Event-Based Modeling with High-Dimensional Imaging Biomarkers for Estimating Spatial Progression of Dementia
Paper and Code
Mar 08, 2019
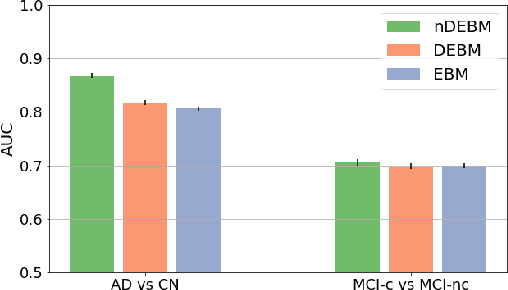
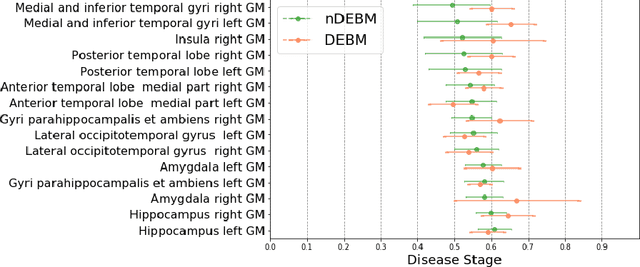
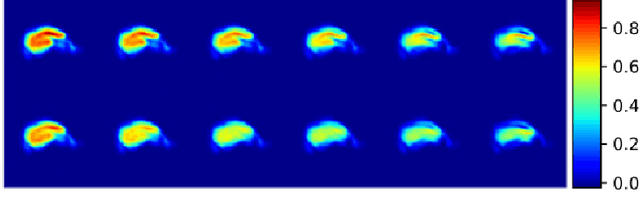
Event-based models (EBM) are a class of disease progression models that can be used to estimate temporal ordering of neuropathological changes from cross-sectional data. Current EBMs only handle scalar biomarkers, such as regional volumes, as inputs. However, regional aggregates are a crude summary of the underlying high-resolution images, potentially limiting the accuracy of EBM. Therefore, we propose a novel method that exploits high-dimensional voxel-wise imaging biomarkers: n-dimensional discriminative EBM (nDEBM). nDEBM is based on an insight that mixture modeling, which is a key element of conventional EBMs, can be replaced by a more scalable semi-supervised support vector machine (SVM) approach. This SVM is used to estimate the degree of abnormality of each region which is then used to obtain subject-specific disease progression patterns. These patterns are in turn used for estimating the mean ordering by fitting a generalized Mallows model. In order to validate the biomarker ordering obtained using nDEBM, we also present a framework for Simulation of Imaging Biomarkers' Temporal Evolution (SImBioTE) that mimics neurodegeneration in brain regions. SImBioTE trains variational auto-encoders (VAE) in different brain regions independently to simulate images at varying stages of disease progression. We also validate nDEBM clinically using data from the Alzheimer's Disease Neuroimaging Initiative (ADNI). In both experiments, nDEBM using high-dimensional features gave better performance than state-of-the-art EBM methods using regional volume biomarkers. This suggests that nDEBM is a promising approach for disease progression modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge