Reinhold Schmidt
Explainable concept mappings of MRI: Revealing the mechanisms underlying deep learning-based brain disease classification
Apr 16, 2024
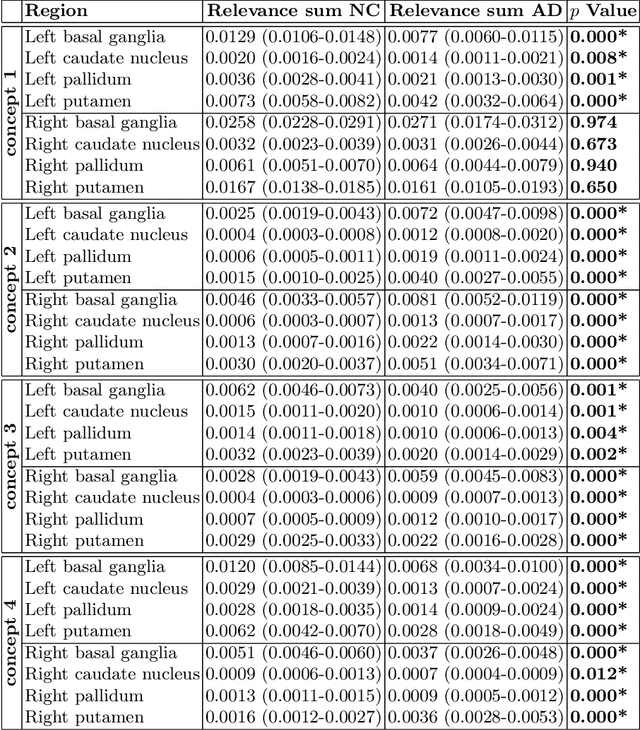
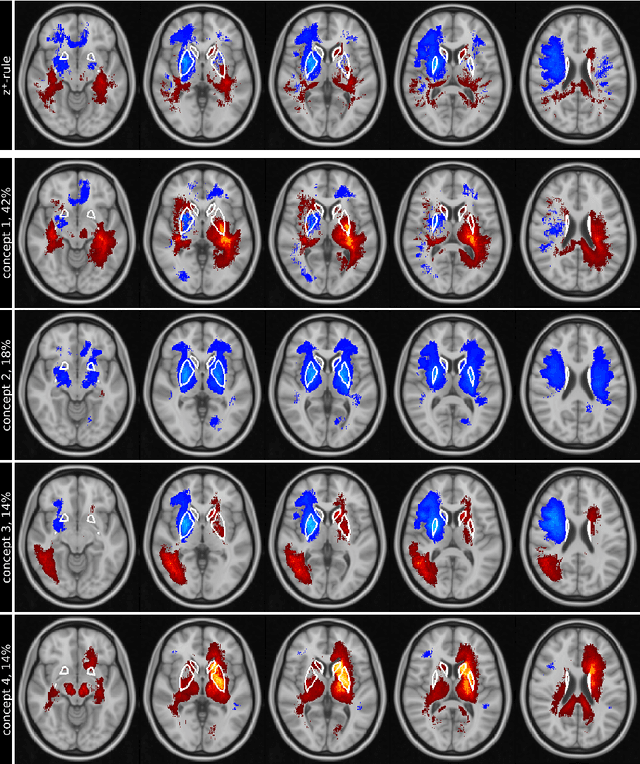
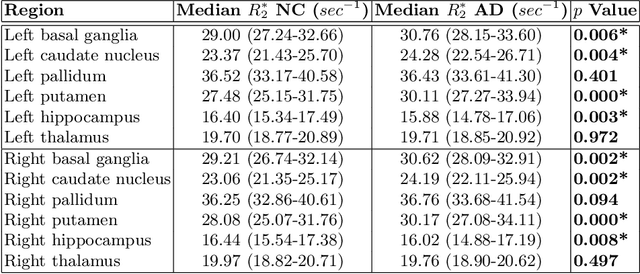
Abstract:Motivation. While recent studies show high accuracy in the classification of Alzheimer's disease using deep neural networks, the underlying learned concepts have not been investigated. Goals. To systematically identify changes in brain regions through concepts learned by the deep neural network for model validation. Approach. Using quantitative R2* maps we separated Alzheimer's patients (n=117) from normal controls (n=219) by using a convolutional neural network and systematically investigated the learned concepts using Concept Relevance Propagation and compared these results to a conventional region of interest-based analysis. Results. In line with established histological findings and the region of interest-based analyses, highly relevant concepts were primarily found in and adjacent to the basal ganglia. Impact. The identification of concepts learned by deep neural networks for disease classification enables validation of the models and could potentially improve reliability.
Analyzing hierarchical multi-view MRI data with StaPLR: An application to Alzheimer's disease classification
Aug 12, 2021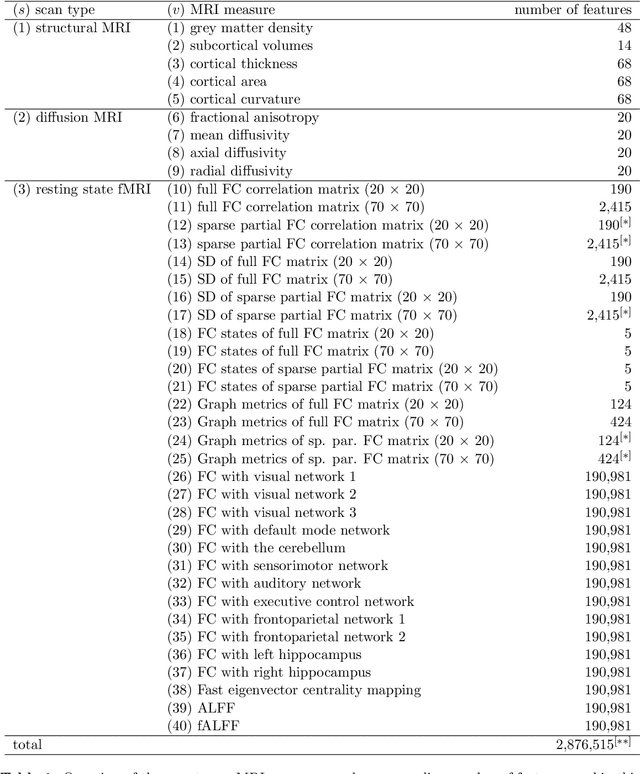
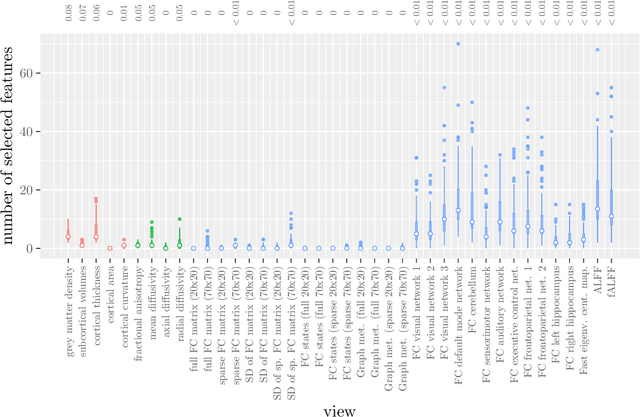
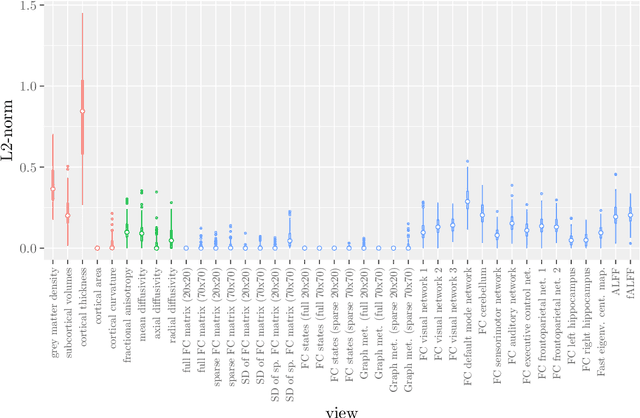
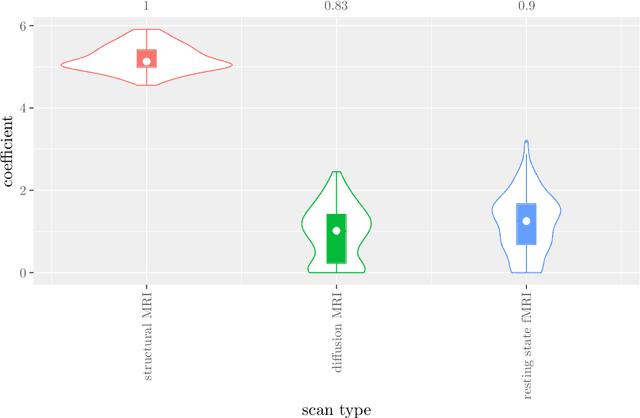
Abstract:Multi-view data refers to a setting where features are divided into feature sets, for example because they correspond to different sources. Stacked penalized logistic regression (StaPLR) is a recently introduced method that can be used for classification and automatically selecting the views that are most important for prediction. We show how this method can easily be extended to a setting where the data has a hierarchical multi-view structure. We apply StaPLR to Alzheimer's disease classification where different MRI measures have been calculated from three scan types: structural MRI, diffusion-weighted MRI, and resting-state fMRI. StaPLR can identify which scan types and which MRI measures are most important for classification, and it outperforms elastic net regression in classification performance.
Bayesian deep neural networks for low-cost neurophysiological markers of Alzheimer's disease severity
Dec 13, 2018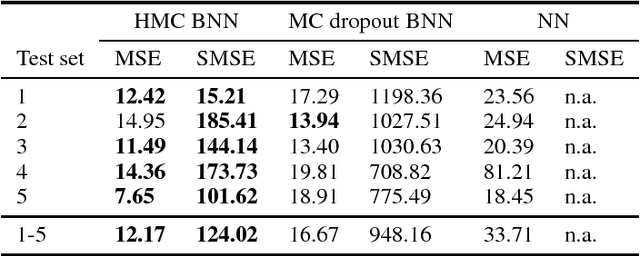
Abstract:As societies around the world are ageing, the number of Alzheimer's disease (AD) patients is rapidly increasing. To date, no low-cost, non-invasive biomarkers have been established to advance the objectivization of AD diagnosis and progression assessment. Here, we utilize Bayesian neural networks to develop a multivariate predictor for AD severity using a wide range of quantitative EEG (QEEG) markers. The Bayesian treatment of neural networks both automatically controls model complexity and provides a predictive distribution over the target function, giving uncertainty bounds for our regression task. It is therefore well suited to clinical neuroscience, where data sets are typically sparse and practitioners require a precise assessment of the predictive uncertainty. We use data of one of the largest prospective AD EEG trials ever conducted to demonstrate the potential of Bayesian deep learning in this domain, while comparing two distinct Bayesian neural network approaches, i.e., Monte Carlo dropout and Hamiltonian Monte Carlo.
Riemannian tangent space mapping and elastic net regularization for cost-effective EEG markers of brain atrophy in Alzheimer's disease
Nov 22, 2017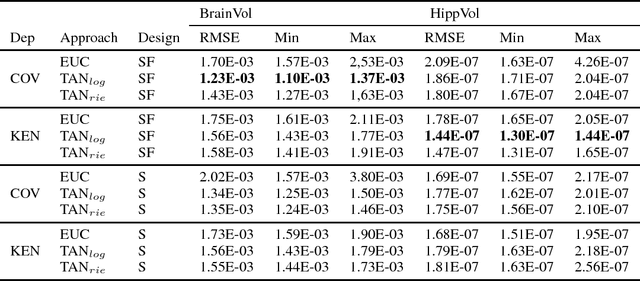
Abstract:The diagnosis of Alzheimer's disease (AD) in routine clinical practice is most commonly based on subjective clinical interpretations. Quantitative electroencephalography (QEEG) measures have been shown to reflect neurodegenerative processes in AD and might qualify as affordable and thereby widely available markers to facilitate the objectivization of AD assessment. Here, we present a novel framework combining Riemannian tangent space mapping and elastic net regression for the development of brain atrophy markers. While most AD QEEG studies are based on small sample sizes and psychological test scores as outcome measures, here we train and test our models using data of one of the largest prospective EEG AD trials ever conducted, including MRI biomarkers of brain atrophy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge