Lei Du
Experimental robustness benchmark of quantum neural network on a superconducting quantum processor
May 22, 2025Abstract:Quantum machine learning (QML) models, like their classical counterparts, are vulnerable to adversarial attacks, hindering their secure deployment. Here, we report the first systematic experimental robustness benchmark for 20-qubit quantum neural network (QNN) classifiers executed on a superconducting processor. Our benchmarking framework features an efficient adversarial attack algorithm designed for QNNs, enabling quantitative characterization of adversarial robustness and robustness bounds. From our analysis, we verify that adversarial training reduces sensitivity to targeted perturbations by regularizing input gradients, significantly enhancing QNN's robustness. Additionally, our analysis reveals that QNNs exhibit superior adversarial robustness compared to classical neural networks, an advantage attributed to inherent quantum noise. Furthermore, the empirical upper bound extracted from our attack experiments shows a minimal deviation ($3 \times 10^{-3}$) from the theoretical lower bound, providing strong experimental confirmation of the attack's effectiveness and the tightness of fidelity-based robustness bounds. This work establishes a critical experimental framework for assessing and improving quantum adversarial robustness, paving the way for secure and reliable QML applications.
Chat2Brain: A Method for Mapping Open-Ended Semantic Queries to Brain Activation Maps
Sep 10, 2023Abstract:Over decades, neuroscience has accumulated a wealth of research results in the text modality that can be used to explore cognitive processes. Meta-analysis is a typical method that successfully establishes a link from text queries to brain activation maps using these research results, but it still relies on an ideal query environment. In practical applications, text queries used for meta-analyses may encounter issues such as semantic redundancy and ambiguity, resulting in an inaccurate mapping to brain images. On the other hand, large language models (LLMs) like ChatGPT have shown great potential in tasks such as context understanding and reasoning, displaying a high degree of consistency with human natural language. Hence, LLMs could improve the connection between text modality and neuroscience, resolving existing challenges of meta-analyses. In this study, we propose a method called Chat2Brain that combines LLMs to basic text-2-image model, known as Text2Brain, to map open-ended semantic queries to brain activation maps in data-scarce and complex query environments. By utilizing the understanding and reasoning capabilities of LLMs, the performance of the mapping model is optimized by transferring text queries to semantic queries. We demonstrate that Chat2Brain can synthesize anatomically plausible neural activation patterns for more complex tasks of text queries.
Knowledge-aware Attention Network for Protein-Protein Interaction Extraction
Jan 07, 2020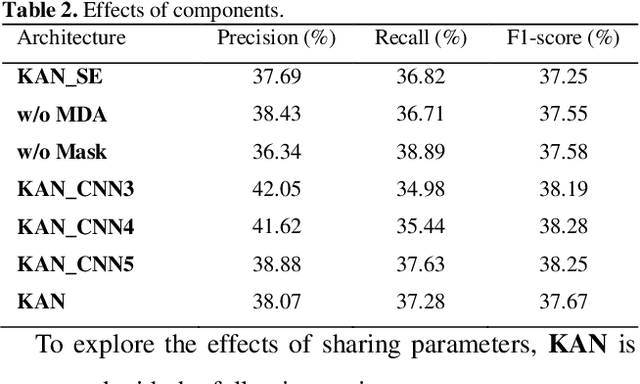
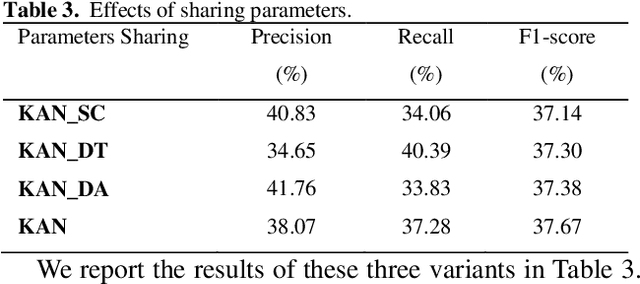
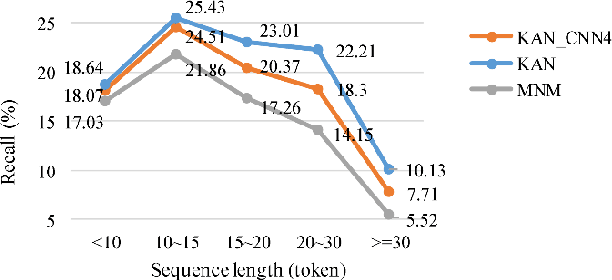
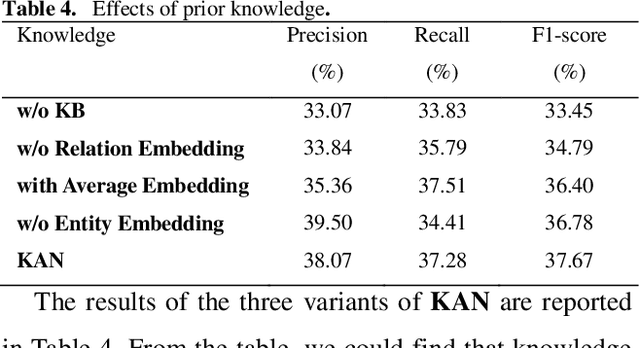
Abstract:Protein-protein interaction (PPI) extraction from published scientific literature provides additional support for precision medicine efforts. However, many of the current PPI extraction methods need extensive feature engineering and cannot make full use of the prior knowledge in knowledge bases (KB). KBs contain huge amounts of structured information about entities and relationships, therefore plays a pivotal role in PPI extraction. This paper proposes a knowledge-aware attention network (KAN) to fuse prior knowledge about protein-protein pairs and context information for PPI extraction. The proposed model first adopts a diagonal-disabled multi-head attention mechanism to encode context sequence along with knowledge representations learned from KB. Then a novel multi-dimensional attention mechanism is used to select the features that can best describe the encoded context. Experiment results on the BioCreative VI PPI dataset show that the proposed approach could acquire knowledge-aware dependencies between different words in a sequence and lead to a new state-of-the-art performance.
* Published on Journal of Biomedical Informatics, 14 pages, 5 figures
Knowledge-guided Convolutional Networks for Chemical-Disease Relation Extraction
Dec 23, 2019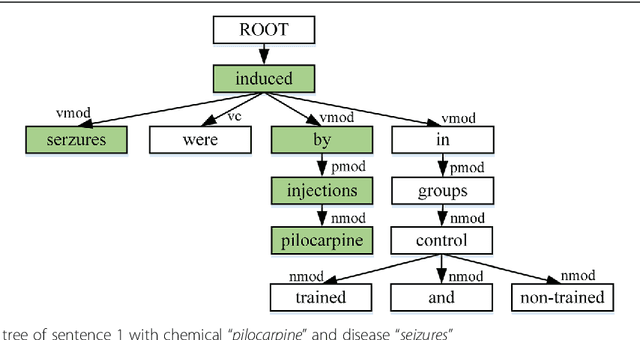
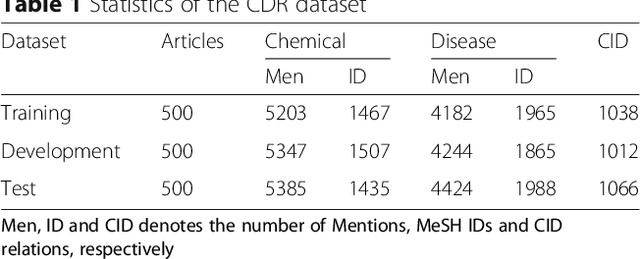
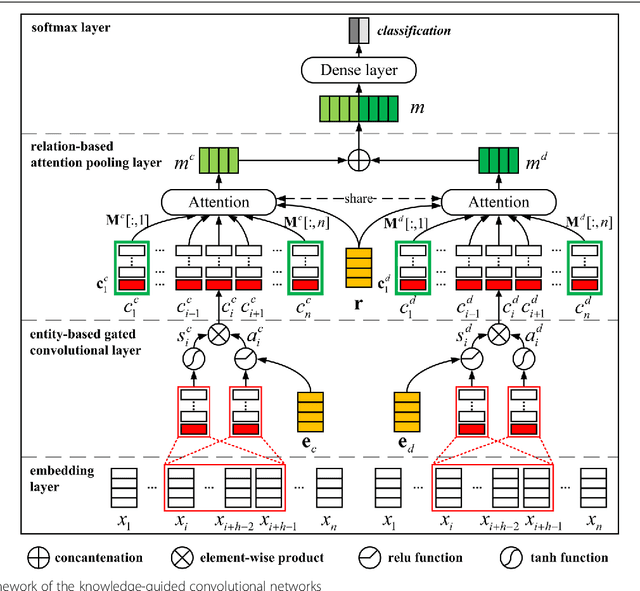
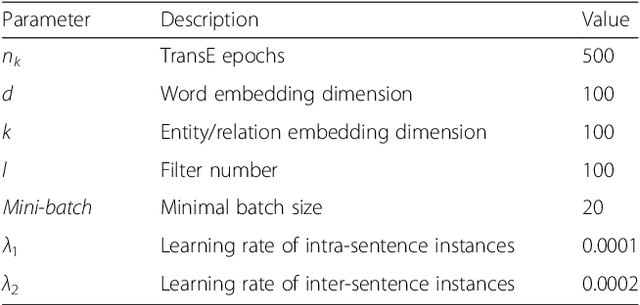
Abstract:Background: Automatic extraction of chemical-disease relations (CDR) from unstructured text is of essential importance for disease treatment and drug development. Meanwhile, biomedical experts have built many highly-structured knowledge bases (KBs), which contain prior knowledge about chemicals and diseases. Prior knowledge provides strong support for CDR extraction. How to make full use of it is worth studying. Results: This paper proposes a novel model called "Knowledge-guided Convolutional Networks (KCN)" to leverage prior knowledge for CDR extraction. The proposed model first learns knowledge representations including entity embeddings and relation embeddings from KBs. Then, entity embeddings are used to control the propagation of context features towards a chemical-disease pair with gated convolutions. After that, relation embeddings are employed to further capture the weighted context features by a shared attention pooling. Finally, the weighted context features containing additional knowledge information are used for CDR extraction. Experiments on the BioCreative V CDR dataset show that the proposed KCN achieves 71.28% F1-score, which outperforms most of the state-of-the-art systems. Conclusions: This paper proposes a novel CDR extraction model KCN to make full use of prior knowledge. Experimental results demonstrate that KCN could effectively integrate prior knowledge and contexts for the performance improvement.
* Published on BMC Bioinformatics, 16 pages, 5 figures
Improving Neural Protein-Protein Interaction Extraction with Knowledge Selection
Dec 11, 2019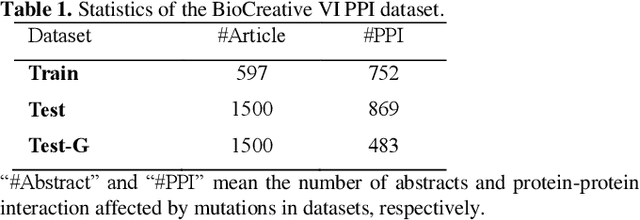
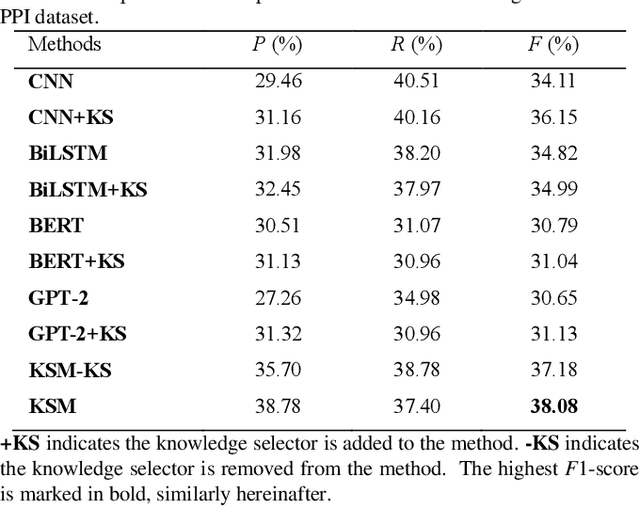
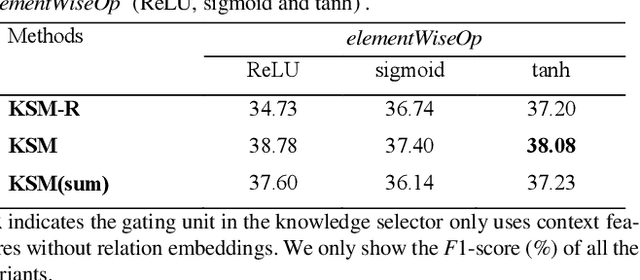
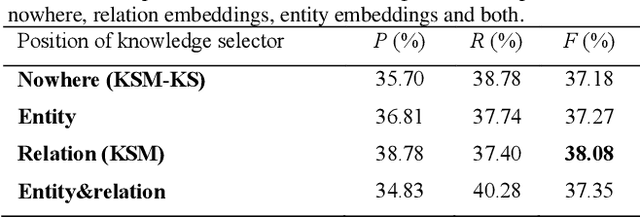
Abstract:Protein-protein interaction (PPI) extraction from published scientific literature provides additional support for precision medicine efforts. Meanwhile, knowledge bases (KBs) contain huge amounts of structured information of protein entities and their relations, which can be encoded in entity and relation embeddings to help PPI extraction. However, the prior knowledge of protein-protein pairs must be selectively used so that it is suitable for different contexts. This paper proposes a Knowledge Selection Model (KSM) to fuse the selected prior knowledge and context information for PPI extraction. Firstly, two Transformers encode the context sequence of a protein pair according to each protein embedding, respectively. Then, the two outputs are fed to a mutual attention to capture the important context features towards the protein pair. Next, the context features are used to distill the relation embedding by a knowledge selector. Finally, the selected relation embedding and the context features are concatenated for PPI extraction. Experiments on the BioCreative VI PPI dataset show that KSM achieves a new state-of-the-art performance (38.08% F1-score) by adding knowledge selection.
* Published in Computational Biology and Chemistry; 14 pages, 2 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge