Yiwei Pan
for the ALFA study
Where is VALDO? VAscular Lesions Detection and segmentatiOn challenge at MICCAI 2021
Aug 15, 2022
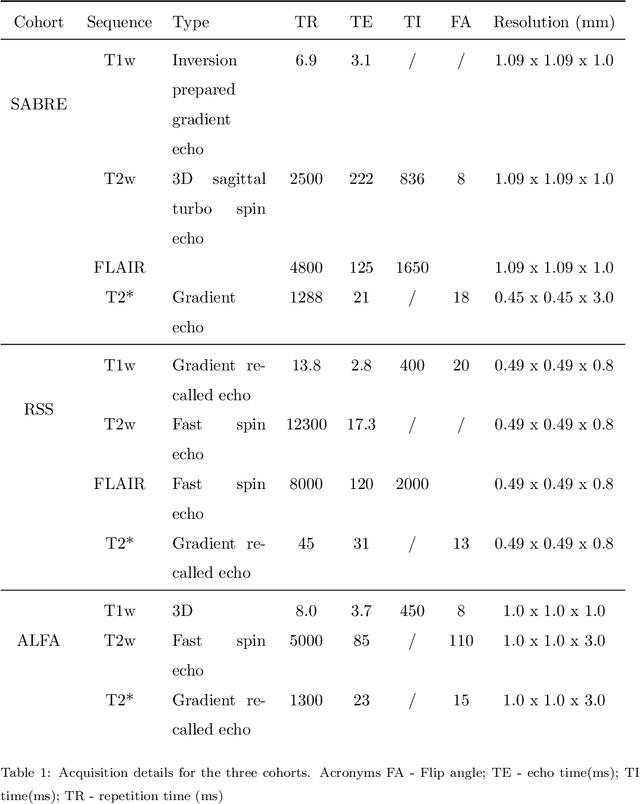
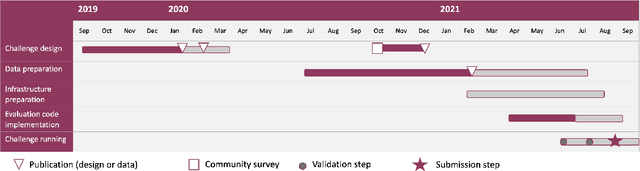
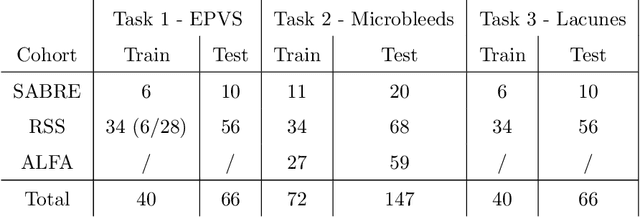
Abstract:Imaging markers of cerebral small vessel disease provide valuable information on brain health, but their manual assessment is time-consuming and hampered by substantial intra- and interrater variability. Automated rating may benefit biomedical research, as well as clinical assessment, but diagnostic reliability of existing algorithms is unknown. Here, we present the results of the \textit{VAscular Lesions DetectiOn and Segmentation} (\textit{Where is VALDO?}) challenge that was run as a satellite event at the international conference on Medical Image Computing and Computer Aided Intervention (MICCAI) 2021. This challenge aimed to promote the development of methods for automated detection and segmentation of small and sparse imaging markers of cerebral small vessel disease, namely enlarged perivascular spaces (EPVS) (Task 1), cerebral microbleeds (Task 2) and lacunes of presumed vascular origin (Task 3) while leveraging weak and noisy labels. Overall, 12 teams participated in the challenge proposing solutions for one or more tasks (4 for Task 1 - EPVS, 9 for Task 2 - Microbleeds and 6 for Task 3 - Lacunes). Multi-cohort data was used in both training and evaluation. Results showed a large variability in performance both across teams and across tasks, with promising results notably for Task 1 - EPVS and Task 2 - Microbleeds and not practically useful results yet for Task 3 - Lacunes. It also highlighted the performance inconsistency across cases that may deter use at an individual level, while still proving useful at a population level.
Uncertainty Quantification in Medical Image Segmentation with Multi-decoder U-Net
Sep 15, 2021
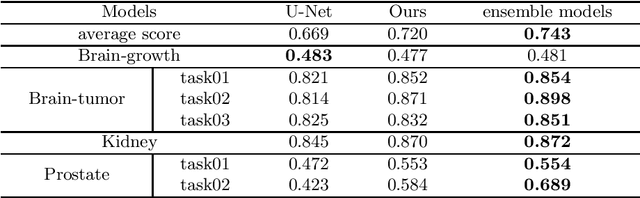
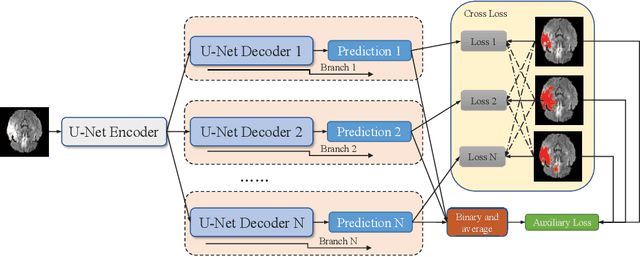
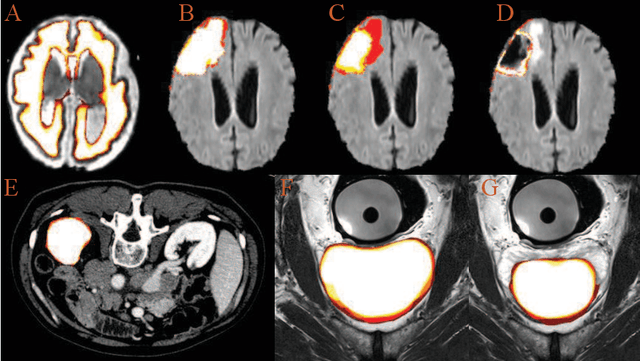
Abstract:Accurate medical image segmentation is crucial for diagnosis and analysis. However, the models without calibrated uncertainty estimates might lead to errors in downstream analysis and exhibit low levels of robustness. Estimating the uncertainty in the measurement is vital to making definite, informed conclusions. Especially, it is difficult to make accurate predictions on ambiguous areas and focus boundaries for both models and radiologists, even harder to reach a consensus with multiple annotations. In this work, the uncertainty under these areas is studied, which introduces significant information with anatomical structure and is as important as segmentation performance. We exploit the medical image segmentation uncertainty quantification by measuring segmentation performance with multiple annotations in a supervised learning manner and propose a U-Net based architecture with multiple decoders, where the image representation is encoded with the same encoder, and segmentation referring to each annotation is estimated with multiple decoders. Nevertheless, a cross-loss function is proposed for bridging the gap between different branches. The proposed architecture is trained in an end-to-end manner and able to improve predictive uncertainty estimates. The model achieves comparable performance with fewer parameters to the integrated training model that ranked the runner-up in the MICCAI-QUBIQ 2020 challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge