Sophie Chiron
Cross-scale Multi-instance Learning for Pathological Image Diagnosis
Apr 01, 2023



Abstract:Analyzing high resolution whole slide images (WSIs) with regard to information across multiple scales poses a significant challenge in digital pathology. Multi-instance learning (MIL) is a common solution for working with high resolution images by classifying bags of objects (i.e. sets of smaller image patches). However, such processing is typically performed at a single scale (e.g., 20x magnification) of WSIs, disregarding the vital inter-scale information that is key to diagnoses by human pathologists. In this study, we propose a novel cross-scale MIL algorithm to explicitly aggregate inter-scale relationships into a single MIL network for pathological image diagnosis. The contribution of this paper is three-fold: (1) A novel cross-scale MIL (CS-MIL) algorithm that integrates the multi-scale information and the inter-scale relationships is proposed; (2) A toy dataset with scale-specific morphological features is created and released to examine and visualize differential cross-scale attention; (3) Superior performance on both in-house and public datasets is demonstrated by our simple cross-scale MIL strategy. The official implementation is publicly available at https://github.com/hrlblab/CS-MIL.
Cross-scale Attention Guided Multi-instance Learning for Crohn's Disease Diagnosis with Pathological Images
Aug 15, 2022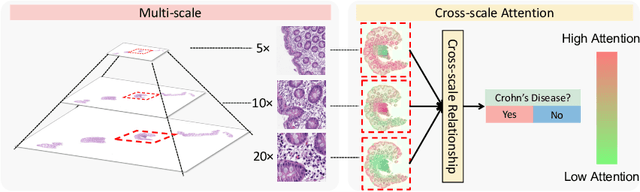
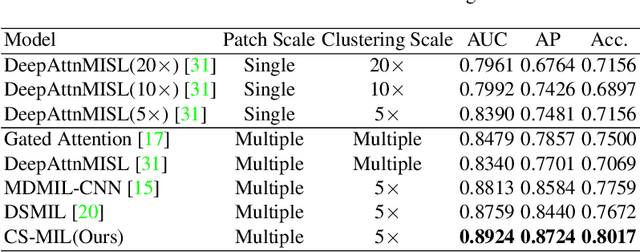
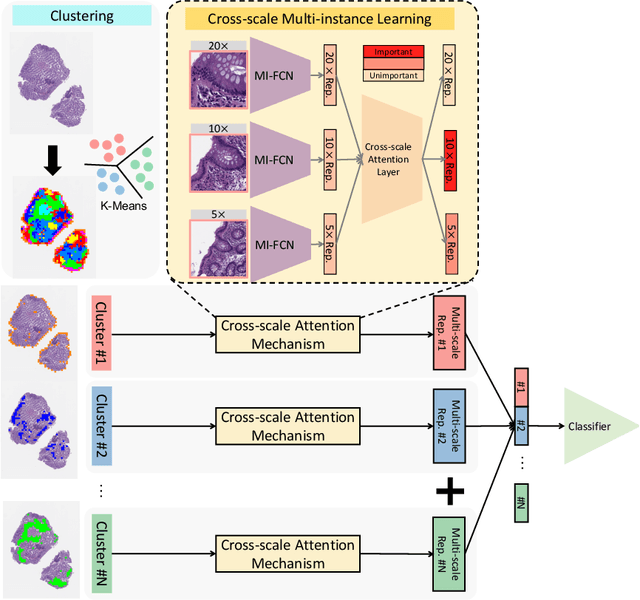
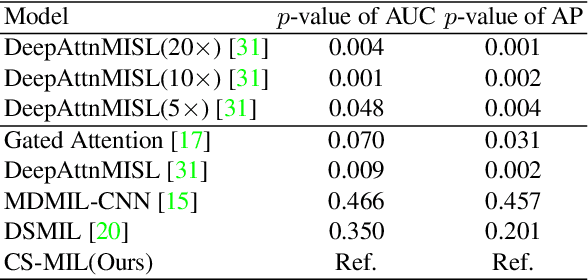
Abstract:Multi-instance learning (MIL) is widely used in the computer-aided interpretation of pathological Whole Slide Images (WSIs) to solve the lack of pixel-wise or patch-wise annotations. Often, this approach directly applies "natural image driven" MIL algorithms which overlook the multi-scale (i.e. pyramidal) nature of WSIs. Off-the-shelf MIL algorithms are typically deployed on a single-scale of WSIs (e.g., 20x magnification), while human pathologists usually aggregate the global and local patterns in a multi-scale manner (e.g., by zooming in and out between different magnifications). In this study, we propose a novel cross-scale attention mechanism to explicitly aggregate inter-scale interactions into a single MIL network for Crohn's Disease (CD), which is a form of inflammatory bowel disease. The contribution of this paper is two-fold: (1) a cross-scale attention mechanism is proposed to aggregate features from different resolutions with multi-scale interaction; and (2) differential multi-scale attention visualizations are generated to localize explainable lesion patterns. By training ~250,000 H&E-stained Ascending Colon (AC) patches from 20 CD patient and 30 healthy control samples at different scales, our approach achieved a superior Area under the Curve (AUC) score of 0.8924 compared with baseline models. The official implementation is publicly available at https://github.com/hrlblab/CS-MIL.
Random Multi-Channel Image Synthesis for Multiplexed Immunofluorescence Imaging
Sep 18, 2021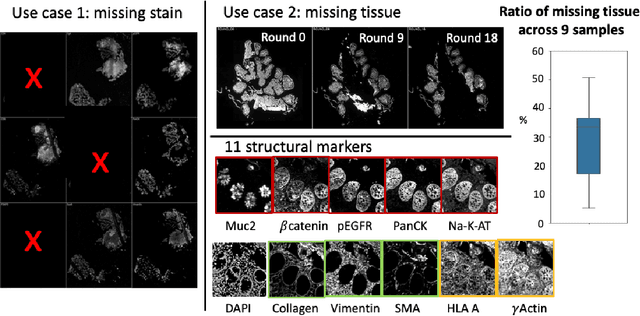
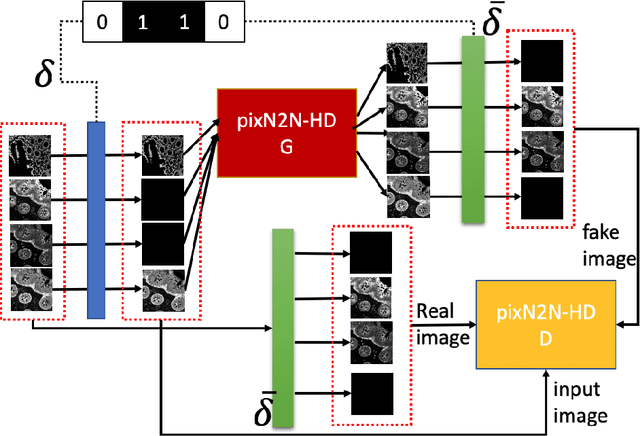
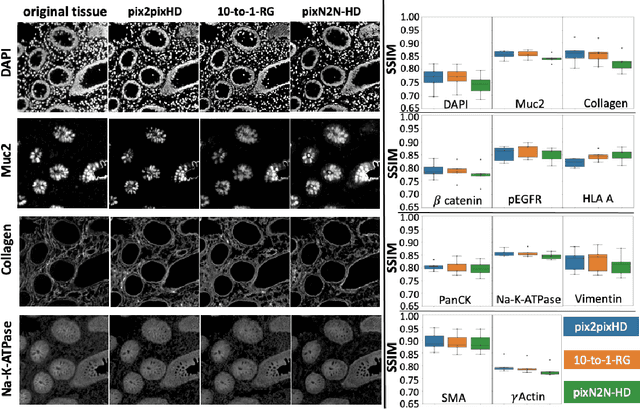
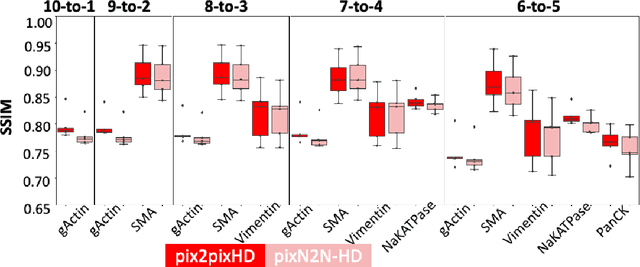
Abstract:Multiplex immunofluorescence (MxIF) is an emerging imaging technique that produces the high sensitivity and specificity of single-cell mapping. With a tenet of 'seeing is believing', MxIF enables iterative staining and imaging extensive antibodies, which provides comprehensive biomarkers to segment and group different cells on a single tissue section. However, considerable depletion of the scarce tissue is inevitable from extensive rounds of staining and bleaching ('missing tissue'). Moreover, the immunofluorescence (IF) imaging can globally fail for particular rounds ('missing stain''). In this work, we focus on the 'missing stain' issue. It would be appealing to develop digital image synthesis approaches to restore missing stain images without losing more tissue physically. Herein, we aim to develop image synthesis approaches for eleven MxIF structural molecular markers (i.e., epithelial and stromal) on real samples. We propose a novel multi-channel high-resolution image synthesis approach, called pixN2N-HD, to tackle possible missing stain scenarios via a high-resolution generative adversarial network (GAN). Our contribution is three-fold: (1) a single deep network framework is proposed to tackle missing stain in MxIF; (2) the proposed 'N-to-N' strategy reduces theoretical four years of computational time to 20 hours when covering all possible missing stains scenarios, with up to five missing stains (e.g., '(N-1)-to-1', '(N-2)-to-2'); and (3) this work is the first comprehensive experimental study of investigating cross-stain synthesis in MxIF. Our results elucidate a promising direction of advancing MxIF imaging with deep image synthesis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge