Laura Pede
Institute of Computer Science and Campus Institute Data Science, University of Goettingen, Germany
The Sensorium competition on predicting large-scale mouse primary visual cortex activity
Jun 17, 2022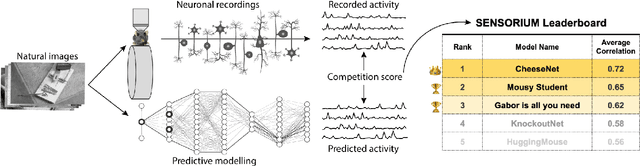
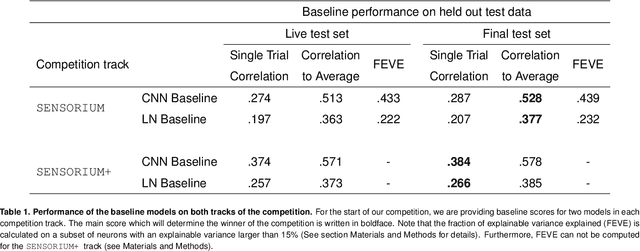
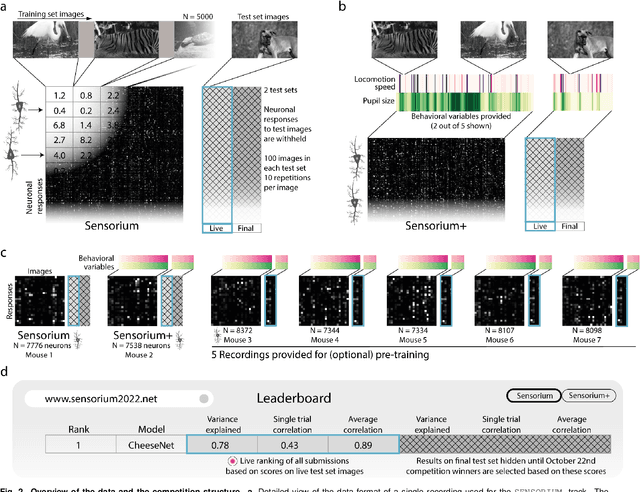
Abstract:The neural underpinning of the biological visual system is challenging to study experimentally, in particular as the neuronal activity becomes increasingly nonlinear with respect to visual input. Artificial neural networks (ANNs) can serve a variety of goals for improving our understanding of this complex system, not only serving as predictive digital twins of sensory cortex for novel hypothesis generation in silico, but also incorporating bio-inspired architectural motifs to progressively bridge the gap between biological and machine vision. The mouse has recently emerged as a popular model system to study visual information processing, but no standardized large-scale benchmark to identify state-of-the-art models of the mouse visual system has been established. To fill this gap, we propose the Sensorium benchmark competition. We collected a large-scale dataset from mouse primary visual cortex containing the responses of more than 28,000 neurons across seven mice stimulated with thousands of natural images, together with simultaneous behavioral measurements that include running speed, pupil dilation, and eye movements. The benchmark challenge will rank models based on predictive performance for neuronal responses on a held-out test set, and includes two tracks for model input limited to either stimulus only (Sensorium) or stimulus plus behavior (Sensorium+). We provide a starting kit to lower the barrier for entry, including tutorials, pre-trained baseline models, and APIs with one line commands for data loading and submission. We would like to see this as a starting point for regular challenges and data releases, and as a standard tool for measuring progress in large-scale neural system identification models of the mouse visual system and beyond.
Self-supervised Representation Learning of Neuronal Morphologies
Jan 18, 2022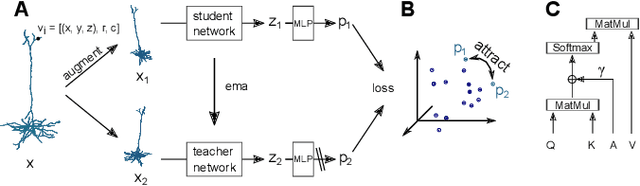
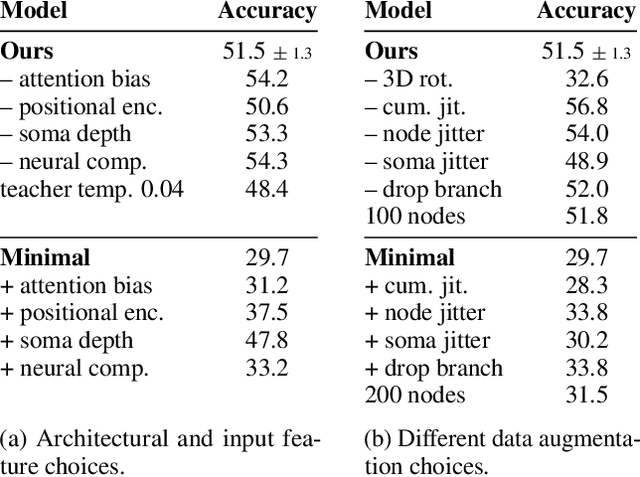
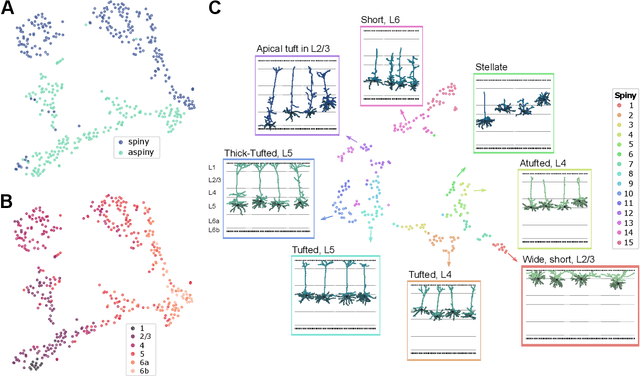
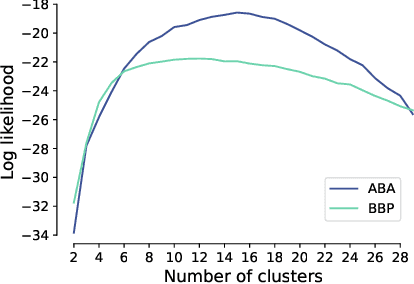
Abstract:Understanding the diversity of cell types and their function in the brain is one of the key challenges in neuroscience. The advent of large-scale datasets has given rise to the need of unbiased and quantitative approaches to cell type classification. We present GraphDINO, a purely data-driven approach to learning a low dimensional representation of the 3D morphology of neurons. GraphDINO is a novel graph representation learning method for spatial graphs utilizing self-supervised learning on transformer models. It combines attention-based global interaction between nodes and classic graph convolutional processing. We show, in two different species and cortical areas, that this method is able to yield morphological cell type clustering that is comparable to manual feature-based classification and shows a good correspondence to expert-labeled cell types. Our method is applicable beyond neuroscience in settings where samples in a dataset are graphs and graph-level embeddings are desired.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge