Tianrun Cai
Representation Learning to Advance Multi-institutional Studies with Electronic Health Record Data
Feb 12, 2025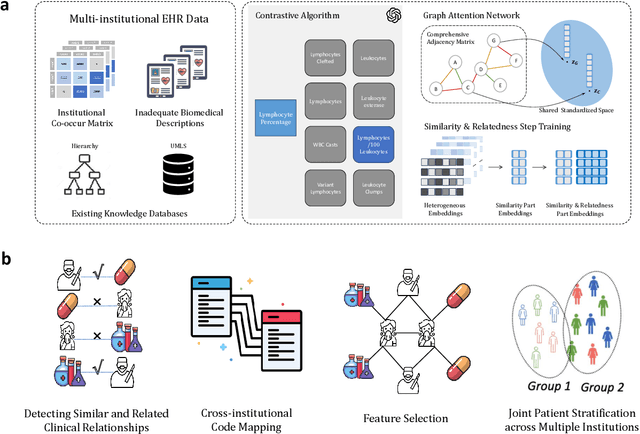
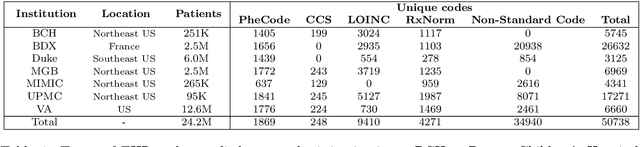
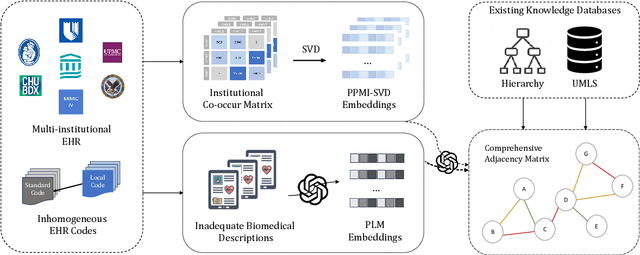
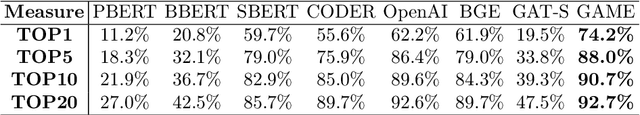
Abstract:The adoption of EHRs has expanded opportunities to leverage data-driven algorithms in clinical care and research. A major bottleneck in effectively conducting multi-institutional EHR studies is the data heterogeneity across systems with numerous codes that either do not exist or represent different clinical concepts across institutions. The need for data privacy further limits the feasibility of including multi-institutional patient-level data required to study similarities and differences across patient subgroups. To address these challenges, we developed the GAME algorithm. Tested and validated across 7 institutions and 2 languages, GAME integrates data in several levels: (1) at the institutional level with knowledge graphs to establish relationships between codes and existing knowledge sources, providing the medical context for standard codes and their relationship to each other; (2) between institutions, leveraging language models to determine the relationships between institution-specific codes with established standard codes; and (3) quantifying the strength of the relationships between codes using a graph attention network. Jointly trained embeddings are created using transfer and federated learning to preserve data privacy. In this study, we demonstrate the applicability of GAME in selecting relevant features as inputs for AI-driven algorithms in a range of conditions, e.g., heart failure, rheumatoid arthritis. We then highlight the application of GAME harmonized multi-institutional EHR data in a study of Alzheimer's disease outcomes and suicide risk among patients with mental health disorders, without sharing patient-level data outside individual institutions.
LATTE: Label-efficient Incident Phenotyping from Longitudinal Electronic Health Records
May 19, 2023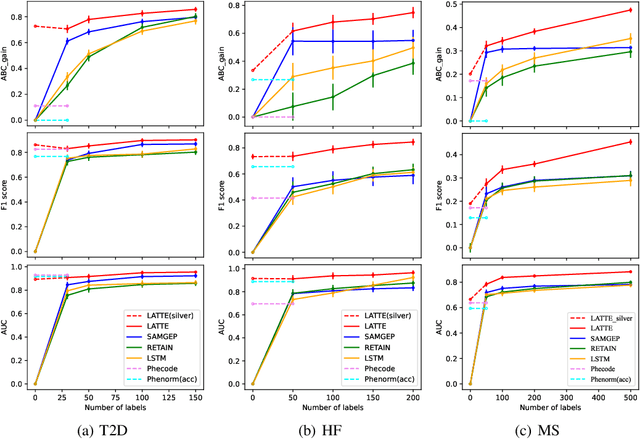
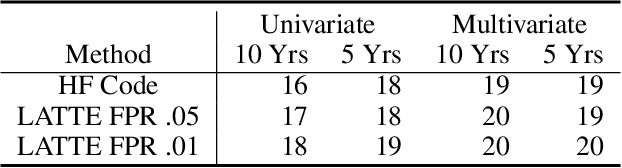
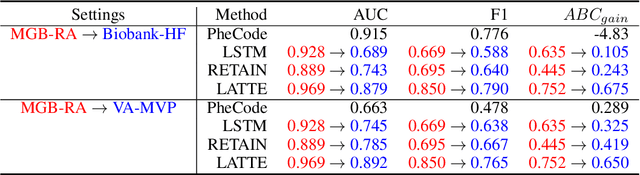
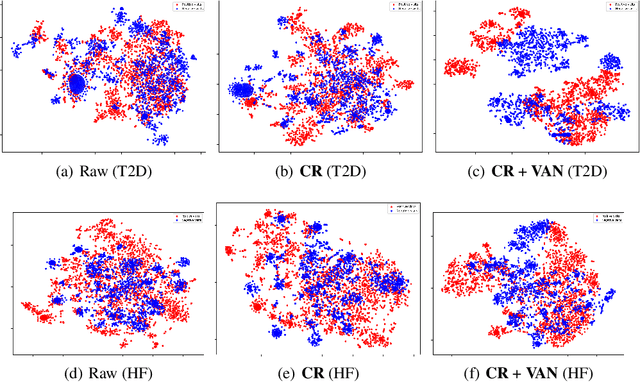

Abstract:Electronic health record (EHR) data are increasingly used to support real-world evidence (RWE) studies. Yet its ability to generate reliable RWE is limited by the lack of readily available precise information on the timing of clinical events such as the onset time of heart failure. We propose a LAbel-efficienT incidenT phEnotyping (LATTE) algorithm to accurately annotate the timing of clinical events from longitudinal EHR data. By leveraging the pre-trained semantic embedding vectors from large-scale EHR data as prior knowledge, LATTE selects predictive EHR features in a concept re-weighting module by mining their relationship to the target event and compresses their information into longitudinal visit embeddings through a visit attention learning network. LATTE employs a recurrent neural network to capture the sequential dependency between the target event and visit embeddings before/after it. To improve label efficiency, LATTE constructs highly informative longitudinal silver-standard labels from large-scale unlabeled patients to perform unsupervised pre-training and semi-supervised joint training. Finally, LATTE enhances cross-site portability via contrastive representation learning. LATTE is evaluated on three analyses: the onset of type-2 diabetes, heart failure, and the onset and relapses of multiple sclerosis. We use various evaluation metrics present in the literature including the $ABC_{gain}$, the proportion of reduction in the area between the observed event indicator and the predicted cumulative incidences in reference to the prediction per incident prevalence. LATTE consistently achieves substantial improvement over benchmark methods such as SAMGEP and RETAIN in all settings.
Unsupervised Ensemble Learning via Ising Model Approximation with Application to Phenotyping Prediction
Oct 15, 2018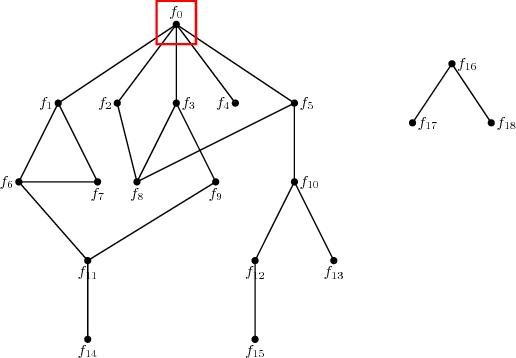
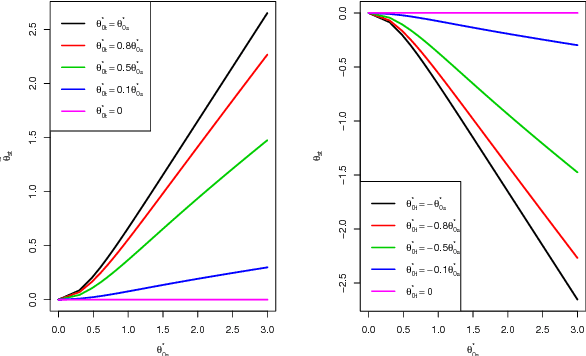
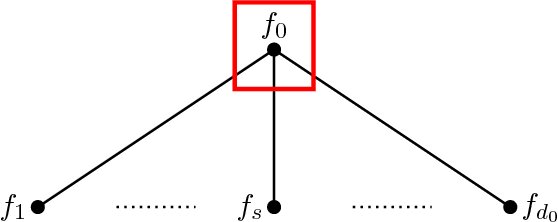
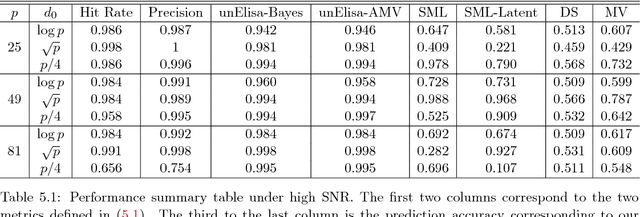
Abstract:Unsupervised ensemble learning has long been an interesting yet challenging problem that comes to prominence in recent years with the increasing demand of crowdsourcing in various applications. In this paper, we propose a novel method-- unsupervised ensemble learning via Ising model approximation (unElisa) that combines a pruning step with a predicting step. We focus on the binary case and use an Ising model to characterize interactions between the ensemble and the underlying true classifier. The presence of an edge between an observed classifier and the true classifier indicates a direct dependence whereas the absence indicates the corresponding one provides no additional information and shall be eliminated. This observation leads to the pruning step where the key is to recover the neighborhood of the true classifier. We show that it can be recovered successfully with exponentially decaying error in the high-dimensional setting by performing nodewise $\ell_1$-regularized logistic regression. The pruned ensemble allows us to get a consistent estimate of the Bayes classifier for predicting. We also propose an augmented version of majority voting by reversing all labels given by a subgroup of the pruned ensemble. We demonstrate the efficacy of our method through extensive numerical experiments and through the application to EHR-based phenotyping prediction on Rheumatoid Arthritis (RA) using data from Partners Healthcare System.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge