Kanyakorn Veerakanjana
Med-CoReasoner: Reducing Language Disparities in Medical Reasoning via Language-Informed Co-Reasoning
Jan 13, 2026Abstract:While reasoning-enhanced large language models perform strongly on English medical tasks, a persistent multilingual gap remains, with substantially weaker reasoning in local languages, limiting equitable global medical deployment. To bridge this gap, we introduce Med-CoReasoner, a language-informed co-reasoning framework that elicits parallel English and local-language reasoning, abstracts them into structured concepts, and integrates local clinical knowledge into an English logical scaffold via concept-level alignment and retrieval. This design combines the structural robustness of English reasoning with the practice-grounded expertise encoded in local languages. To evaluate multilingual medical reasoning beyond multiple-choice settings, we construct MultiMed-X, a benchmark covering seven languages with expert-annotated long-form question answering and natural language inference tasks, comprising 350 instances per language. Experiments across three benchmarks show that Med-CoReasoner improves multilingual reasoning performance by an average of 5%, with particularly substantial gains in low-resource languages. Moreover, model distillation and expert evaluation analysis further confirm that Med-CoReasoner produces clinically sound and culturally grounded reasoning traces.
Crowdsource, Crawl, or Generate? Creating SEA-VL, a Multicultural Vision-Language Dataset for Southeast Asia
Mar 10, 2025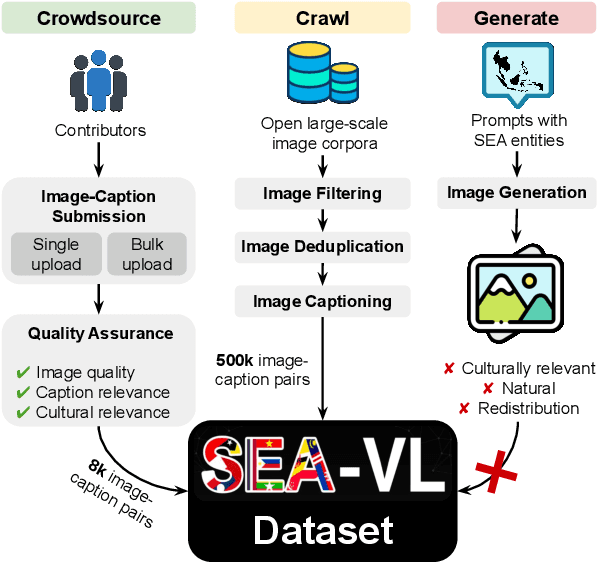
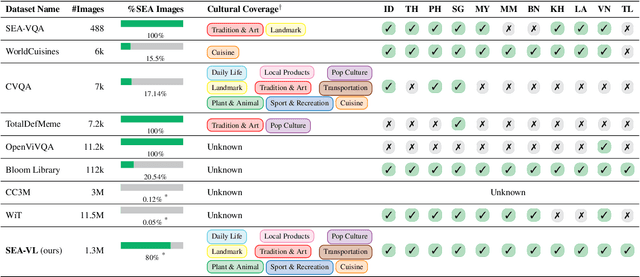
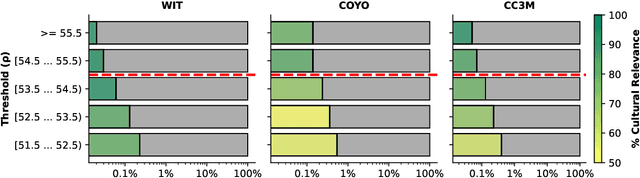

Abstract:Southeast Asia (SEA) is a region of extraordinary linguistic and cultural diversity, yet it remains significantly underrepresented in vision-language (VL) research. This often results in artificial intelligence (AI) models that fail to capture SEA cultural nuances. To fill this gap, we present SEA-VL, an open-source initiative dedicated to developing high-quality, culturally relevant data for SEA languages. By involving contributors from SEA countries, SEA-VL aims to ensure better cultural relevance and diversity, fostering greater inclusivity of underrepresented languages in VL research. Beyond crowdsourcing, our initiative goes one step further in the exploration of the automatic collection of culturally relevant images through crawling and image generation. First, we find that image crawling achieves approximately ~85% cultural relevance while being more cost- and time-efficient than crowdsourcing. Second, despite the substantial progress in generative vision models, synthetic images remain unreliable in accurately reflecting SEA cultures. The generated images often fail to reflect the nuanced traditions and cultural contexts of the region. Collectively, we gather 1.28M SEA culturally-relevant images, more than 50 times larger than other existing datasets. Through SEA-VL, we aim to bridge the representation gap in SEA, fostering the development of more inclusive AI systems that authentically represent diverse cultures across SEA.
On Creating an English-Thai Code-switched Machine Translation in Medical Domain
Oct 21, 2024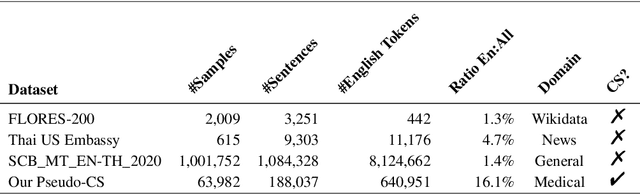
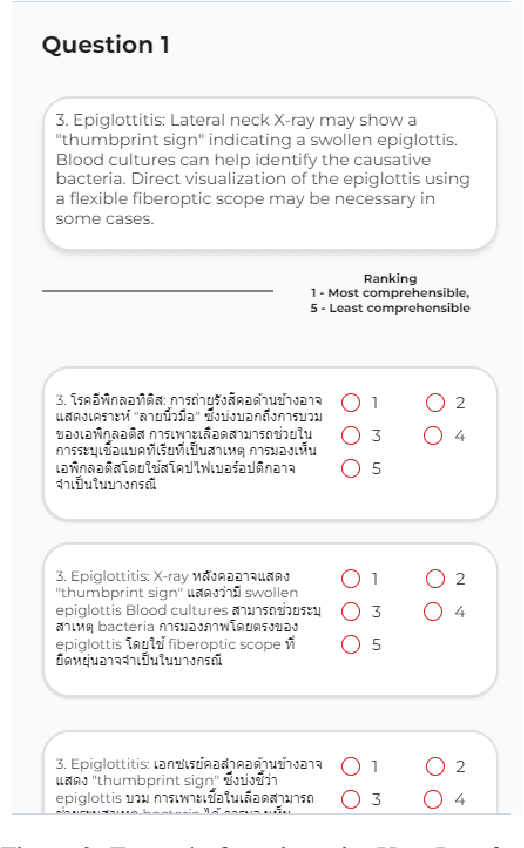

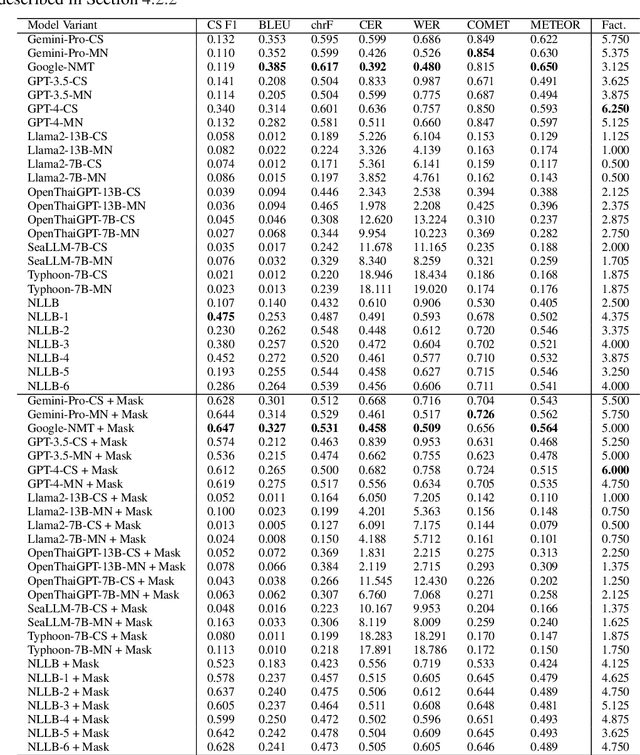
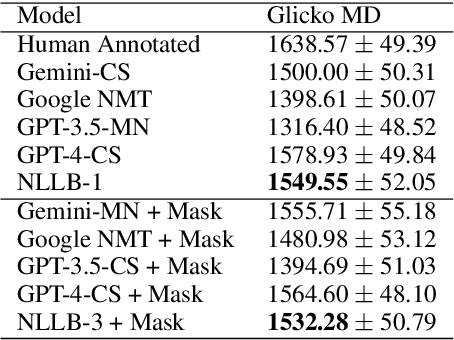
Abstract:Machine translation (MT) in the medical domain plays a pivotal role in enhancing healthcare quality and disseminating medical knowledge. Despite advancements in English-Thai MT technology, common MT approaches often underperform in the medical field due to their inability to precisely translate medical terminologies. Our research prioritizes not merely improving translation accuracy but also maintaining medical terminology in English within the translated text through code-switched (CS) translation. We developed a method to produce CS medical translation data, fine-tuned a CS translation model with this data, and evaluated its performance against strong baselines, such as Google Neural Machine Translation (NMT) and GPT-3.5/GPT-4. Our model demonstrated competitive performance in automatic metrics and was highly favored in human preference evaluations. Our evaluation result also shows that medical professionals significantly prefer CS translations that maintain critical English terms accurately, even if it slightly compromises fluency. Our code and test set are publicly available https://github.com/preceptorai-org/NLLB_CS_EM_NLP2024.
PseudoCell: Hard Negative Mining as Pseudo Labeling for Deep Learning-Based Centroblast Cell Detection
Jul 06, 2023Abstract:Patch classification models based on deep learning have been utilized in whole-slide images (WSI) of H&E-stained tissue samples to assist pathologists in grading follicular lymphoma patients. However, these approaches still require pathologists to manually identify centroblast cells and provide refined labels for optimal performance. To address this, we propose PseudoCell, an object detection framework to automate centroblast detection in WSI (source code is available at https://github.com/IoBT-VISTEC/PseudoCell.git). This framework incorporates centroblast labels from pathologists and combines them with pseudo-negative labels obtained from undersampled false-positive predictions using the cell's morphological features. By employing PseudoCell, pathologists' workload can be reduced as it accurately narrows down the areas requiring their attention during examining tissue. Depending on the confidence threshold, PseudoCell can eliminate 58.18-99.35% of non-centroblasts tissue areas on WSI. This study presents a practical centroblast prescreening method that does not require pathologists' refined labels for improvement. Detailed guidance on the practical implementation of PseudoCell is provided in the discussion section.
PACMAN: a framework for pulse oximeter digit detection and reading in a low-resource setting
Dec 09, 2022



Abstract:In light of the COVID-19 pandemic, patients were required to manually input their daily oxygen saturation (SpO2) and pulse rate (PR) values into a health monitoring system-unfortunately, such a process trend to be an error in typing. Several studies attempted to detect the physiological value from the captured image using optical character recognition (OCR). However, the technology has limited availability with high cost. Thus, this study aimed to propose a novel framework called PACMAN (Pandemic Accelerated Human-Machine Collaboration) with a low-resource deep learning-based computer vision. We compared state-of-the-art object detection algorithms (scaled YOLOv4, YOLOv5, and YOLOR), including the commercial OCR tools for digit recognition on the captured images from pulse oximeter display. All images were derived from crowdsourced data collection with varying quality and alignment. YOLOv5 was the best-performing model against the given model comparison across all datasets, notably the correctly orientated image dataset. We further improved the model performance with the digits auto-orientation algorithm and applied a clustering algorithm to extract SpO2 and PR values. The accuracy performance of YOLOv5 with the implementations was approximately 81.0-89.5%, which was enhanced compared to without any additional implementation. Accordingly, this study highlighted the completion of PACMAN framework to detect and read digits in real-world datasets. The proposed framework has been currently integrated into the patient monitoring system utilized by hospitals nationwide.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge