Thomas Booth
Beyond one-hot encoding? Journey into compact encoding for large multi-class segmentation
Oct 01, 2025Abstract:This work presents novel methods to reduce computational and memory requirements for medical image segmentation with a large number of classes. We curiously observe challenges in maintaining state-of-the-art segmentation performance with all of the explored options. Standard learning-based methods typically employ one-hot encoding of class labels. The computational complexity and memory requirements thus increase linearly with the number of classes. We propose a family of binary encoding approaches instead of one-hot encoding to reduce the computational complexity and memory requirements to logarithmic in the number of classes. In addition to vanilla binary encoding, we investigate the effects of error-correcting output codes (ECOCs), class weighting, hard/soft decoding, class-to-codeword assignment, and label embedding trees. We apply the methods to the use case of whole brain parcellation with 108 classes based on 3D MRI images. While binary encodings have proven efficient in so-called extreme classification problems in computer vision, we faced challenges in reaching state-of-the-art segmentation quality with binary encodings. Compared to one-hot encoding (Dice Similarity Coefficient (DSC) = 82.4 (2.8)), we report reduced segmentation performance with the binary segmentation approaches, achieving DSCs in the range from 39.3 to 73.8. Informative negative results all too often go unpublished. We hope that this work inspires future research of compact encoding strategies for large multi-class segmentation tasks.
Learning-Based Autonomous Navigation, Benchmark Environments and Simulation Framework for Endovascular Interventions
Oct 02, 2024



Abstract:Endovascular interventions are a life-saving treatment for many diseases, yet suffer from drawbacks such as radiation exposure and potential scarcity of proficient physicians. Robotic assistance during these interventions could be a promising support towards these problems. Research focusing on autonomous endovascular interventions utilizing artificial intelligence-based methodologies is gaining popularity. However, variability in assessment environments hinders the ability to compare and contrast the efficacy of different approaches, primarily due to each study employing a unique evaluation framework. In this study, we present deep reinforcement learning-based autonomous endovascular device navigation on three distinct digital benchmark interventions: BasicWireNav, ArchVariety, and DualDeviceNav. The benchmark interventions were implemented with our modular simulation framework stEVE (simulated EndoVascular Environment). Autonomous controllers were trained solely in simulation and evaluated in simulation and on physical test benches with camera and fluoroscopy feedback. Autonomous control for BasicWireNav and ArchVariety reached high success rates and was successfully transferred from the simulated training environment to the physical test benches, while autonomous control for DualDeviceNav reached a moderate success rate. The experiments demonstrate the feasibility of stEVE and its potential for transferring controllers trained in simulation to real-world scenarios. Nevertheless, they also reveal areas that offer opportunities for future research. This study demonstrates the transferability of autonomous controllers from simulation to the real world in endovascular navigation and lowers the entry barriers and increases the comparability of research on endovascular assistance systems by providing open-source training scripts, benchmarks and the stEVE framework.
Machine Learning and Glioblastoma: Treatment Response Monitoring Biomarkers in 2021
Apr 15, 2021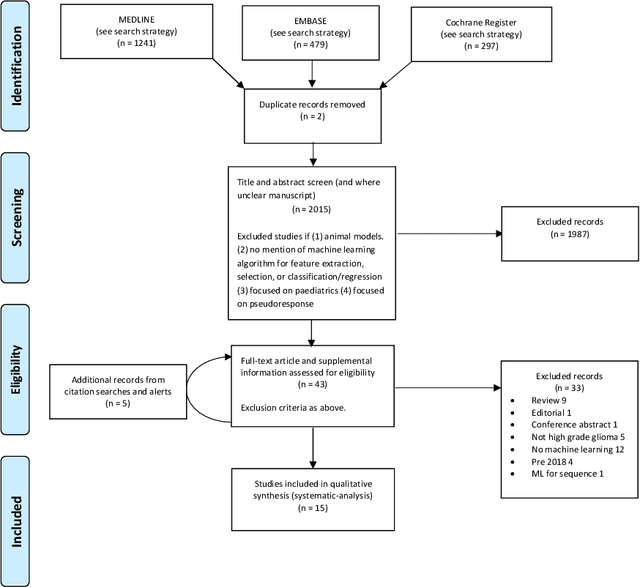
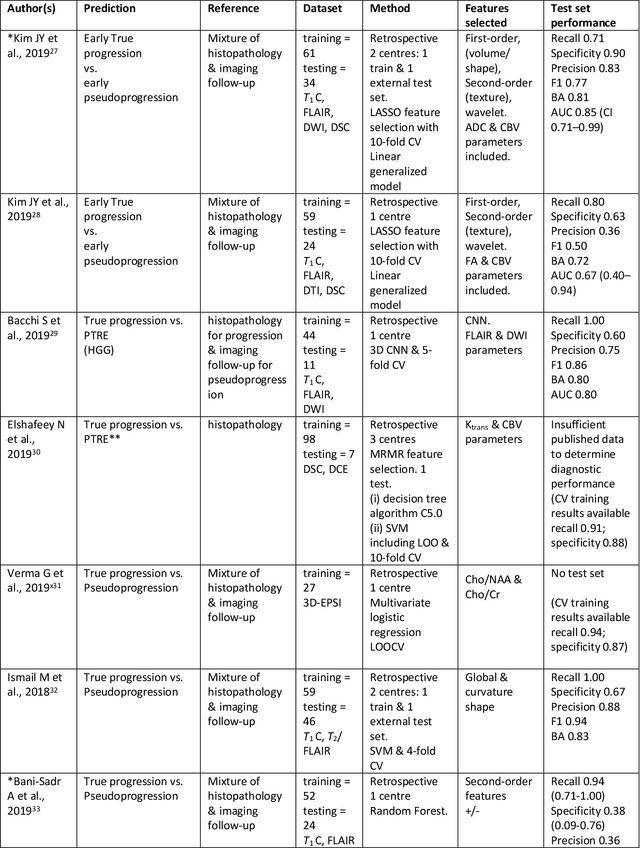
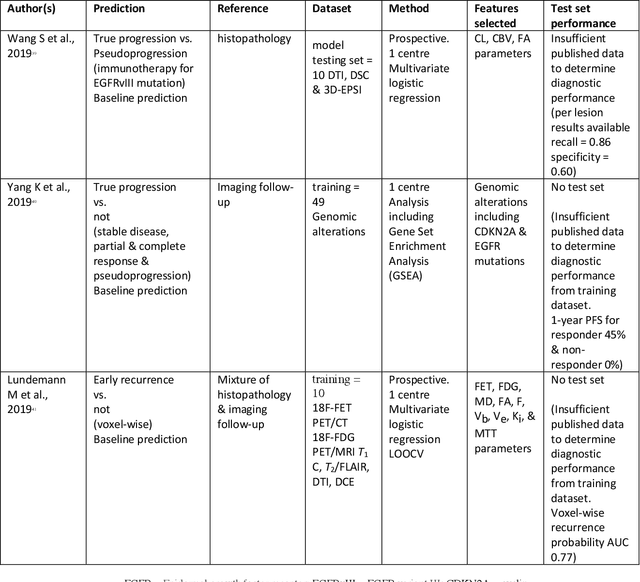
Abstract:The aim of the systematic review was to assess recently published studies on diagnostic test accuracy of glioblastoma treatment response monitoring biomarkers in adults, developed through machine learning (ML). Articles were searched for using MEDLINE, EMBASE, and the Cochrane Register. Included study participants were adult patients with high grade glioma who had undergone standard treatment (maximal resection, radiotherapy with concomitant and adjuvant temozolomide) and subsequently underwent follow-up imaging to determine treatment response status. Risk of bias and applicability was assessed with QUADAS 2 methodology. Contingency tables were created for hold-out test sets and recall, specificity, precision, F1-score, balanced accuracy calculated. Fifteen studies were included with 1038 patients in training sets and 233 in test sets. To determine whether there was progression or a mimic, the reference standard combination of follow-up imaging and histopathology at re-operation was applied in 67% of studies. The small numbers of patient included in studies, the high risk of bias and concerns of applicability in the study designs (particularly in relation to the reference standard and patient selection due to confounding), and the low level of evidence, suggest that limited conclusions can be drawn from the data. There is likely good diagnostic performance of machine learning models that use MRI features to distinguish between progression and mimics. The diagnostic performance of ML using implicit features did not appear to be superior to ML using explicit features. There are a range of ML-based solutions poised to become treatment response monitoring biomarkers for glioblastoma. To achieve this, the development and validation of ML models require large, well-annotated datasets where the potential for confounding in the study design has been carefully considered.
Learning joint segmentation of tissues and brain lesions from task-specific hetero-modal domain-shifted datasets
Sep 08, 2020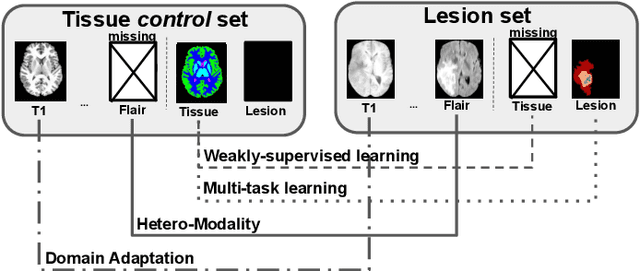

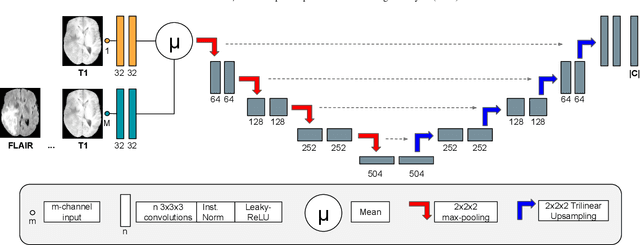

Abstract:Brain tissue segmentation from multimodal MRI is a key building block of many neuroimaging analysis pipelines. Established tissue segmentation approaches have, however, not been developed to cope with large anatomical changes resulting from pathology, such as white matter lesions or tumours, and often fail in these cases. In the meantime, with the advent of deep neural networks (DNNs), segmentation of brain lesions has matured significantly. However, few existing approaches allow for the joint segmentation of normal tissue and brain lesions. Developing a DNN for such a joint task is currently hampered by the fact that annotated datasets typically address only one specific task and rely on task-specific imaging protocols including a task-specific set of imaging modalities. In this work, we propose a novel approach to build a joint tissue and lesion segmentation model from aggregated task-specific hetero-modal domain-shifted and partially-annotated datasets. Starting from a variational formulation of the joint problem, we show how the expected risk can be decomposed and optimised empirically. We exploit an upper bound of the risk to deal with heterogeneous imaging modalities across datasets. To deal with potential domain shift, we integrated and tested three conventional techniques based on data augmentation, adversarial learning and pseudo-healthy generation. For each individual task, our joint approach reaches comparable performance to task-specific and fully-supervised models. The proposed framework is assessed on two different types of brain lesions: White matter lesions and gliomas. In the latter case, lacking a joint ground-truth for quantitative assessment purposes, we propose and use a novel clinically-relevant qualitative assessment methodology.
NEURO-DRAM: a 3D recurrent visual attention model for interpretable neuroimaging classification
Oct 18, 2019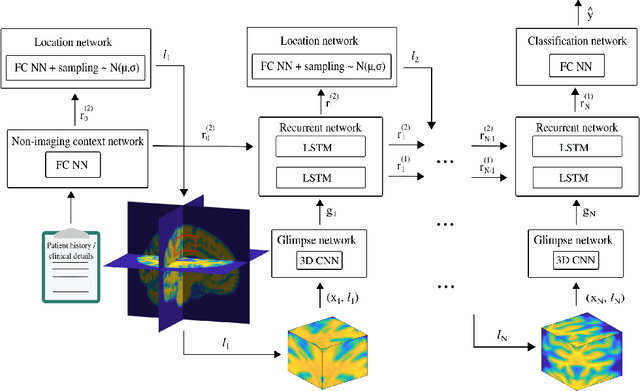
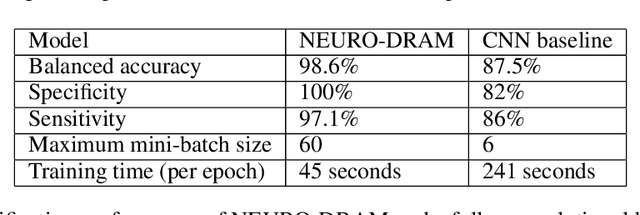

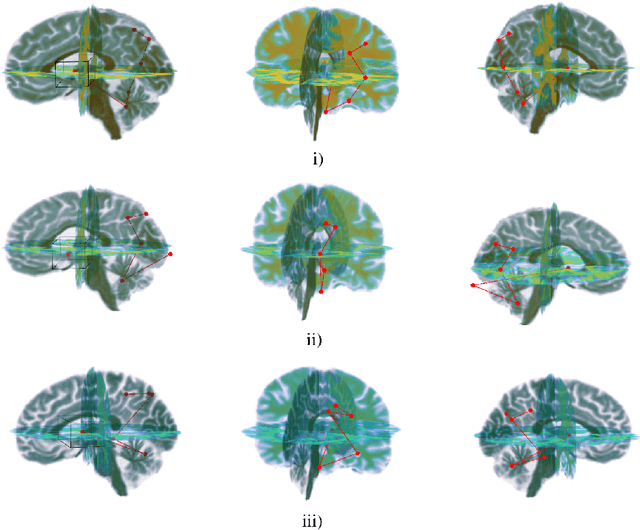
Abstract:Deep learning is attracting significant interest in the neuroimaging community as a means to diagnose psychiatric and neurological disorders from structural magnetic resonance images. However, there is a tendency amongst researchers to adopt architectures optimized for traditional computer vision tasks, rather than design networks customized for neuroimaging data. We address this by introducing NEURO-DRAM, a 3D recurrent visual attention model tailored for neuroimaging classification. The model comprises an agent which, trained by reinforcement learning, learns to navigate through volumetric images, selectively attending to the most informative regions for a given task. When applied to Alzheimer's disease prediction, NEURODRAM achieves state-of-the-art classification accuracy on an out-of-sample dataset, significantly outperforming a baseline convolutional neural network. When further applied to the task of predicting which patients with mild cognitive impairment will be diagnosed with Alzheimer's disease within two years, the model achieves state-of-the-art accuracy with no additional training. Encouragingly, the agent learns, without explicit instruction, a search policy in agreement with standardized radiological hallmarks of Alzheimer's disease, suggesting a route to automated biomarker discovery for more poorly understood disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge