Lindsay Edwards
No Foundations without Foundations -- Why semi-mechanistic models are essential for regulatory biology
Jan 31, 2025Abstract:Despite substantial efforts, deep learning has not yet delivered a transformative impact on elucidating regulatory biology, particularly in the realm of predicting gene expression profiles. Here, we argue that genuine "foundation models" of regulatory biology will remain out of reach unless guided by frameworks that integrate mechanistic insight with principled experimental design. We present one such ground-up, semi-mechanistic framework that unifies perturbation-based experimental designs across both in vitro and in vivo CRISPR screens, accounting for differentiating and non-differentiating cellular systems. By revealing previously unrecognised assumptions in published machine learning methods, our approach clarifies links with popular techniques such as variational autoencoders and structural causal models. In practice, this framework suggests a modified loss function that we demonstrate can improve predictive performance, and further suggests an error analysis that informs batching strategies. Ultimately, since cellular regulation emerges from innumerable interactions amongst largely uncharted molecular components, we contend that systems-level understanding cannot be achieved through structural biology alone. Instead, we argue that real progress will require a first-principles perspective on how experiments capture biological phenomena, how data are generated, and how these processes can be reflected in more faithful modelling architectures.
BioNeMo Framework: a modular, high-performance library for AI model development in drug discovery
Nov 15, 2024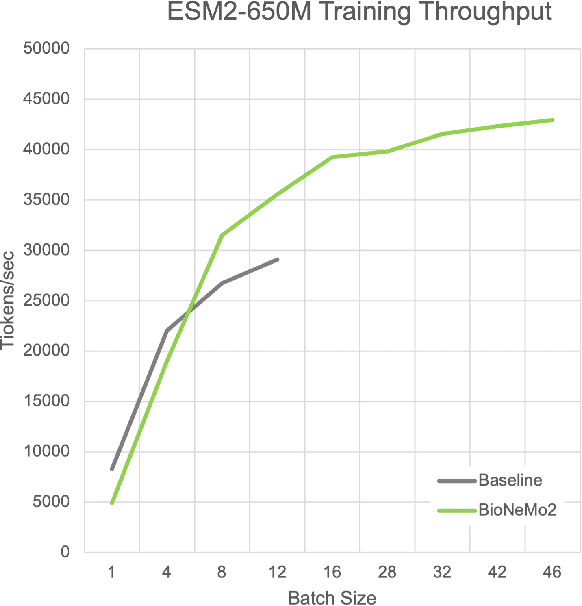
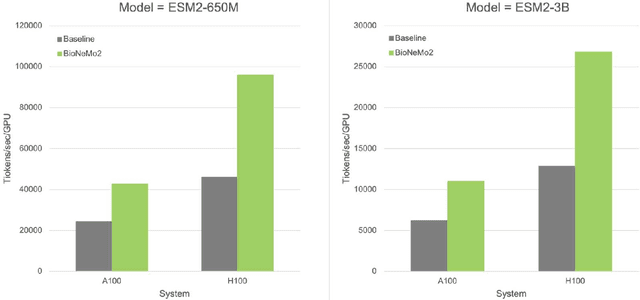
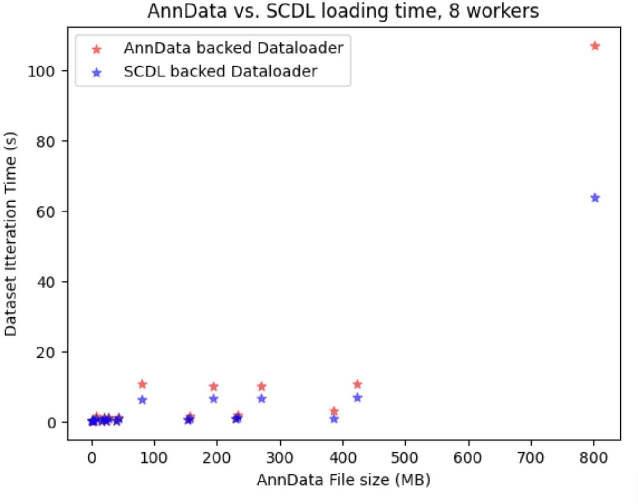
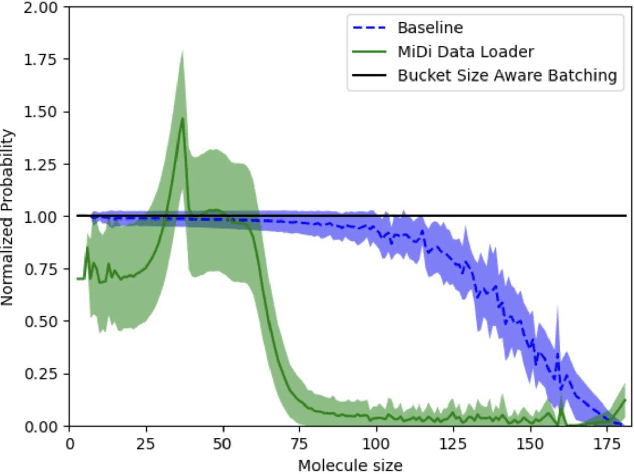
Abstract:Artificial Intelligence models encoding biology and chemistry are opening new routes to high-throughput and high-quality in-silico drug development. However, their training increasingly relies on computational scale, with recent protein language models (pLM) training on hundreds of graphical processing units (GPUs). We introduce the BioNeMo Framework to facilitate the training of computational biology and chemistry AI models across hundreds of GPUs. Its modular design allows the integration of individual components, such as data loaders, into existing workflows and is open to community contributions. We detail technical features of the BioNeMo Framework through use cases such as pLM pre-training and fine-tuning. On 256 NVIDIA A100s, BioNeMo Framework trains a three billion parameter BERT-based pLM on over one trillion tokens in 4.2 days. The BioNeMo Framework is open-source and free for everyone to use.
Season combinatorial intervention predictions with Salt & Peper
Apr 25, 2024
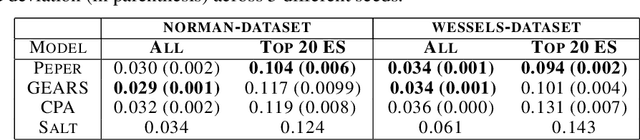
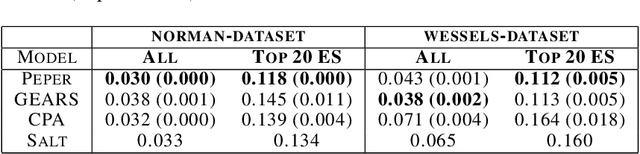
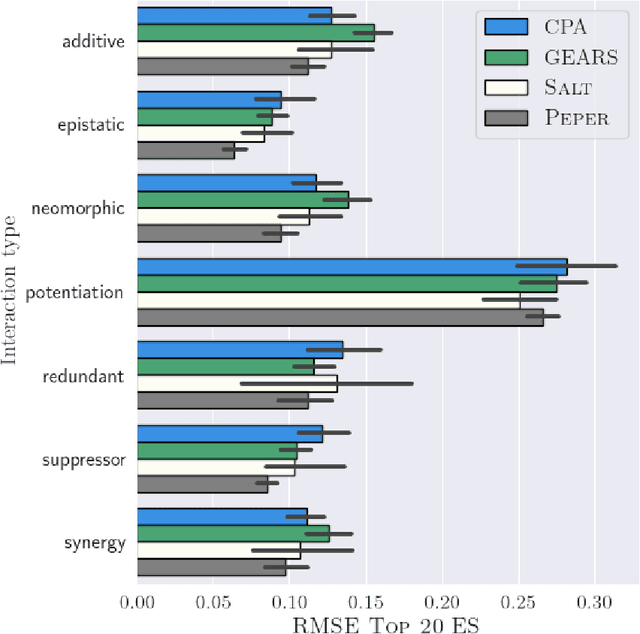
Abstract:Interventions play a pivotal role in the study of complex biological systems. In drug discovery, genetic interventions (such as CRISPR base editing) have become central to both identifying potential therapeutic targets and understanding a drug's mechanism of action. With the advancement of CRISPR and the proliferation of genome-scale analyses such as transcriptomics, a new challenge is to navigate the vast combinatorial space of concurrent genetic interventions. Addressing this, our work concentrates on estimating the effects of pairwise genetic combinations on the cellular transcriptome. We introduce two novel contributions: Salt, a biologically-inspired baseline that posits the mostly additive nature of combination effects, and Peper, a deep learning model that extends Salt's additive assumption to achieve unprecedented accuracy. Our comprehensive comparison against existing state-of-the-art methods, grounded in diverse metrics, and our out-of-distribution analysis highlight the limitations of current models in realistic settings. This analysis underscores the necessity for improved modelling techniques and data acquisition strategies, paving the way for more effective exploration of genetic intervention effects.
PyRelationAL: A Library for Active Learning Research and Development
May 23, 2022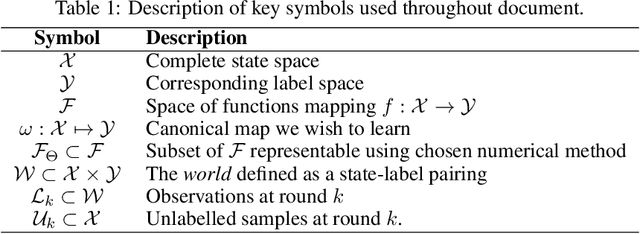
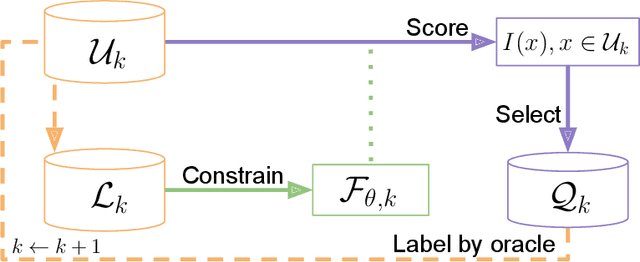
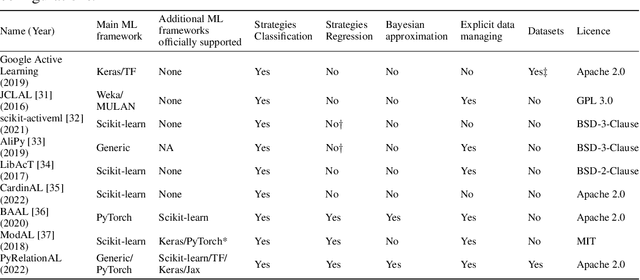
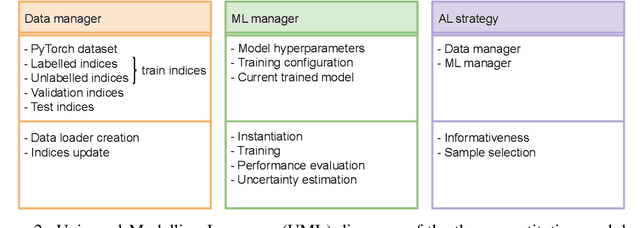
Abstract:In constrained real-world scenarios where it is challenging or costly to generate data, disciplined methods for acquiring informative new data points are of fundamental importance for the efficient training of machine learning (ML) models. Active learning (AL) is a subfield of ML focused on the development of methods to iteratively and economically acquire data through strategically querying new data points that are the most useful for a particular task. Here, we introduce PyRelationAL, an open source library for AL research. We describe a modular toolkit that is compatible with diverse ML frameworks (e.g. PyTorch, Scikit-Learn, TensorFlow, JAX). Furthermore, to help accelerate research and development in the field, the library implements a number of published methods and provides API access to wide-ranging benchmark datasets and AL task configurations based on existing literature. The library is supplemented by an expansive set of tutorials, demos, and documentation to help users get started. We perform experiments on the PyRelationAL collection of benchmark datasets and showcase the considerable economies that AL can provide. PyRelationAL is maintained using modern software engineering practices - with an inclusive contributor code of conduct - to promote long term library quality and utilisation.
SubTab: Subsetting Features of Tabular Data for Self-Supervised Representation Learning
Oct 27, 2021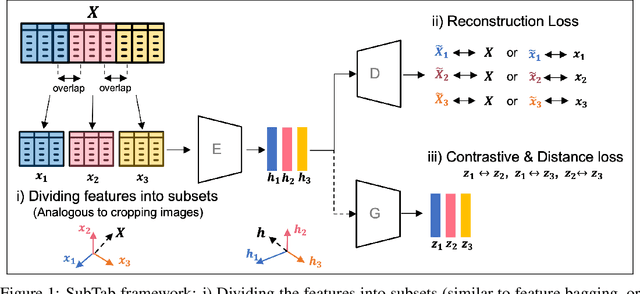
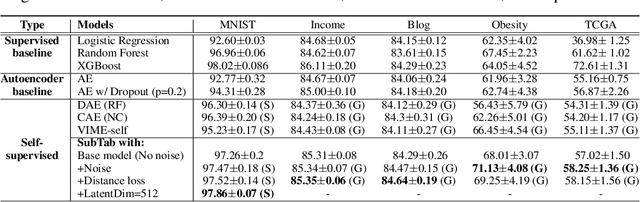
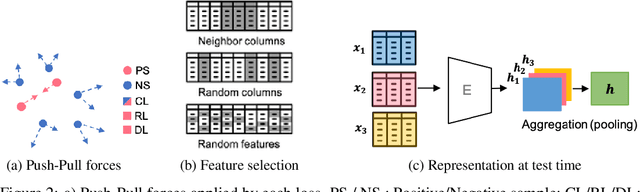
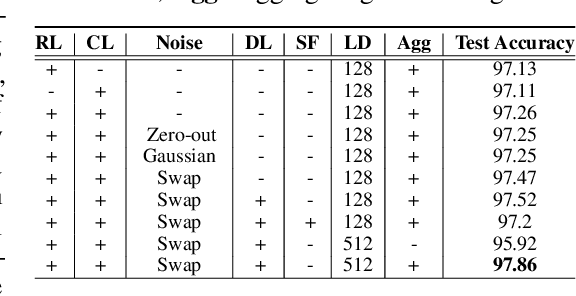
Abstract:Self-supervised learning has been shown to be very effective in learning useful representations, and yet much of the success is achieved in data types such as images, audio, and text. The success is mainly enabled by taking advantage of spatial, temporal, or semantic structure in the data through augmentation. However, such structure may not exist in tabular datasets commonly used in fields such as healthcare, making it difficult to design an effective augmentation method, and hindering a similar progress in tabular data setting. In this paper, we introduce a new framework, Subsetting features of Tabular data (SubTab), that turns the task of learning from tabular data into a multi-view representation learning problem by dividing the input features to multiple subsets. We argue that reconstructing the data from the subset of its features rather than its corrupted version in an autoencoder setting can better capture its underlying latent representation. In this framework, the joint representation can be expressed as the aggregate of latent variables of the subsets at test time, which we refer to as collaborative inference. Our experiments show that the SubTab achieves the state of the art (SOTA) performance of 98.31% on MNIST in tabular setting, on par with CNN-based SOTA models, and surpasses existing baselines on three other real-world datasets by a significant margin.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge