Suqi Liu
Graph Attention Network for Node Regression on Random Geometric Graphs with Erdős--Rényi contamination
Jan 30, 2026Abstract:Graph attention networks (GATs) are widely used and often appear robust to noise in node covariates and edges, yet rigorous statistical guarantees demonstrating a provable advantage of GATs over non-attention graph neural networks~(GNNs) are scarce. We partially address this gap for node regression with graph-based errors-in-variables models under simultaneous covariate and edge corruption: responses are generated from latent node-level covariates, but only noise-perturbed versions of the latent covariates are observed; and the sample graph is a random geometric graph created from the node covariates but contaminated by independent Erdős--Rényi edges. We propose and analyze a carefully designed, task-specific GAT that constructs denoised proxy features for regression. We prove that regressing the response variables on the proxies achieves lower error asymptotically in (a) estimating the regression coefficient compared to the ordinary least squares (OLS) estimator on the noisy node covariates, and (b) predicting the response for an unlabelled node compared to a vanilla graph convolutional network~(GCN) -- under mild growth conditions. Our analysis leverages high-dimensional geometric tail bounds and concentration for neighbourhood counts and sample covariances. We verify our theoretical findings through experiments on synthetically generated data. We also perform experiments on real-world graphs and demonstrate the effectiveness of the attention mechanism in several node regression tasks.
Representation Learning to Advance Multi-institutional Studies with Electronic Health Record Data
Feb 12, 2025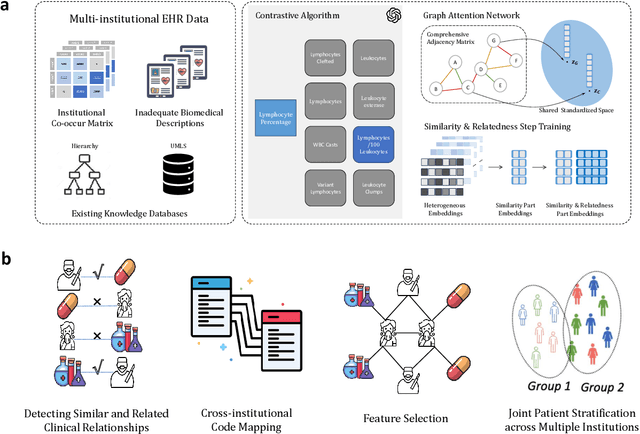
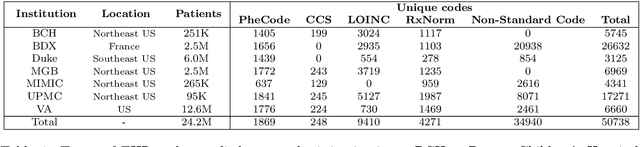
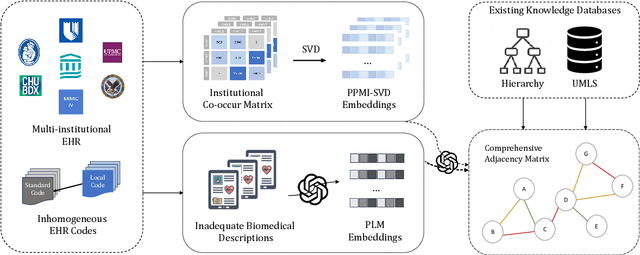
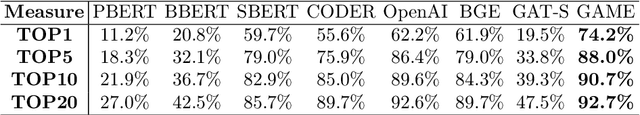
Abstract:The adoption of EHRs has expanded opportunities to leverage data-driven algorithms in clinical care and research. A major bottleneck in effectively conducting multi-institutional EHR studies is the data heterogeneity across systems with numerous codes that either do not exist or represent different clinical concepts across institutions. The need for data privacy further limits the feasibility of including multi-institutional patient-level data required to study similarities and differences across patient subgroups. To address these challenges, we developed the GAME algorithm. Tested and validated across 7 institutions and 2 languages, GAME integrates data in several levels: (1) at the institutional level with knowledge graphs to establish relationships between codes and existing knowledge sources, providing the medical context for standard codes and their relationship to each other; (2) between institutions, leveraging language models to determine the relationships between institution-specific codes with established standard codes; and (3) quantifying the strength of the relationships between codes using a graph attention network. Jointly trained embeddings are created using transfer and federated learning to preserve data privacy. In this study, we demonstrate the applicability of GAME in selecting relevant features as inputs for AI-driven algorithms in a range of conditions, e.g., heart failure, rheumatoid arthritis. We then highlight the application of GAME harmonized multi-institutional EHR data in a study of Alzheimer's disease outcomes and suicide risk among patients with mental health disorders, without sharing patient-level data outside individual institutions.
Unified Representation of Genomic and Biomedical Concepts through Multi-Task, Multi-Source Contrastive Learning
Oct 14, 2024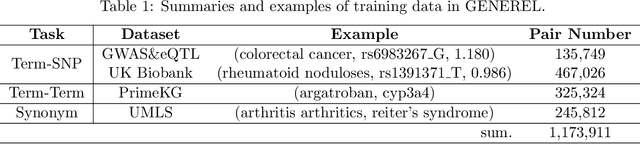
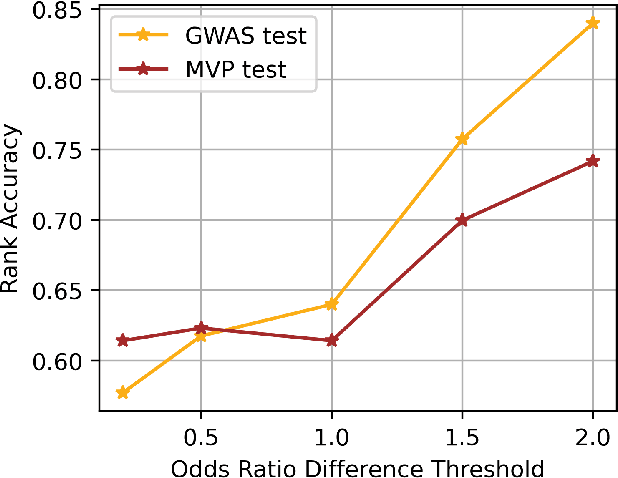
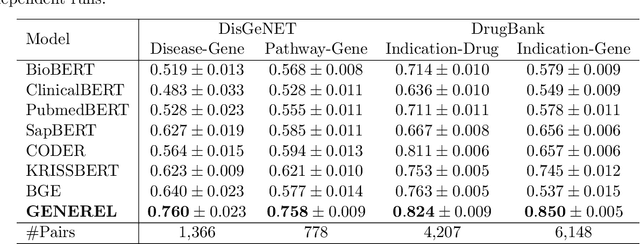
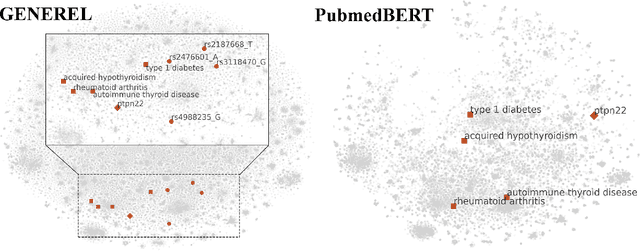
Abstract:We introduce GENomic Encoding REpresentation with Language Model (GENEREL), a framework designed to bridge genetic and biomedical knowledge bases. What sets GENEREL apart is its ability to fine-tune language models to infuse biological knowledge behind clinical concepts such as diseases and medications. This fine-tuning enables the model to capture complex biomedical relationships more effectively, enriching the understanding of how genomic data connects to clinical outcomes. By constructing a unified embedding space for biomedical concepts and a wide range of common SNPs from sources such as patient-level data, biomedical knowledge graphs, and GWAS summaries, GENEREL aligns the embeddings of SNPs and clinical concepts through multi-task contrastive learning. This allows the model to adapt to diverse natural language representations of biomedical concepts while bypassing the limitations of traditional code mapping systems across different data sources. Our experiments demonstrate GENEREL's ability to effectively capture the nuanced relationships between SNPs and clinical concepts. GENEREL also emerges to discern the degree of relatedness, potentially allowing for a more refined identification of concepts. This pioneering approach in constructing a unified embedding system for both SNPs and biomedical concepts enhances the potential for data integration and discovery in biomedical research.
Representation-Enhanced Neural Knowledge Integration with Application to Large-Scale Medical Ontology Learning
Oct 09, 2024Abstract:A large-scale knowledge graph enhances reproducibility in biomedical data discovery by providing a standardized, integrated framework that ensures consistent interpretation across diverse datasets. It improves generalizability by connecting data from various sources, enabling broader applicability of findings across different populations and conditions. Generating reliable knowledge graph, leveraging multi-source information from existing literature, however, is challenging especially with a large number of node sizes and heterogeneous relations. In this paper, we propose a general theoretically guaranteed statistical framework, called RENKI, to enable simultaneous learning of multiple relation types. RENKI generalizes various network models widely used in statistics and computer science. The proposed framework incorporates representation learning output into initial entity embedding of a neural network that approximates the score function for the knowledge graph and continuously trains the model to fit observed facts. We prove nonasymptotic bounds for in-sample and out-of-sample weighted MSEs in relation to the pseudo-dimension of the knowledge graph function class. Additionally, we provide pseudo-dimensions for score functions based on multilayer neural networks with ReLU activation function, in the scenarios when the embedding parameters either fixed or trainable. Finally, we complement our theoretical results with numerical studies and apply the method to learn a comprehensive medical knowledge graph combining a pretrained language model representation with knowledge graph links observed in several medical ontologies. The experiments justify our theoretical findings and demonstrate the effect of weighting in the presence of heterogeneous relations and the benefit of incorporating representation learning in nonparametric models.
Random Geometric Graph Alignment with Graph Neural Networks
Feb 12, 2024Abstract:We characterize the performance of graph neural networks for graph alignment problems in the presence of vertex feature information. More specifically, given two graphs that are independent perturbations of a single random geometric graph with noisy sparse features, the task is to recover an unknown one-to-one mapping between the vertices of the two graphs. We show under certain conditions on the sparsity and noise level of the feature vectors, a carefully designed one-layer graph neural network can with high probability recover the correct alignment between the vertices with the help of the graph structure. We also prove that our conditions on the noise level are tight up to logarithmic factors. Finally we compare the performance of the graph neural network to directly solving an assignment problem on the noisy vertex features. We demonstrate that when the noise level is at least constant this direct matching fails to have perfect recovery while the graph neural network can tolerate noise level growing as fast as a power of the size of the graph.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge