine-grained reward assignment. This design evaluates both high-level action types and the correctness of fine-grained sub-action details (attributes and values), rewarding outputs proportionally to their difficulty. Experimental results show that our method achieves a relative improvement of over 65% compared to the baseline.
PIG: Physically-based Multi-Material Interaction with 3D Gaussians
Jun 09, 2025Abstract:3D Gaussian Splatting has achieved remarkable success in reconstructing both static and dynamic 3D scenes. However, in a scene represented by 3D Gaussian primitives, interactions between objects suffer from inaccurate 3D segmentation, imprecise deformation among different materials, and severe rendering artifacts. To address these challenges, we introduce PIG: Physically-Based Multi-Material Interaction with 3D Gaussians, a novel approach that combines 3D object segmentation with the simulation of interacting objects in high precision. Firstly, our method facilitates fast and accurate mapping from 2D pixels to 3D Gaussians, enabling precise 3D object-level segmentation. Secondly, we assign unique physical properties to correspondingly segmented objects within the scene for multi-material coupled interactions. Finally, we have successfully embedded constraint scales into deformation gradients, specifically clamping the scaling and rotation properties of the Gaussian primitives to eliminate artifacts and achieve geometric fidelity and visual consistency. Experimental results demonstrate that our method not only outperforms the state-of-the-art (SOTA) in terms of visual quality, but also opens up new directions and pipelines for the field of physically realistic scene generation.
When and Why does a Model Fail? A Human-in-the-loop Error Detection Framework for Sentiment Analysis
Jun 02, 2021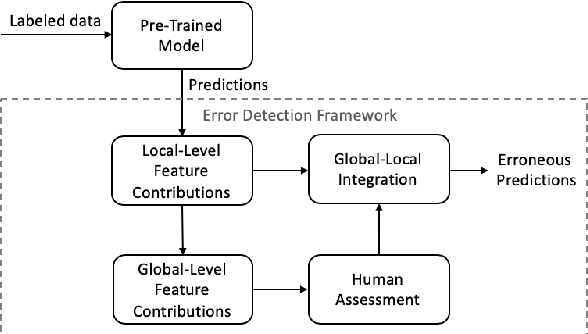


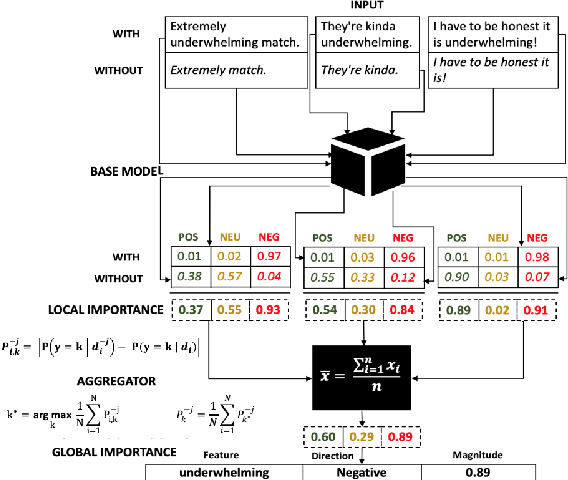
Abstract:Although deep neural networks have been widely employed and proven effective in sentiment analysis tasks, it remains challenging for model developers to assess their models for erroneous predictions that might exist prior to deployment. Once deployed, emergent errors can be hard to identify in prediction run-time and impossible to trace back to their sources. To address such gaps, in this paper we propose an error detection framework for sentiment analysis based on explainable features. We perform global-level feature validation with human-in-the-loop assessment, followed by an integration of global and local-level feature contribution analysis. Experimental results show that, given limited human-in-the-loop intervention, our method is able to identify erroneous model predictions on unseen data with high precision.
Bimodal network architectures for automatic generation of image annotation from text
Sep 05, 2018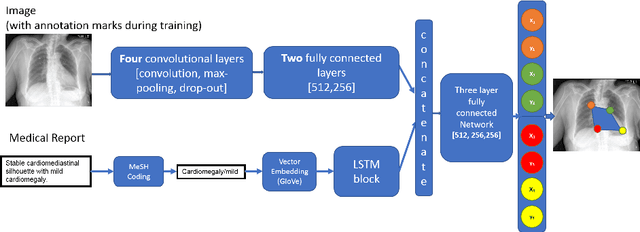

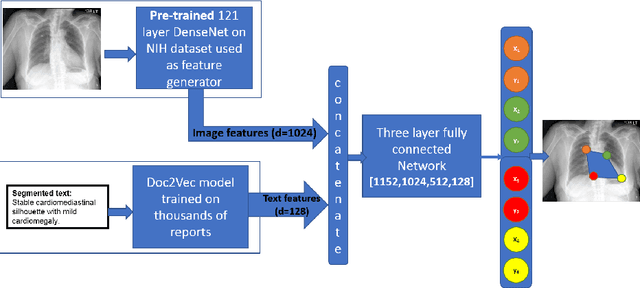
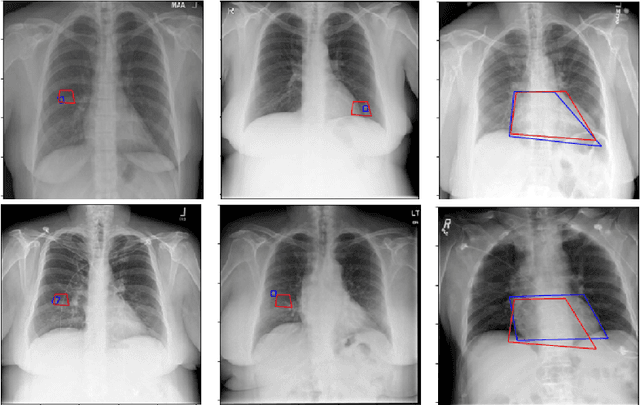
Abstract:Medical image analysis practitioners have embraced big data methodologies. This has created a need for large annotated datasets. The source of big data is typically large image collections and clinical reports recorded for these images. In many cases, however, building algorithms aimed at segmentation and detection of disease requires a training dataset with markings of the areas of interest on the image that match with the described anomalies. This process of annotation is expensive and needs the involvement of clinicians. In this work we propose two separate deep neural network architectures for automatic marking of a region of interest (ROI) on the image best representing a finding location, given a textual report or a set of keywords. One architecture consists of LSTM and CNN components and is trained end to end with images, matching text, and markings of ROIs for those images. The output layer estimates the coordinates of the vertices of a polygonal region. The second architecture uses a network pre-trained on a large dataset of the same image types for learning feature representations of the findings of interest. We show that for a variety of findings from chest X-ray images, both proposed architectures learn to estimate the ROI, as validated by clinical annotations. There is a clear advantage obtained from the architecture with pre-trained imaging network. The centroids of the ROIs marked by this network were on average at a distance equivalent to 5.1% of the image width from the centroids of the ground truth ROIs.
* Accepted to MICCAI 2018, LNCS 11070
Erratum: Link prediction in drug-target interactions network using similarity indices
Nov 01, 2017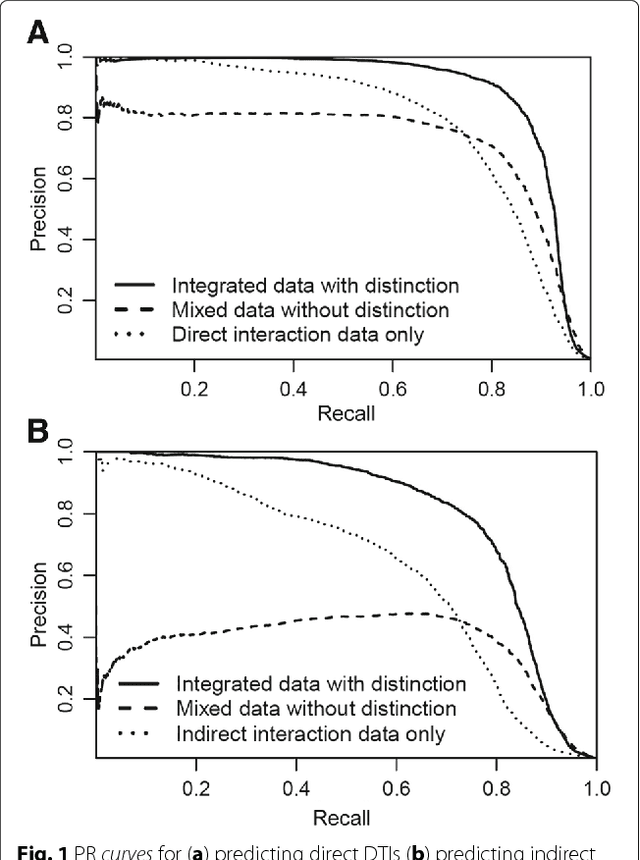

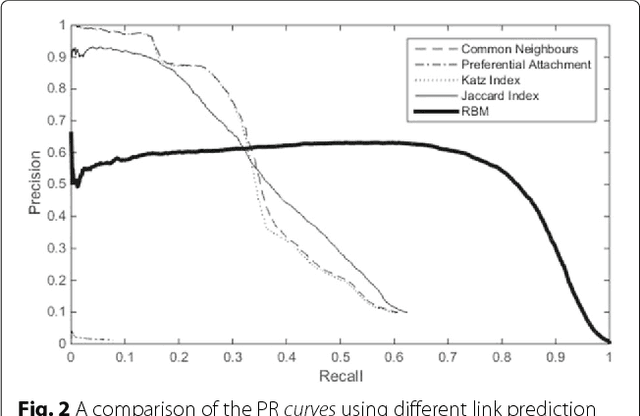
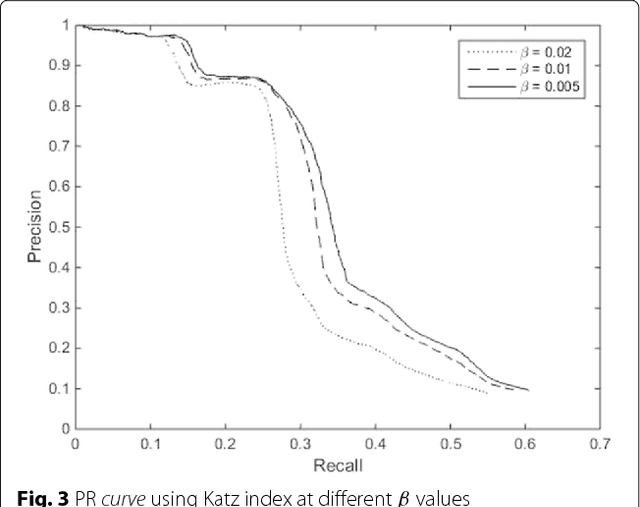
Abstract:Background: In silico drug-target interaction (DTI) prediction plays an integral role in drug repositioning: the discovery of new uses for existing drugs. One popular method of drug repositioning is network-based DTI prediction, which uses complex network theory to predict DTIs from a drug-target network. Currently, most network-based DTI prediction is based on machine learning methods such as Restricted Boltzmann Machines (RBM) or Support Vector Machines (SVM). These methods require additional information about the characteristics of drugs, targets and DTIs, such as chemical structure, genome sequence, binding types, causes of interactions, etc., and do not perform satisfactorily when such information is unavailable. We propose a new, alternative method for DTI prediction that makes use of only network topology information attempting to solve this problem. Results: We compare our method for DTI prediction against the well-known RBM approach. We show that when applied to the MATADOR database, our approach based on node neighborhoods yield higher precision for high-ranking predictions than RBM when no information regarding DTI types is available. Conclusion: This demonstrates that approaches purely based on network topology provide a more suitable approach to DTI prediction in the many real-life situations where little or no prior knowledge is available about the characteristics of drugs, targets, or their interactions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge