Weixiong Zhang
LEDA: Latent Semantic Distribution Alignment for Multi-domain Graph Pre-training
Feb 26, 2026Abstract:Recent advances in generic large models, such as GPT and DeepSeek, have motivated the introduction of universality to graph pre-training, aiming to learn rich and generalizable knowledge across diverse domains using graph representations to improve performance in various downstream applications. However, most existing methods face challenges in learning effective knowledge from generic graphs, primarily due to simplistic data alignment and limited training guidance. The issue of simplistic data alignment arises from the use of a straightforward unification for highly diverse graph data, which fails to align semantics and misleads pre-training models. The problem with limited training guidance lies in the arbitrary application of in-domain pre-training paradigms to cross-domain scenarios. While it is effective in enhancing discriminative representation in one data space, it struggles to capture effective knowledge from many graphs. To address these challenges, we propose a novel Latent sEmantic Distribution Alignment (LEDA) model for universal graph pre-training. Specifically, we first introduce a dimension projection unit to adaptively align diverse domain features into a shared semantic space with minimal information loss. Furthermore, we design a variational semantic inference module to obtain the shared latent distribution. The distribution is then adopted to guide the domain projection, aligning it with shared semantics across domains and ensuring cross-domain semantic learning. LEDA exhibits strong performance across a broad range of graphs and downstream tasks. Remarkably, in few-shot cross-domain settings, it significantly outperforms in-domain baselines and advanced universal pre-training models.
MUG: Meta-path-aware Universal Heterogeneous Graph Pre-Training
Feb 26, 2026Abstract:Universal graph pre-training has emerged as a key paradigm in graph representation learning, offering a promising way to train encoders to learn transferable representations from unlabeled graphs and to effectively generalize across a wide range of downstream tasks. However, recent explorations in universal graph pre-training primarily focus on homogeneous graphs and it remains unexplored for heterogeneous graphs, which exhibit greater structural and semantic complexity. This heterogeneity makes it highly challenging to train a universal encoder for diverse heterogeneous graphs: (i) the diverse types with dataset-specific semantics hinder the construction of a unified representation space; (ii) the number and semantics of meta-paths vary across datasets, making encoding and aggregation patterns learned from one dataset difficult to apply to others. To address these challenges, we propose a novel Meta-path-aware Universal heterogeneous Graph pre-training (MUG) approach. Specifically, for challenge (i), MUG introduces a input unification module that integrates information from multiple node and relation types within each heterogeneous graph into a unified representation.This representation is then projected into a shared space by a dimension-aware encoder, enabling alignment across graphs with diverse schemas.Furthermore, for challenge (ii), MUG trains a shared encoder to capture consistent structural patterns across diverse meta-path views rather than relying on dataset-specific aggregation strategies, while a global objective encourages discriminability and reduces dataset-specific biases. Extensive experiments demonstrate the effectiveness of MUG on some real datasets.
Unified Multi-Domain Graph Pre-training for Homogeneous and Heterogeneous Graphs via Domain-Specific Expert Encoding
Feb 13, 2026Abstract:Graph pre-training has achieved remarkable success in recent years, delivering transferable representations for downstream adaptation. However, most existing methods are designed for either homogeneous or heterogeneous graphs, thereby hindering unified graph modeling across diverse graph types. This separation contradicts real-world applications, where mixed homogeneous and heterogeneous graphs are ubiquitous, and distribution shifts between upstream pre-training and downstream deployment are common. In this paper, we empirically demonstrate that a balanced mixture of homogeneous and heterogeneous graph pre-training benefits downstream tasks and propose a unified multi-domain \textbf{G}raph \textbf{P}re-training method across \textbf{H}omogeneous and \textbf{H}eterogeneous graphs ($\mathbf{GPH^{2}}$). To address the lack of a unified encoder for homogeneous and heterogeneous graphs, we propose a Unified Multi-View Graph Construction that simultaneously encodes both without explicit graph-type-specific designs. To cope with the increased cross-domain distribution discrepancies arising from mixed graphs, we introduce domain-specific expert encoding. Each expert is independently pre-trained on a single graph to capture domain-specific knowledge, thereby shielding the pre-training encoder from the adverse effects of cross-domain discrepancies. For downstream tasks, we further design a Task-oriented Expert Fusion Strategy that adaptively integrates multiple experts based on their discriminative strengths. Extensive experiments on mixed graphs demonstrate that $\text{GPH}^{2}$ enables stable transfer across graph types and domains, significantly outperforming existing graph pre-training methods.
Transformer-based toxin-protein interaction analysis prioritizes airborne particulate matter components with potential adverse health effects
Dec 21, 2024Abstract:Air pollution, particularly airborne particulate matter (PM), poses a significant threat to public health globally. It is crucial to comprehend the association between PM-associated toxic components and their cellular targets in humans to understand the mechanisms by which air pollution impacts health and to establish causal relationships between air pollution and public health consequences. Although many studies have explored the impact of PM on human health, the understanding of the association between toxins and the associated targets remain limited. Leveraging cutting-edge deep learning technologies, we developed tipFormer (toxin-protein interaction prediction based on transformer), a novel deep-learning tool for identifying toxic components capable of penetrating human cells and instigating pathogenic biological activities and signaling cascades. Experimental results show that tipFormer effectively captures interactions between proteins and toxic components. It incorporates dual pre-trained language models to encode protein sequences and chemicals. It employs a convolutional encoder to assimilate the sequential attributes of proteins and chemicals. It then introduces a learning module with a cross-attention mechanism to decode and elucidate the multifaceted interactions pivotal for the hotspots binding proteins and chemicals. Experimental results show that tipFormer effectively captures interactions between proteins and toxic components. This approach offers significant value to air quality and toxicology researchers by allowing high-throughput identification and prioritization of hazards. It supports more targeted laboratory studies and field measurements, ultimately enhancing our understanding of how air pollution impacts human health.
Pixel-global Self-supervised Learning with Uncertainty-aware Context Stabilizer
Oct 02, 2022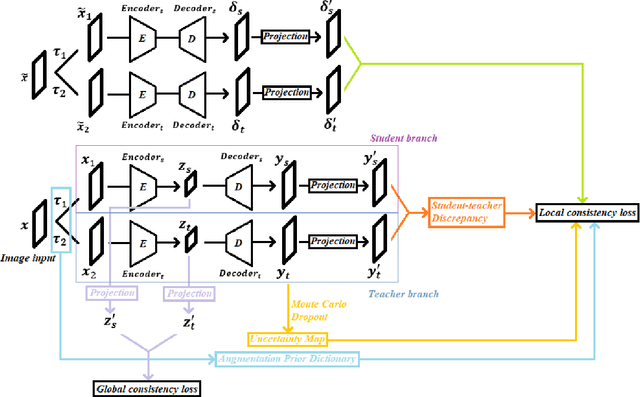


Abstract:We developed a novel SSL approach to capture global consistency and pixel-level local consistencies between differently augmented views of the same images to accommodate downstream discriminative and dense predictive tasks. We adopted the teacher-student architecture used in previous contrastive SSL methods. In our method, the global consistency is enforced by aggregating the compressed representations of augmented views of the same image. The pixel-level consistency is enforced by pursuing similar representations for the same pixel in differently augmented views. Importantly, we introduced an uncertainty-aware context stabilizer to adaptively preserve the context gap created by the two views from different augmentations. Moreover, we used Monte Carlo dropout in the stabilizer to measure uncertainty and adaptively balance the discrepancy between the representations of the same pixels in different views.
RAW-GNN: RAndom Walk Aggregation based Graph Neural Network
Jun 28, 2022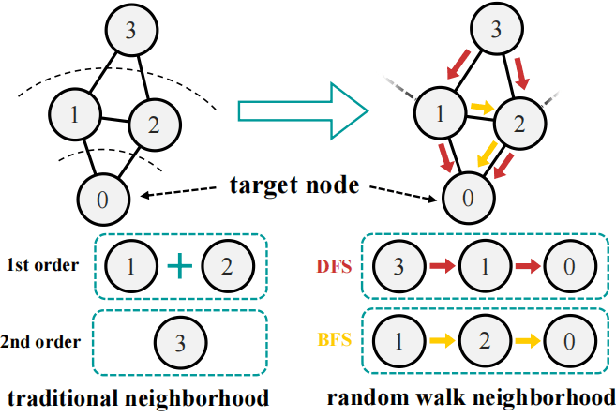
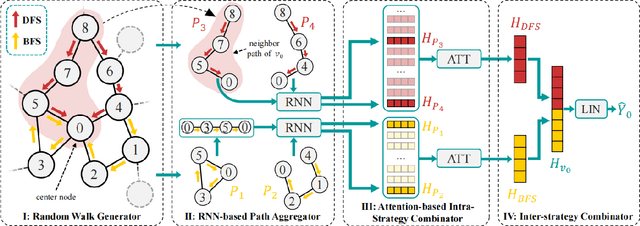
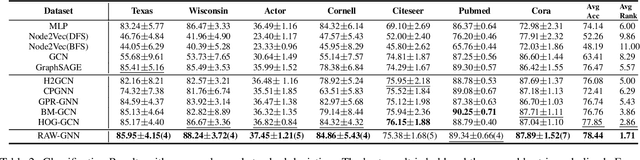
Abstract:Graph-Convolution-based methods have been successfully applied to representation learning on homophily graphs where nodes with the same label or similar attributes tend to connect with one another. Due to the homophily assumption of Graph Convolutional Networks (GCNs) that these methods use, they are not suitable for heterophily graphs where nodes with different labels or dissimilar attributes tend to be adjacent. Several methods have attempted to address this heterophily problem, but they do not change the fundamental aggregation mechanism of GCNs because they rely on summation operators to aggregate information from neighboring nodes, which is implicitly subject to the homophily assumption. Here, we introduce a novel aggregation mechanism and develop a RAndom Walk Aggregation-based Graph Neural Network (called RAW-GNN) method. The proposed approach integrates the random walk strategy with graph neural networks. The new method utilizes breadth-first random walk search to capture homophily information and depth-first search to collect heterophily information. It replaces the conventional neighborhoods with path-based neighborhoods and introduces a new path-based aggregator based on Recurrent Neural Networks. These designs make RAW-GNN suitable for both homophily and heterophily graphs. Extensive experimental results showed that the new method achieved state-of-the-art performance on a variety of homophily and heterophily graphs.
Response to: Significance and stability of deep learning-based identification of subtypes within major psychiatric disorders. Molecular Psychiatry (2022)
Jun 10, 2022Abstract:Recently, Winter and Hahn [1] commented on our work on identifying subtypes of major psychiatry disorders (MPDs) based on neurobiological features using machine learning [2]. They questioned the generalizability of our methods and the statistical significance, stability, and overfitting of the results, and proposed a pipeline for disease subtyping. We appreciate their earnest consideration of our work, however, we need to point out their misconceptions of basic machine-learning concepts and delineate some key issues involved.
Heterogeneous Graph Neural Networks using Self-supervised Reciprocally Contrastive Learning
Apr 30, 2022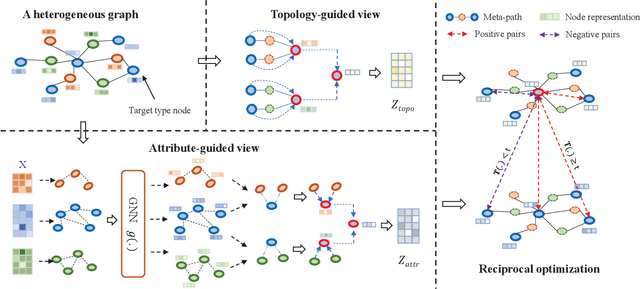
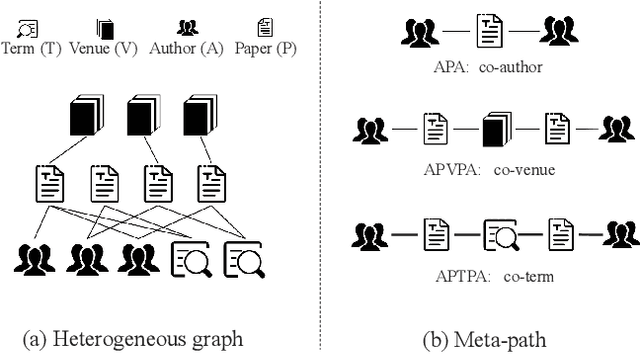
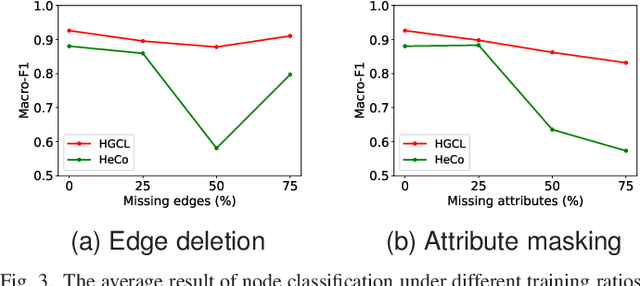
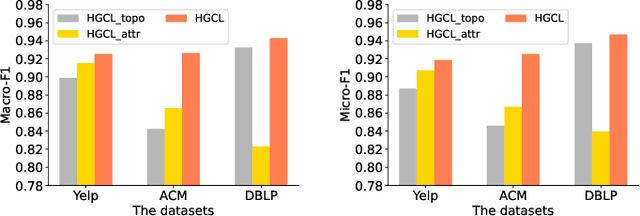
Abstract:Heterogeneous graph neural network (HGNN) is a very popular technique for the modeling and analysis of heterogeneous graphs. Most existing HGNN-based approaches are supervised or semi-supervised learning methods requiring graphs to be annotated, which is costly and time-consuming. Self-supervised contrastive learning has been proposed to address the problem of requiring annotated data by mining intrinsic information hidden within the given data. However, the existing contrastive learning methods are inadequate for heterogeneous graphs because they construct contrastive views only based on data perturbation or pre-defined structural properties (e.g., meta-path) in graph data while ignore the noises that may exist in both node attributes and graph topologies. We develop for the first time a novel and robust heterogeneous graph contrastive learning approach, namely HGCL, which introduces two views on respective guidance of node attributes and graph topologies and integrates and enhances them by reciprocally contrastive mechanism to better model heterogeneous graphs. In this new approach, we adopt distinct but most suitable attribute and topology fusion mechanisms in the two views, which are conducive to mining relevant information in attributes and topologies separately. We further use both attribute similarity and topological correlation to construct high-quality contrastive samples. Extensive experiments on three large real-world heterogeneous graphs demonstrate the superiority and robustness of HGCL over state-of-the-art methods.
Weaving Attention U-net: A Novel Hybrid CNN and Attention-based Method for Organs-at-risk Segmentation in Head and Neck CT Images
Jul 10, 2021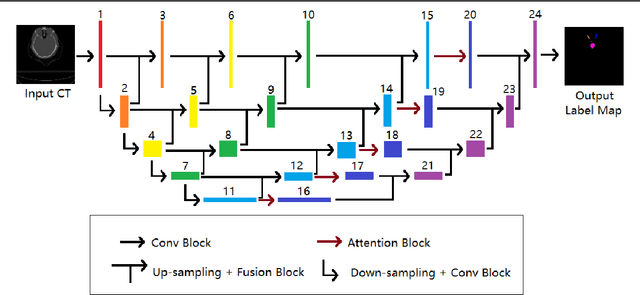
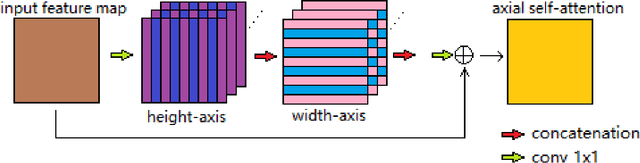
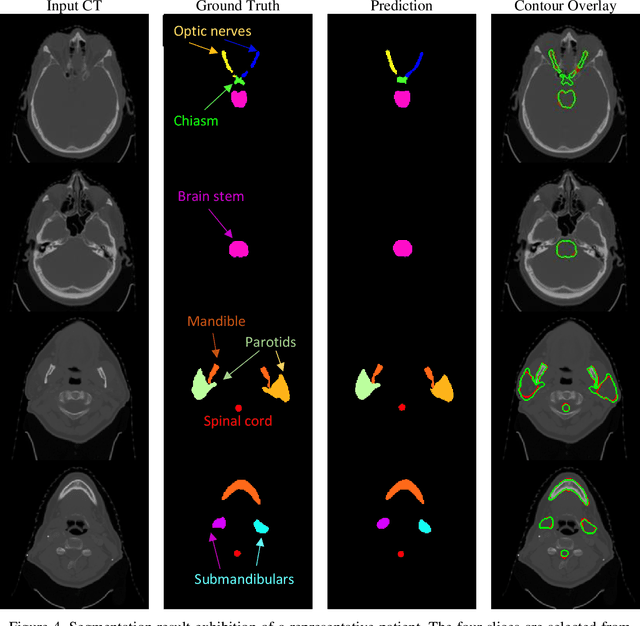
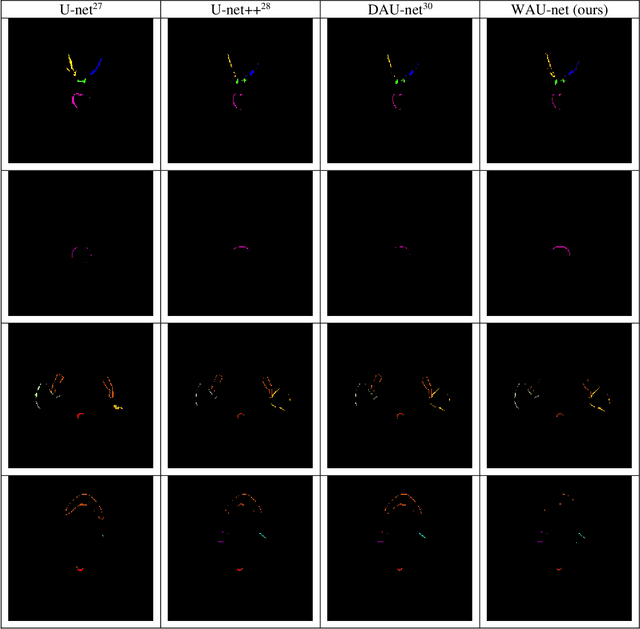
Abstract:In radiotherapy planning, manual contouring is labor-intensive and time-consuming. Accurate and robust automated segmentation models improve the efficiency and treatment outcome. We aim to develop a novel hybrid deep learning approach, combining convolutional neural networks (CNNs) and the self-attention mechanism, for rapid and accurate multi-organ segmentation on head and neck computed tomography (CT) images. Head and neck CT images with manual contours of 115 patients were retrospectively collected and used. We set the training/validation/testing ratio to 81/9/25 and used the 10-fold cross-validation strategy to select the best model parameters. The proposed hybrid model segmented ten organs-at-risk (OARs) altogether for each case. The performance of the model was evaluated by three metrics, i.e., the Dice Similarity Coefficient (DSC), Hausdorff distance 95% (HD95), and mean surface distance (MSD). We also tested the performance of the model on the Head and Neck 2015 challenge dataset and compared it against several state-of-the-art automated segmentation algorithms. The proposed method generated contours that closely resemble the ground truth for ten OARs. Our results of the new Weaving Attention U-net demonstrate superior or similar performance on the segmentation of head and neck CT images.
Pyramid Medical Transformer for Medical Image Segmentation
Apr 29, 2021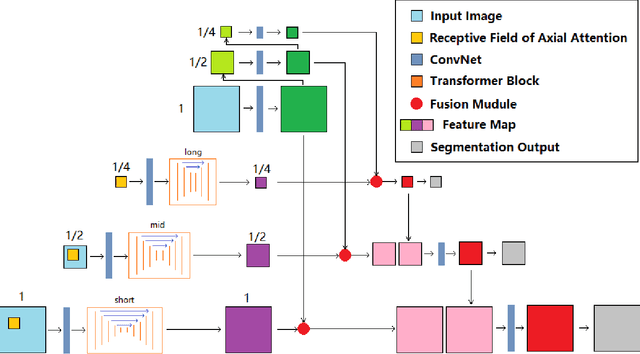

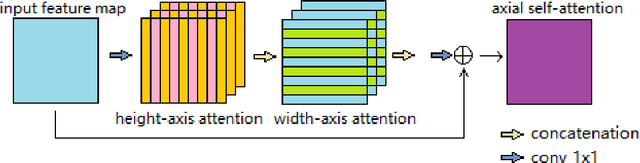
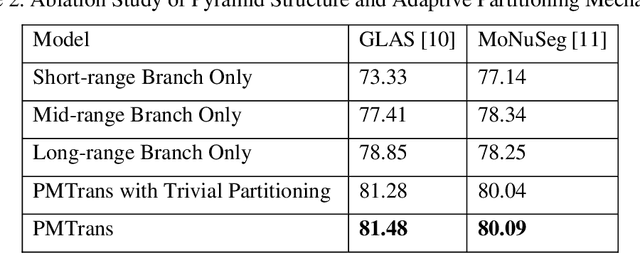
Abstract:Deep neural networks have been a prevailing technique in the field of medical image processing. However, the most popular convolutional neural networks (CNNs) based methods for medical image segmentation are imperfect because they cannot adequately model long-range pixel relations. Transformers and the self-attention mechanism are recently proposed to effectively learn long-range dependencies by modeling all pairs of word-to-word attention regardless of their positions. The idea has also been extended to the computer vision field by creating and treating image patches as embeddings. Considering the computation complexity for whole image self-attention, current transformer-based models settle for a rigid partitioning scheme that would potentially lose informative relations. Besides, current medical transformers model global context on full resolution images, leading to unnecessary computation costs. To address these issues, we developed a novel method to integrate multi-scale attention and CNN feature extraction using a pyramidal network architecture, namely Pyramid Medical Transformer (PMTrans). The PMTrans captured multi-range relations by working on multi-resolution images. An adaptive partitioning scheme was implemented to retain informative relations and to access different receptive fields efficiently. Experimental results on two medical image datasets, gland segmentation and MoNuSeg datasets, showed that PMTrans outperformed the latest CNN-based and transformer-based models for medical image segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge