Leon Lenchik
Comp2Comp: Open-Source Software with FDA-Cleared Artificial Intelligence Algorithms for Computed Tomography Image Analysis
Feb 10, 2026Abstract:Artificial intelligence allows automatic extraction of imaging biomarkers from already-acquired radiologic images. This paradigm of opportunistic imaging adds value to medical imaging without additional imaging costs or patient radiation exposure. However, many open-source image analysis solutions lack rigorous validation while commercial solutions lack transparency, leading to unexpected failures when deployed. Here, we report development and validation for two of the first fully open-sourced, FDA-510(k)-cleared deep learning pipelines to mitigate both challenges: Abdominal Aortic Quantification (AAQ) and Bone Mineral Density (BMD) estimation are both offered within the Comp2Comp package for opportunistic analysis of computed tomography scans. AAQ segments the abdominal aorta to assess aneurysm size; BMD segments vertebral bodies to estimate trabecular bone density and osteoporosis risk. AAQ-derived maximal aortic diameters were compared against radiologist ground-truth measurements on 258 patient scans enriched for abdominal aortic aneurysms from four external institutions. BMD binary classifications (low vs. normal bone density) were compared against concurrent DXA scan ground truths obtained on 371 patient scans from four external institutions. AAQ had an overall mean absolute error of 1.57 mm (95% CI 1.38-1.80 mm). BMD had a sensitivity of 81.0% (95% CI 74.0-86.8%) and specificity of 78.4% (95% CI 72.3-83.7%). Comp2Comp AAQ and BMD demonstrated sufficient accuracy for clinical use. Open-sourcing these algorithms improves transparency of typically opaque FDA clearance processes, allows hospitals to test the algorithms before cumbersome clinical pilots, and provides researchers with best-in-class methods.
Comp2Comp: Open-Source Body Composition Assessment on Computed Tomography
Feb 13, 2023Abstract:Computed tomography (CT) is routinely used in clinical practice to evaluate a wide variety of medical conditions. While CT scans provide diagnoses, they also offer the ability to extract quantitative body composition metrics to analyze tissue volume and quality. Extracting quantitative body composition measures manually from CT scans is a cumbersome and time-consuming task. Proprietary software has been developed recently to automate this process, but the closed-source nature impedes widespread use. There is a growing need for fully automated body composition software that is more accessible and easier to use, especially for clinicians and researchers who are not experts in medical image processing. To this end, we have built Comp2Comp, an open-source Python package for rapid and automated body composition analysis of CT scans. This package offers models, post-processing heuristics, body composition metrics, automated batching, and polychromatic visualizations. Comp2Comp currently computes body composition measures for bone, skeletal muscle, visceral adipose tissue, and subcutaneous adipose tissue on CT scans of the abdomen. We have created two pipelines for this purpose. The first pipeline computes vertebral measures, as well as muscle and adipose tissue measures, at the T12 - L5 vertebral levels from abdominal CT scans. The second pipeline computes muscle and adipose tissue measures on user-specified 2D axial slices. In this guide, we discuss the architecture of the Comp2Comp pipelines, provide usage instructions, and report internal and external validation results to measure the quality of segmentations and body composition measures. Comp2Comp can be found at https://github.com/StanfordMIMI/Comp2Comp.
Data-Limited Tissue Segmentation using Inpainting-Based Self-Supervised Learning
Oct 14, 2022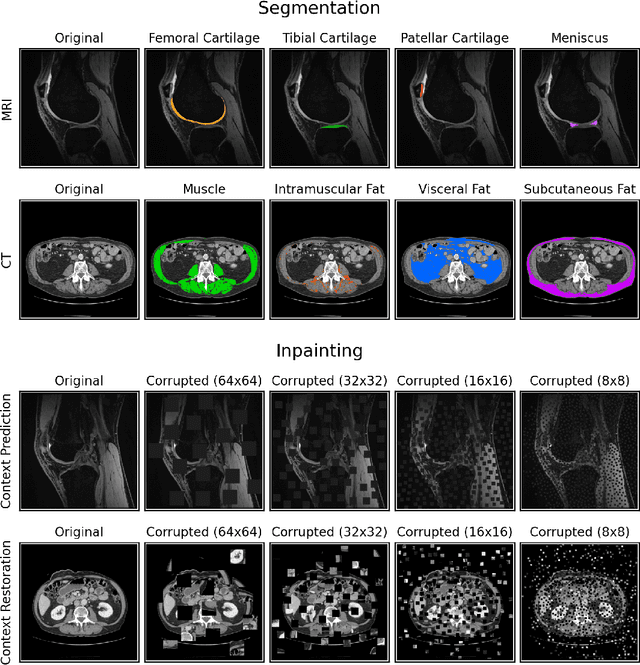
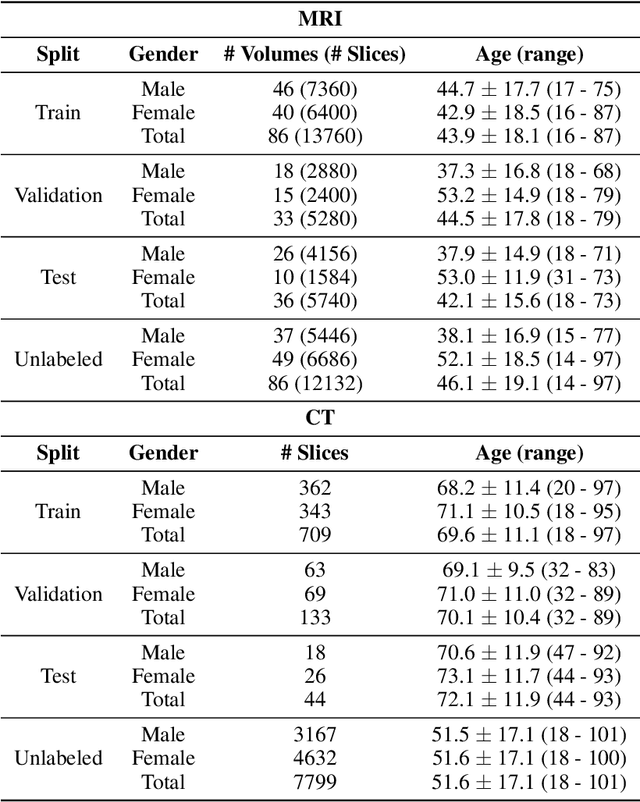
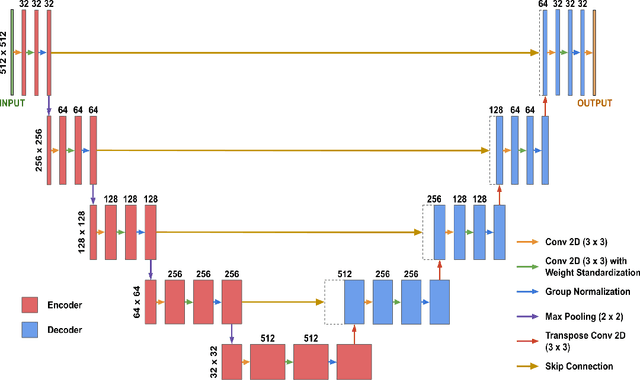
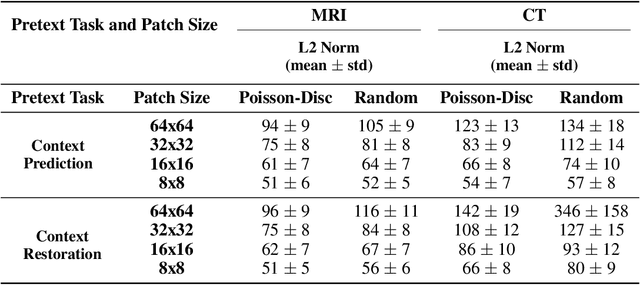
Abstract:Although supervised learning has enabled high performance for image segmentation, it requires a large amount of labeled training data, which can be difficult to obtain in the medical imaging field. Self-supervised learning (SSL) methods involving pretext tasks have shown promise in overcoming this requirement by first pretraining models using unlabeled data. In this work, we evaluate the efficacy of two SSL methods (inpainting-based pretext tasks of context prediction and context restoration) for CT and MRI image segmentation in label-limited scenarios, and investigate the effect of implementation design choices for SSL on downstream segmentation performance. We demonstrate that optimally trained and easy-to-implement inpainting-based SSL segmentation models can outperform classically supervised methods for MRI and CT tissue segmentation in label-limited scenarios, for both clinically-relevant metrics and the traditional Dice score.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge