Naveen Valluri
Distilling Large Language Models for Biomedical Knowledge Extraction: A Case Study on Adverse Drug Events
Jul 12, 2023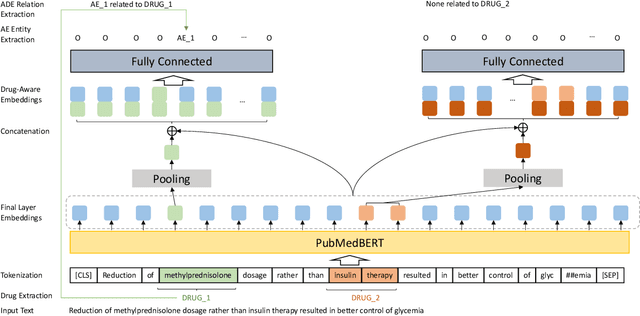
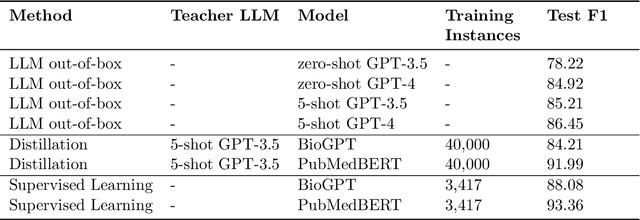
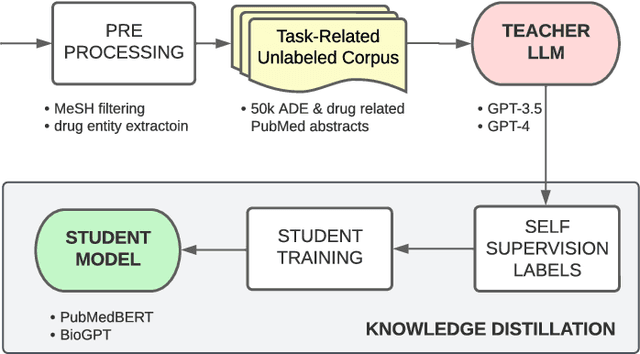
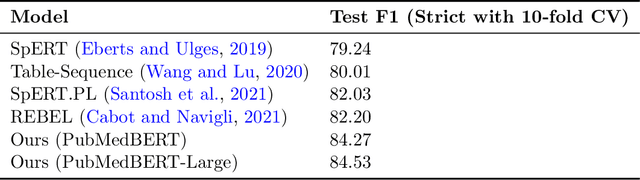
Abstract:Large language models (LLMs), such as GPT-4, have demonstrated remarkable capabilities across a wide range of tasks, including health applications. In this paper, we study how LLMs can be used to scale biomedical knowledge curation. We find that while LLMs already possess decent competency in structuring biomedical text, by distillation into a task-specific student model through self-supervised learning, substantial gains can be attained over out-of-box LLMs, with additional advantages such as cost, efficiency, and white-box model access. We conduct a case study on adverse drug event (ADE) extraction, which is an important area for improving care. On standard ADE extraction evaluation, a GPT-3.5 distilled PubMedBERT model attained comparable accuracy as supervised state-of-the-art models without using any labeled data. Despite being over 1,000 times smaller, the distilled model outperformed its teacher GPT-3.5 by over 6 absolute points in F1 and GPT-4 by over 5 absolute points. Ablation studies on distillation model choice (e.g., PubMedBERT vs BioGPT) and ADE extraction architecture shed light on best practice for biomedical knowledge extraction. Similar gains were attained by distillation for other standard biomedical knowledge extraction tasks such as gene-disease associations and protected health information, further illustrating the promise of this approach.
Large-Scale Domain-Specific Pretraining for Biomedical Vision-Language Processing
Mar 02, 2023Abstract:Contrastive pretraining on parallel image-text data has attained great success in vision-language processing (VLP), as exemplified by CLIP and related methods. However, prior explorations tend to focus on general domains in the web. Biomedical images and text are rather different, but publicly available datasets are small and skew toward chest X-ray, thus severely limiting progress. In this paper, we conducted by far the largest study on biomedical VLP, using 15 million figure-caption pairs extracted from biomedical research articles in PubMed Central. Our dataset (PMC-15M) is two orders of magnitude larger than existing biomedical image-text datasets such as MIMIC-CXR, and spans a diverse range of biomedical images. The standard CLIP method is suboptimal for the biomedical domain. We propose BiomedCLIP with domain-specific adaptations tailored to biomedical VLP. We conducted extensive experiments and ablation studies on standard biomedical imaging tasks from retrieval to classification to visual question-answering (VQA). BiomedCLIP established new state of the art in a wide range of standard datasets, substantially outperformed prior VLP approaches. Surprisingly, BiomedCLIP even outperformed radiology-specific state-of-the-art models such as BioViL on radiology-specific tasks such as RSNA pneumonia detection, thus highlighting the utility in large-scale pretraining across all biomedical image types. We will release our models at https://aka.ms/biomedclip to facilitate future research in biomedical VLP.
Towards Structuring Real-World Data at Scale: Deep Learning for Extracting Key Oncology Information from Clinical Text with Patient-Level Supervision
Mar 20, 2022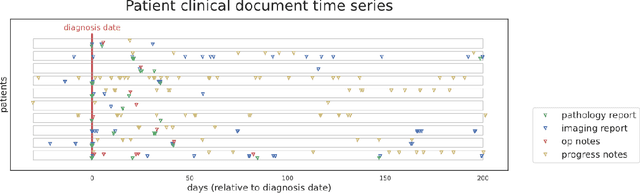
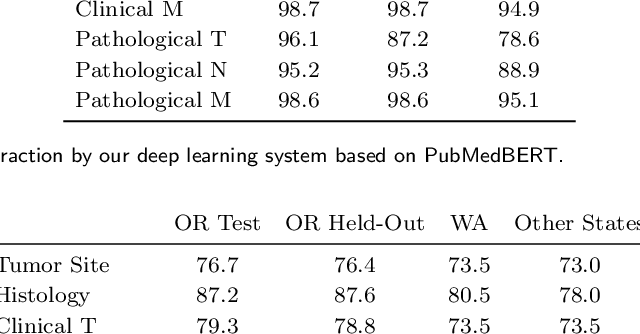
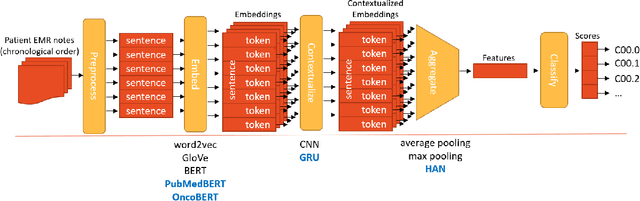
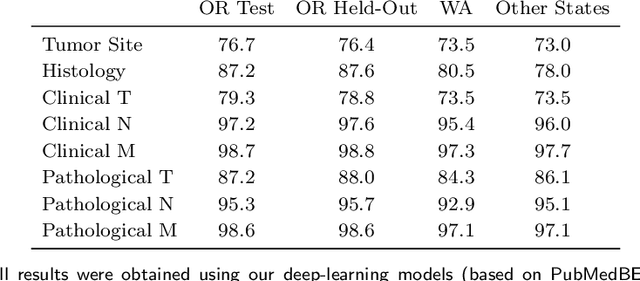
Abstract:Objective: The majority of detailed patient information in real-world data (RWD) is only consistently available in free-text clinical documents. Manual curation is expensive and time-consuming. Developing natural language processing (NLP) methods for structuring RWD is thus essential for scaling real-world evidence generation. Materials and Methods: Traditional rule-based systems are vulnerable to the prevalent linguistic variations and ambiguities in clinical text, and prior applications of machine-learning methods typically require sentence-level or report-level labeled examples that are hard to produce at scale. We propose leveraging patient-level supervision from medical registries, which are often readily available and capture key patient information, for general RWD applications. To combat the lack of sentence-level or report-level annotations, we explore advanced deep-learning methods by combining domain-specific pretraining, recurrent neural networks, and hierarchical attention. Results: We conduct an extensive study on 135,107 patients from the cancer registry of a large integrated delivery network (IDN) comprising healthcare systems in five western US states. Our deep learning methods attain test AUROC of 94-99% for key tumor attributes and comparable performance on held-out data from separate health systems and states. Discussion and Conclusion: Ablation results demonstrate clear superiority of these advanced deep-learning methods over prior approaches. Error analysis shows that our NLP system sometimes even corrects errors in registrar labels. We also conduct a preliminary investigation in accelerating registry curation and general RWD structuring via assisted curation for over 1.2 million cancer patients in this healthcare network.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge