Yu-Xing Tang
Learning from Multiple Datasets with Heterogeneous and Partial Labels for Universal Lesion Detection in CT
Sep 05, 2020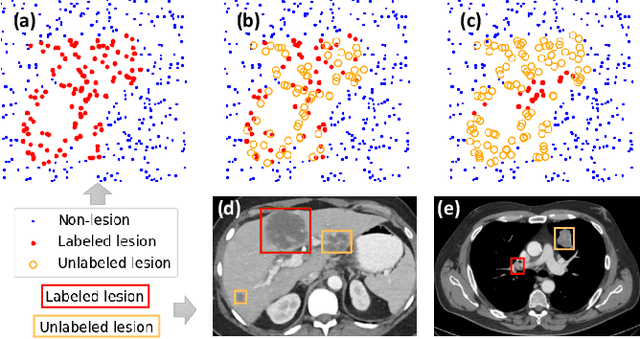
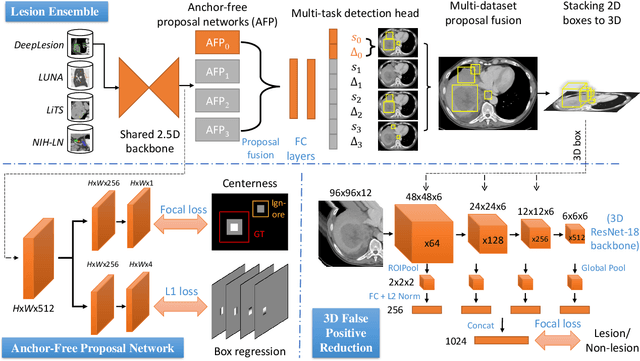
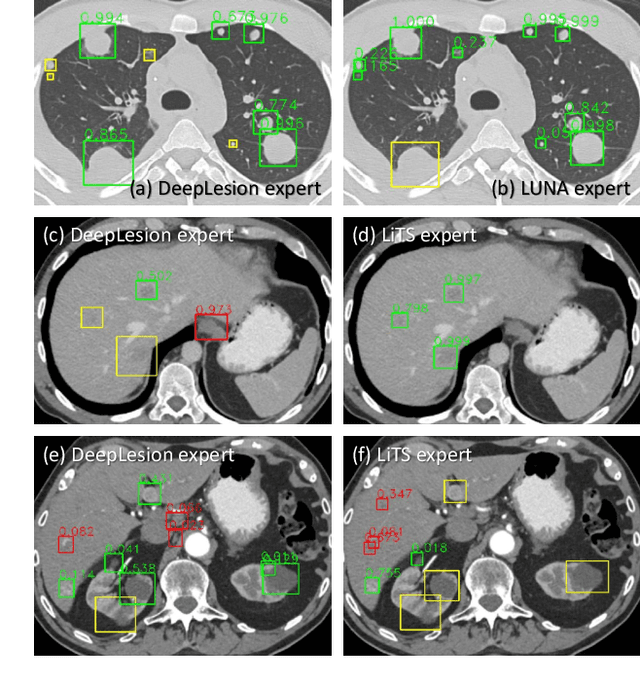
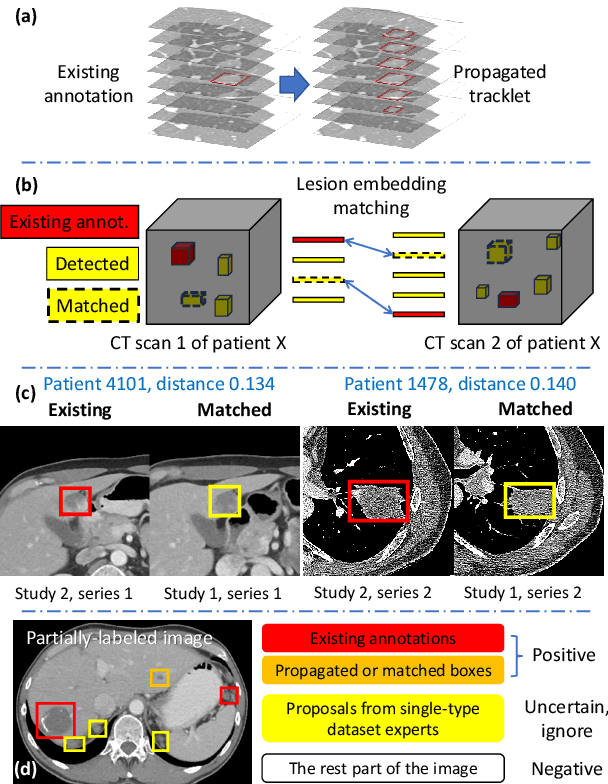
Abstract:Large-scale datasets with high-quality labels are desired for training accurate deep learning models. However, due to annotation costs, medical imaging datasets are often either partially-labeled or small. For example, DeepLesion is a large-scale CT image dataset with lesions of various types, but it also has many unlabeled lesions (missing annotations). When training a lesion detector on a partially-labeled dataset, the missing annotations will generate incorrect negative signals and degrade performance. Besides DeepLesion, there are several small single-type datasets, such as LUNA for lung nodules and LiTS for liver tumors. Such datasets have heterogeneous label scopes, i.e., different lesion types are labeled in different datasets with other types ignored. In this work, we aim to tackle the problem of heterogeneous and partial labels, and develop a universal lesion detection algorithm to detect a comprehensive variety of lesions. First, we build a simple yet effective lesion detection framework named Lesion ENSemble (LENS). LENS can efficiently learn from multiple heterogeneous lesion datasets in a multi-task fashion and leverage their synergy by feature sharing and proposal fusion. Next, we propose strategies to mine missing annotations from partially-labeled datasets by exploiting clinical prior knowledge and cross-dataset knowledge transfer. Finally, we train our framework on four public lesion datasets and evaluate it on 800 manually-labeled sub-volumes in DeepLesion. On this challenging task, our method brings a relative improvement of 49% compared to the current state-of-the-art approach.
COVID-19-CT-CXR: a freely accessible and weakly labeled chest X-ray and CT image collection on COVID-19 from biomedical literature
Jun 11, 2020

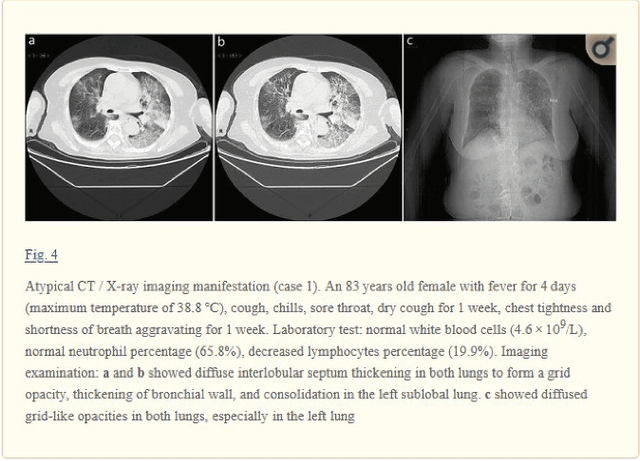
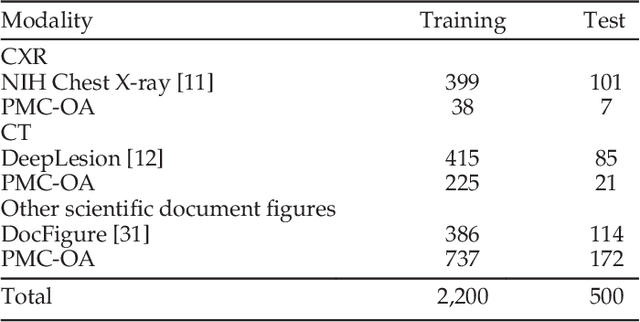
Abstract:The latest threat to global health is the COVID-19 outbreak. Although there exist large datasets of chest X-rays (CXR) and computed tomography (CT) scans, few COVID-19 image collections are currently available due to patient privacy. At the same time, there is a rapid growth of COVID-19-relevant articles in the biomedical literature. Here, we present COVID-19-CT-CXR, a public database of COVID-19 CXR and CT images, which are automatically extracted from COVID-19-relevant articles from the PubMed Central Open Access (PMC-OA) Subset. We extracted figures, associated captions, and relevant figure descriptions in the article and separated compound figures into subfigures. We also designed a deep-learning model to distinguish them from other figure types and to classify them accordingly. The final database includes 1,327 CT and 263 CXR images (as of May 9, 2020) with their relevant text. To demonstrate the utility of COVID-19-CT-CXR, we conducted four case studies. (1) We show that COVID-19-CT-CXR, when used as additional training data, is able to contribute to improved DL performance for the classification of COVID-19 and non-COVID-19 CT. (2) We collected CT images of influenza and trained a DL baseline to distinguish a diagnosis of COVID-19, influenza, or normal or other types of diseases on CT. (3) We trained an unsupervised one-class classifier from non-COVID-19 CXR and performed anomaly detection to detect COVID-19 CXR. (4) From text-mined captions and figure descriptions, we compared clinical symptoms and clinical findings of COVID-19 vs. those of influenza to demonstrate the disease differences in the scientific publications. We believe that our work is complementary to existing resources and hope that it will contribute to medical image analysis of the COVID-19 pandemic. The dataset, code, and DL models are publicly available at https://github.com/ncbi-nlp/COVID-19-CT-CXR.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge