Vikram Venkatraghavan
for the Alzheimer's Disease Neuroimaging Initiative, on behalf of the Parelsnoer Neurodegenerative Diseases study group
Computer-aided diagnosis and prediction in brain disorders
Jun 29, 2022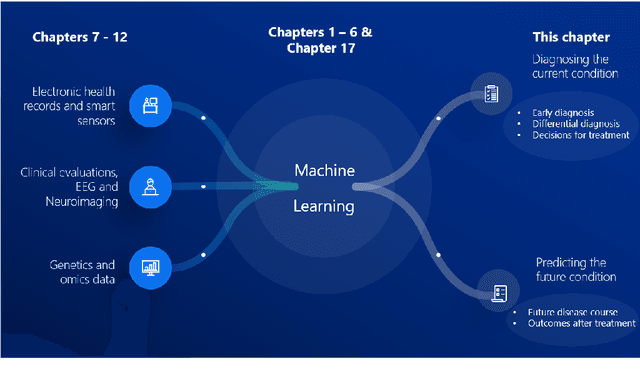
Abstract:Computer-aided methods have shown added value for diagnosing and predicting brain disorders and can thus support decision making in clinical care and treatment planning. This chapter will provide insight into the type of methods, their working, their input data - such as cognitive tests, imaging and genetic data - and the types of output they provide. We will focus on specific use cases for diagnosis, i.e. estimating the current 'condition' of the patient, such as early detection and diagnosis of dementia, differential diagnosis of brain tumours, and decision making in stroke. Regarding prediction, i.e. estimation of the future 'condition' of the patient, we will zoom in on use cases such as predicting the disease course in multiple sclerosis and predicting patient outcomes after treatment in brain cancer. Furthermore, based on these use cases, we will assess the current state-of-the-art methodology and highlight current efforts on benchmarking of these methods and the importance of open science therein. Finally, we assess the current clinical impact of computer-aided methods and discuss the required next steps to increase clinical impact.
Cross-Cohort Generalizability of Deep and Conventional Machine Learning for MRI-based Diagnosis and Prediction of Alzheimer's Disease
Dec 16, 2020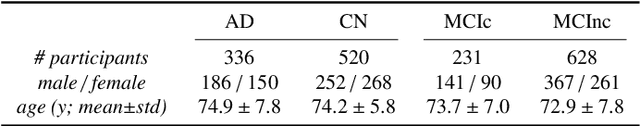
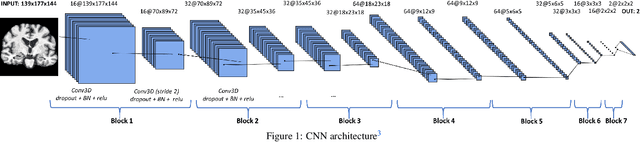
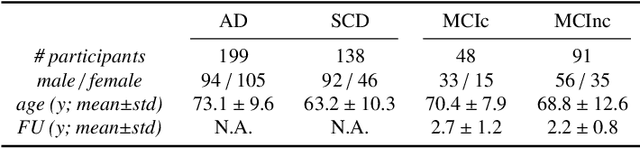
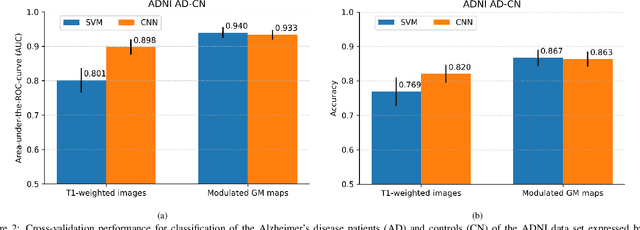
Abstract:This work validates the generalizability of MRI-based classification of Alzheimer's disease (AD) patients and controls (CN) to an external data set and to the task of prediction of conversion to AD in individuals with mild cognitive impairment (MCI). We used a conventional support vector machine (SVM) and a deep convolutional neural network (CNN) approach based on structural MRI scans that underwent either minimal pre-processing or more extensive pre-processing into modulated gray matter (GM) maps. Classifiers were optimized and evaluated using cross-validation in the ADNI (334 AD, 520 CN). Trained classifiers were subsequently applied to predict conversion to AD in ADNI MCI patients (231 converters, 628 non-converters) and in the independent Health-RI Parelsnoer data set. From this multi-center study representing a tertiary memory clinic population, we included 199 AD patients, 139 participants with subjective cognitive decline, 48 MCI patients converting to dementia, and 91 MCI patients who did not convert to dementia. AD-CN classification based on modulated GM maps resulted in a similar AUC for SVM (0.940) and CNN (0.933). Application to conversion prediction in MCI yielded significantly higher performance for SVM (0.756) than for CNN (0.742). In external validation, performance was slightly decreased. For AD-CN, it again gave similar AUCs for SVM (0.896) and CNN (0.876). For prediction in MCI, performances decreased for both SVM (0.665) and CNN (0.702). Both with SVM and CNN, classification based on modulated GM maps significantly outperformed classification based on minimally processed images. Deep and conventional classifiers performed equally well for AD classification and their performance decreased only slightly when applied to the external cohort. We expect that this work on external validation contributes towards translation of machine learning to clinical practice.
Analyzing the effect of APOE on Alzheimer's disease progression using an event-based model for stratified populations
Sep 15, 2020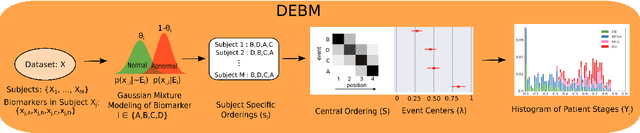
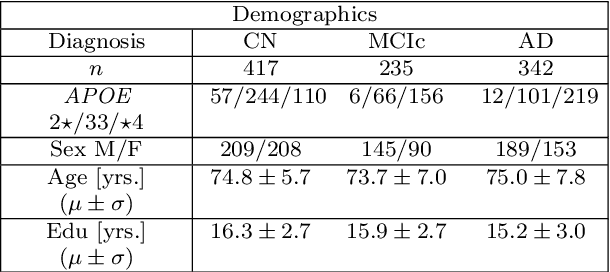
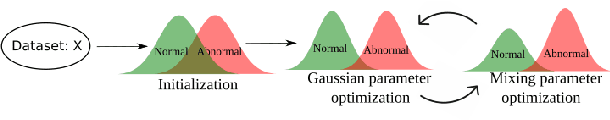

Abstract:Alzheimer's disease (AD) is the most common form of dementia and is phenotypically heterogeneous. APOE is a triallelic gene which correlates with phenotypic heterogeneity in AD. In this work, we determined the effect of APOE alleles on the disease progression timeline of AD using a discriminative event-based model (DEBM). Since DEBM is a data-driven model, stratification into smaller disease subgroups would lead to more inaccurate models as compared to fitting the model on the entire dataset. Hence our secondary aim is to propose and evaluate novel approaches in which we split the different steps of DEBM into group-aspecific and group-specific parts, where the entire dataset is used to train the group-aspecific parts and only the data from a specific group is used to train the group-specific parts of the DEBM. We performed simulation experiments to benchmark the accuracy of the proposed approaches and to select the optimal approach. Subsequently, the chosen approach was applied to the baseline data of 417 cognitively normal, 235 mild cognitively impaired who convert to AD within 3 years, and 342 AD patients from the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset to gain new insights into the effect of APOE carriership on the disease progression timeline of AD. The presented models could aid understanding of the disease, and in selecting homogeneous group of presymptomatic subjects at-risk of developing symptoms for clinical trials.
Event-Based Modeling with High-Dimensional Imaging Biomarkers for Estimating Spatial Progression of Dementia
Mar 08, 2019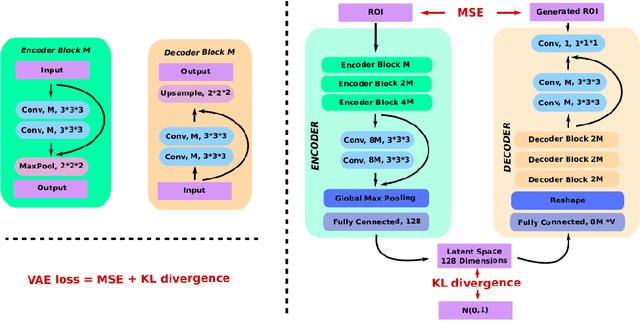
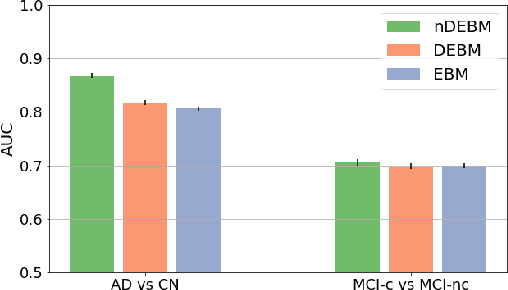
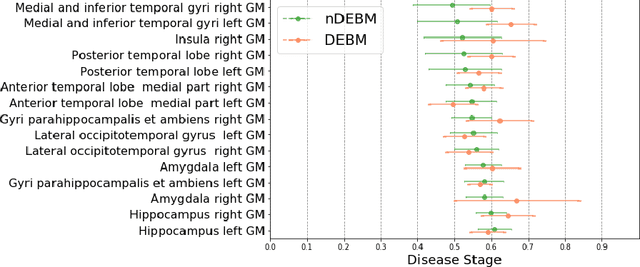
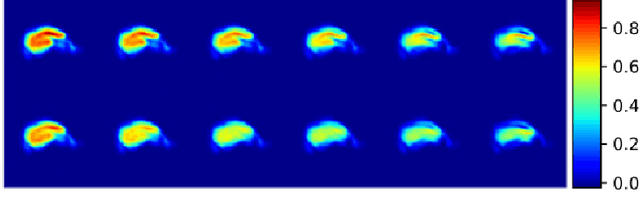
Abstract:Event-based models (EBM) are a class of disease progression models that can be used to estimate temporal ordering of neuropathological changes from cross-sectional data. Current EBMs only handle scalar biomarkers, such as regional volumes, as inputs. However, regional aggregates are a crude summary of the underlying high-resolution images, potentially limiting the accuracy of EBM. Therefore, we propose a novel method that exploits high-dimensional voxel-wise imaging biomarkers: n-dimensional discriminative EBM (nDEBM). nDEBM is based on an insight that mixture modeling, which is a key element of conventional EBMs, can be replaced by a more scalable semi-supervised support vector machine (SVM) approach. This SVM is used to estimate the degree of abnormality of each region which is then used to obtain subject-specific disease progression patterns. These patterns are in turn used for estimating the mean ordering by fitting a generalized Mallows model. In order to validate the biomarker ordering obtained using nDEBM, we also present a framework for Simulation of Imaging Biomarkers' Temporal Evolution (SImBioTE) that mimics neurodegeneration in brain regions. SImBioTE trains variational auto-encoders (VAE) in different brain regions independently to simulate images at varying stages of disease progression. We also validate nDEBM clinically using data from the Alzheimer's Disease Neuroimaging Initiative (ADNI). In both experiments, nDEBM using high-dimensional features gave better performance than state-of-the-art EBM methods using regional volume biomarkers. This suggests that nDEBM is a promising approach for disease progression modeling.
Disease Progression Timeline Estimation for Alzheimer's Disease using Discriminative Event Based Modeling
Aug 10, 2018

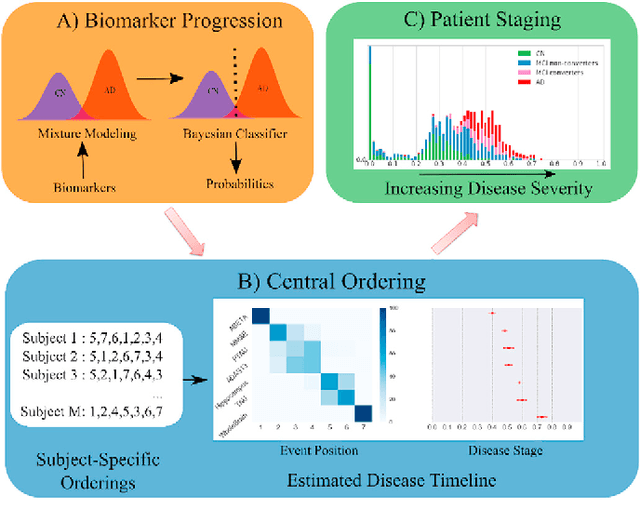
Abstract:Alzheimer's Disease (AD) is characterized by a cascade of biomarkers becoming abnormal, the pathophysiology of which is very complex and largely unknown. Event-based modeling (EBM) is a data-driven technique to estimate the sequence in which biomarkers for a disease become abnormal based on cross-sectional data. It can help in understanding the dynamics of disease progression and facilitate early diagnosis and prognosis. In this work we propose a novel discriminative approach to EBM, which is shown to be more accurate than existing state-of-the-art EBM methods. The method first estimates for each subject an approximate ordering of events. Subsequently, the central ordering over all subjects is estimated by fitting a generalized Mallows model to these approximate subject-specific orderings. We also introduce the concept of relative distance between events which helps in creating a disease progression timeline. Subsequently, we propose a method to stage subjects by placing them on the estimated disease progression timeline. We evaluated the proposed method on Alzheimer's Disease Neuroimaging Initiative (ADNI) data and compared the results with existing state-of-the-art EBM methods. We also performed extensive experiments on synthetic data simulating the progression of Alzheimer's disease. The event orderings obtained on ADNI data seem plausible and are in agreement with the current understanding of progression of AD. The proposed patient staging algorithm performed consistently better than that of state-of-the-art EBM methods. Event orderings obtained in simulation experiments were more accurate than those of other EBM methods and the estimated disease progression timeline was observed to correlate with the timeline of actual disease progression. The results of these experiments are encouraging and suggest that discriminative EBM is a promising approach to disease progression modeling.
A Discriminative Event Based Model for Alzheimer's Disease Progression Modeling
Feb 21, 2017
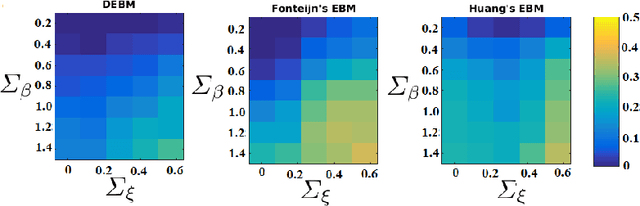
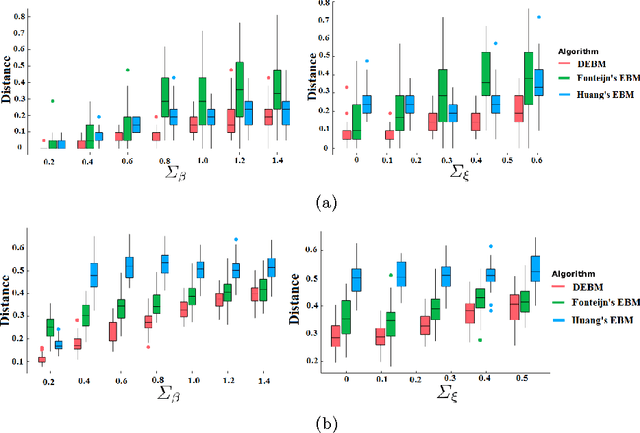
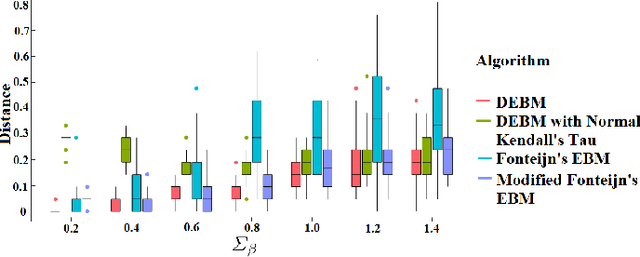
Abstract:The event-based model (EBM) for data-driven disease progression modeling estimates the sequence in which biomarkers for a disease become abnormal. This helps in understanding the dynamics of disease progression and facilitates early diagnosis by staging patients on a disease progression timeline. Existing EBM methods are all generative in nature. In this work we propose a novel discriminative approach to EBM, which is shown to be more accurate as well as computationally more efficient than existing state-of-the art EBM methods. The method first estimates for each subject an approximate ordering of events, by ranking the posterior probabilities of individual biomarkers being abnormal. Subsequently, the central ordering over all subjects is estimated by fitting a generalized Mallows model to these approximate subject-specific orderings based on a novel probabilistic Kendall's Tau distance. To evaluate the accuracy, we performed extensive experiments on synthetic data simulating the progression of Alzheimer's disease. Subsequently, the method was applied to the Alzheimer's Disease Neuroimaging Initiative (ADNI) data to estimate the central event ordering in the dataset. The experiments benchmark the accuracy of the new model under various conditions and compare it with existing state-of-the-art EBM methods. The results indicate that discriminative EBM could be a simple and elegant approach to disease progression modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge