Taliah Muhammad
Department of Neuroscience, Baylor College of Medicine, Houston, USA, Center for Neuroscience and Artificial Intelligence, Baylor College of Medicine, Houston, USA
The Sensorium competition on predicting large-scale mouse primary visual cortex activity
Jun 17, 2022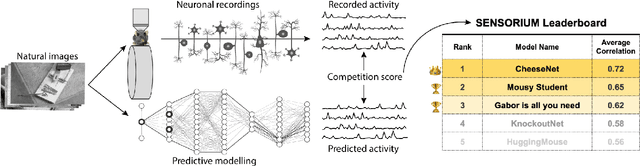
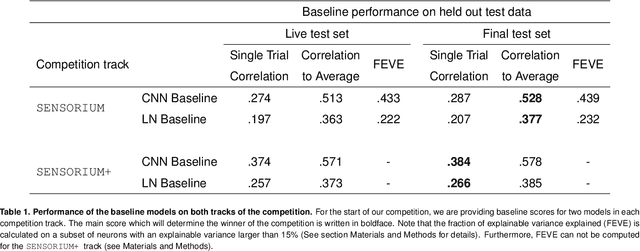
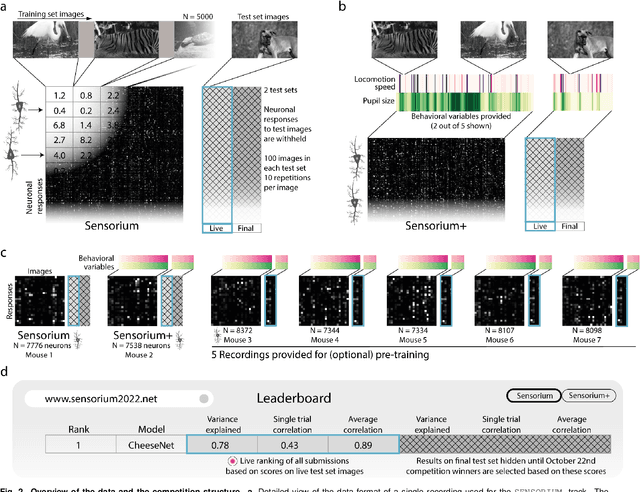
Abstract:The neural underpinning of the biological visual system is challenging to study experimentally, in particular as the neuronal activity becomes increasingly nonlinear with respect to visual input. Artificial neural networks (ANNs) can serve a variety of goals for improving our understanding of this complex system, not only serving as predictive digital twins of sensory cortex for novel hypothesis generation in silico, but also incorporating bio-inspired architectural motifs to progressively bridge the gap between biological and machine vision. The mouse has recently emerged as a popular model system to study visual information processing, but no standardized large-scale benchmark to identify state-of-the-art models of the mouse visual system has been established. To fill this gap, we propose the Sensorium benchmark competition. We collected a large-scale dataset from mouse primary visual cortex containing the responses of more than 28,000 neurons across seven mice stimulated with thousands of natural images, together with simultaneous behavioral measurements that include running speed, pupil dilation, and eye movements. The benchmark challenge will rank models based on predictive performance for neuronal responses on a held-out test set, and includes two tracks for model input limited to either stimulus only (Sensorium) or stimulus plus behavior (Sensorium+). We provide a starting kit to lower the barrier for entry, including tutorials, pre-trained baseline models, and APIs with one line commands for data loading and submission. We would like to see this as a starting point for regular challenges and data releases, and as a standard tool for measuring progress in large-scale neural system identification models of the mouse visual system and beyond.
Learning From Brains How to Regularize Machines
Nov 11, 2019
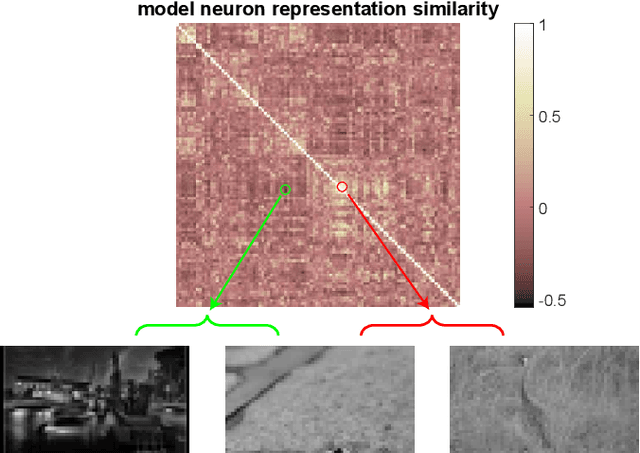


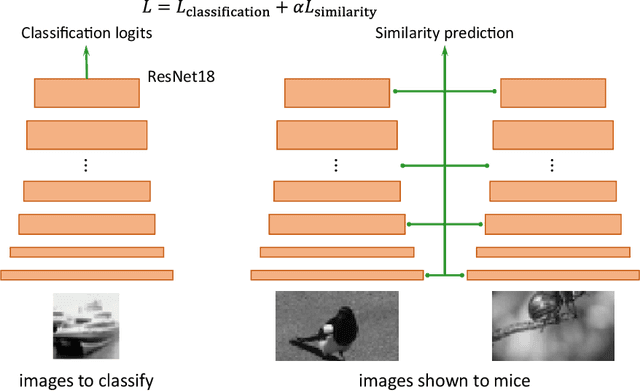
Abstract:Despite impressive performance on numerous visual tasks, Convolutional Neural Networks (CNNs) --- unlike brains --- are often highly sensitive to small perturbations of their input, e.g. adversarial noise leading to erroneous decisions. We propose to regularize CNNs using large-scale neuroscience data to learn more robust neural features in terms of representational similarity. We presented natural images to mice and measured the responses of thousands of neurons from cortical visual areas. Next, we denoised the notoriously variable neural activity using strong predictive models trained on this large corpus of responses from the mouse visual system, and calculated the representational similarity for millions of pairs of images from the model's predictions. We then used the neural representation similarity to regularize CNNs trained on image classification by penalizing intermediate representations that deviated from neural ones. This preserved performance of baseline models when classifying images under standard benchmarks, while maintaining substantially higher performance compared to baseline or control models when classifying noisy images. Moreover, the models regularized with cortical representations also improved model robustness in terms of adversarial attacks. This demonstrates that regularizing with neural data can be an effective tool to create an inductive bias towards more robust inference.
Penalized matrix decomposition for denoising, compression, and improved demixing of functional imaging data
Jul 17, 2018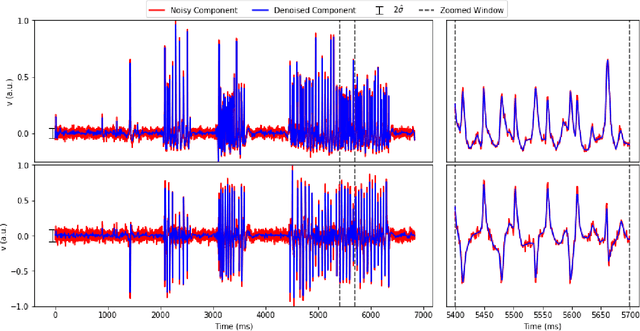
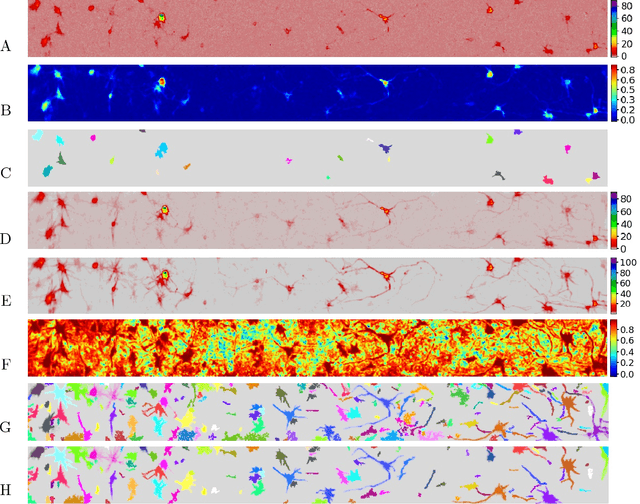
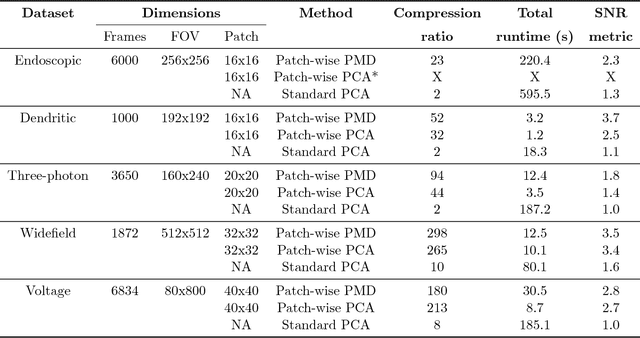

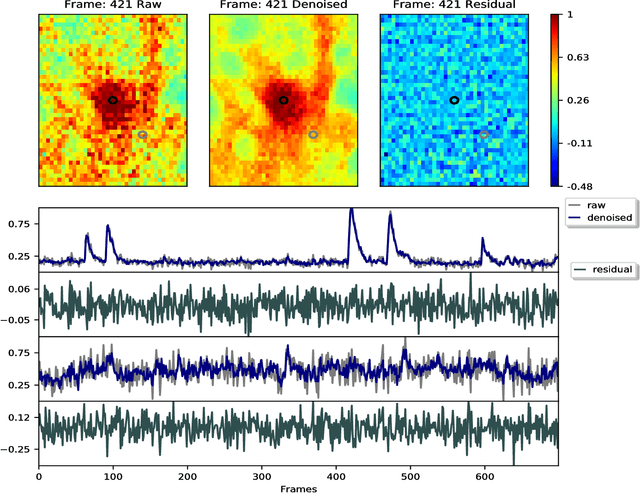
Abstract:Calcium imaging has revolutionized systems neuroscience, providing the ability to image large neural populations with single-cell resolution. The resulting datasets are quite large, which has presented a barrier to routine open sharing of this data, slowing progress in reproducible research. State of the art methods for analyzing this data are based on non-negative matrix factorization (NMF); these approaches solve a non-convex optimization problem, and are effective when good initializations are available, but can break down in low-SNR settings where common initialization approaches fail. Here we introduce an approach to compressing and denoising functional imaging data. The method is based on a spatially-localized penalized matrix decomposition (PMD) of the data to separate (low-dimensional) signal from (temporally-uncorrelated) noise. This approach can be applied in parallel on local spatial patches and is therefore highly scalable, does not impose non-negativity constraints or require stringent identifiability assumptions (leading to significantly more robust results compared to NMF), and estimates all parameters directly from the data, so no hand-tuning is required. We have applied the method to a wide range of functional imaging data (including one-photon, two-photon, three-photon, widefield, somatic, axonal, dendritic, calcium, and voltage imaging datasets): in all cases, we observe ~2-4x increases in SNR and compression rates of 20-300x with minimal visible loss of signal, with no adjustment of hyperparameters; this in turn facilitates the process of demixing the observed activity into contributions from individual neurons. We focus on two challenging applications: dendritic calcium imaging data and voltage imaging data in the context of optogenetic stimulation. In both cases, we show that our new approach leads to faster and much more robust extraction of activity from the data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge