Jorge Samper-González
ARAMIS
Accuracy of MRI Classification Algorithms in a Tertiary Memory Center Clinical Routine Cohort
Mar 19, 2020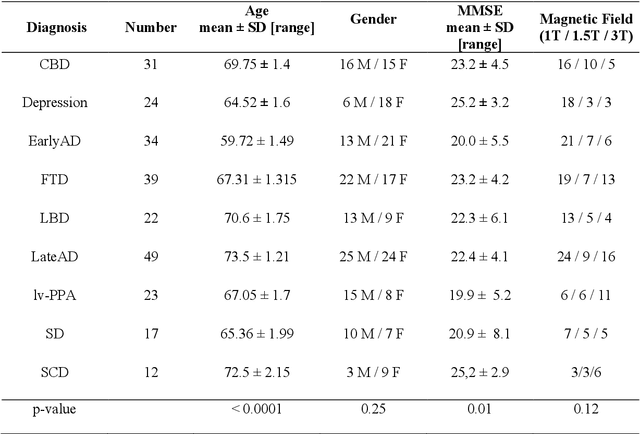
Abstract:BACKGROUND:Automated volumetry software (AVS) has recently become widely available to neuroradiologists. MRI volumetry with AVS may support the diagnosis of dementias by identifying regional atrophy. Moreover, automatic classifiers using machine learning techniques have recently emerged as promising approaches to assist diagnosis. However, the performance of both AVS and automatic classifiers has been evaluated mostly in the artificial setting of research datasets.OBJECTIVE:Our aim was to evaluate the performance of two AVS and an automatic classifier in the clinical routine condition of a memory clinic.METHODS:We studied 239 patients with cognitive troubles from a single memory center cohort. Using clinical routine T1-weighted MRI, we evaluated the classification performance of: 1) univariate volumetry using two AVS (volBrain and Neuroreader$^{TM}$); 2) Support Vector Machine (SVM) automatic classifier, using either the AVS volumes (SVM-AVS), or whole gray matter (SVM-WGM); 3) reading by two neuroradiologists. The performance measure was the balanced diagnostic accuracy. The reference standard was consensus diagnosis by three neurologists using clinical, biological (cerebrospinal fluid) and imaging data and following international criteria.RESULTS:Univariate AVS volumetry provided only moderate accuracies (46% to 71% with hippocampal volume). The accuracy improved when using SVM-AVS classifier (52% to 85%), becoming close to that of SVM-WGM (52 to 90%). Visual classification by neuroradiologists ranged between SVM-AVS and SVM-WGM.CONCLUSION:In the routine practice of a memory clinic, the use of volumetric measures provided by AVS yields only moderate accuracy. Automatic classifiers can improve accuracy and could be a useful tool to assist diagnosis.
Reproducible evaluation of classification methods in Alzheimer's disease: framework and application to MRI and PET data
Aug 20, 2018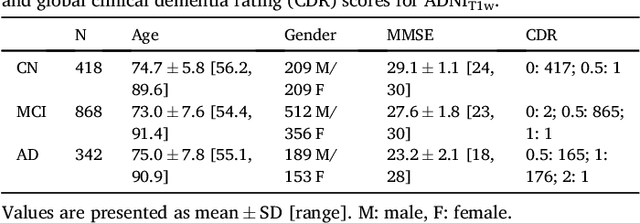
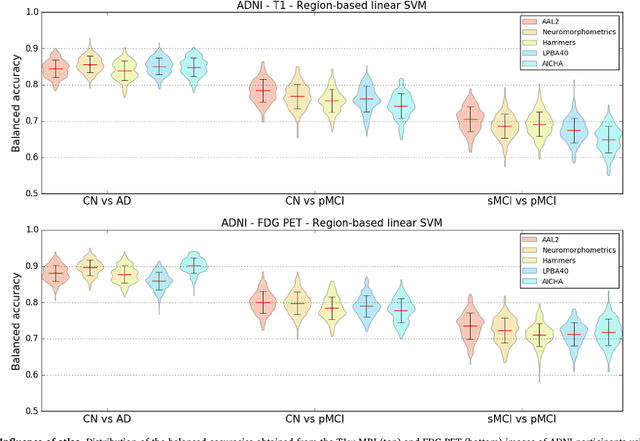
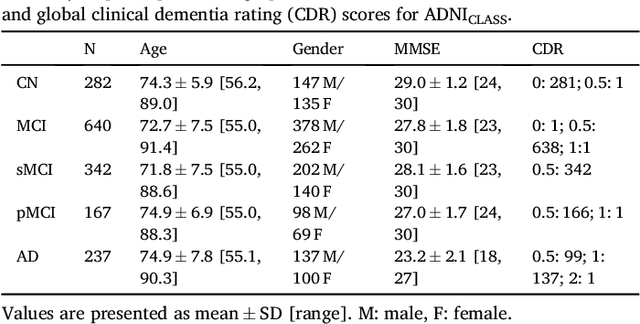
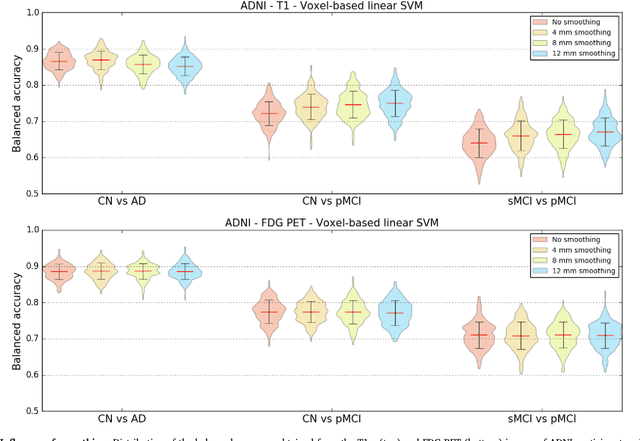
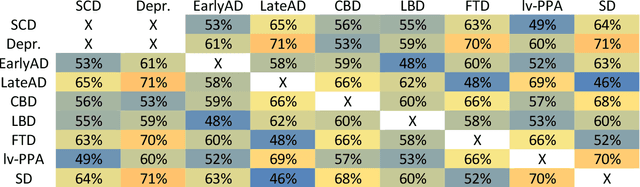
Abstract:A large number of papers have introduced novel machine learning and feature extraction methods for automatic classification of AD. However, they are difficult to reproduce because key components of the validation are often not readily available. These components include selected participants and input data, image preprocessing and cross-validation procedures. The performance of the different approaches is also difficult to compare objectively. In particular, it is often difficult to assess which part of the method provides a real improvement, if any. We propose a framework for reproducible and objective classification experiments in AD using three publicly available datasets (ADNI, AIBL and OASIS). The framework comprises: i) automatic conversion of the three datasets into BIDS format, ii) a modular set of preprocessing pipelines, feature extraction and classification methods, together with an evaluation framework, that provide a baseline for benchmarking the different components. We demonstrate the use of the framework for a large-scale evaluation on 1960 participants using T1 MRI and FDG PET data. In this evaluation, we assess the influence of different modalities, preprocessing, feature types, classifiers, training set sizes and datasets. Performances were in line with the state-of-the-art. FDG PET outperformed T1 MRI for all classification tasks. No difference in performance was found for the use of different atlases, image smoothing, partial volume correction of FDG PET images, or feature type. Linear SVM and L2-logistic regression resulted in similar performance and both outperformed random forests. The classification performance increased along with the number of subjects used for training. Classifiers trained on ADNI generalized well to AIBL and OASIS. All the code of the framework and the experiments is publicly available at: https://gitlab.icm-institute.org/aramislab/AD-ML.
Yet Another ADNI Machine Learning Paper? Paving The Way Towards Fully-reproducible Research on Classification of Alzheimer's Disease
Sep 21, 2017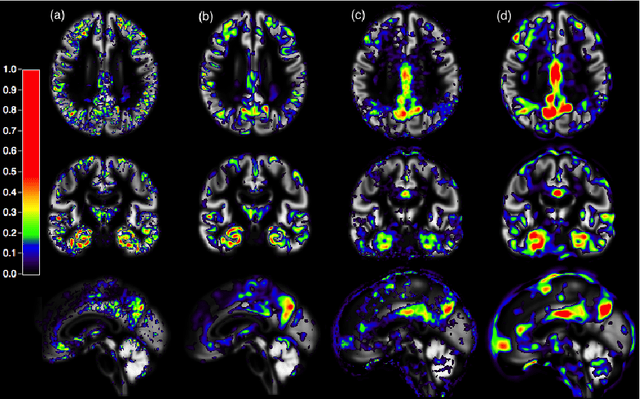
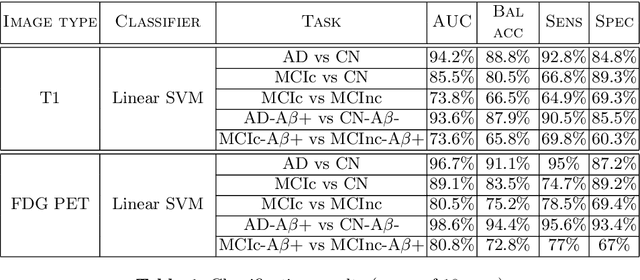
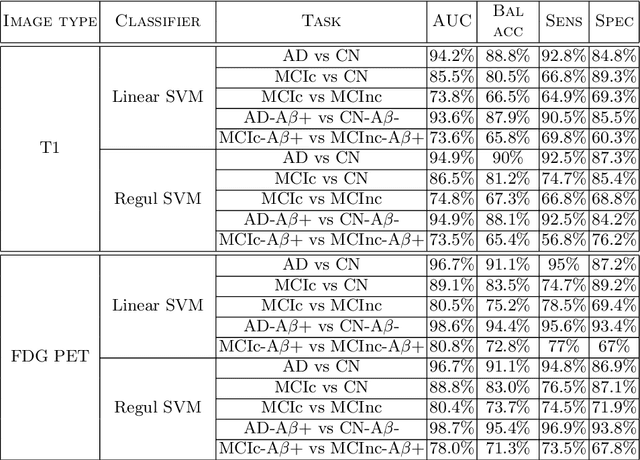

Abstract:In recent years, the number of papers on Alzheimer's disease classification has increased dramatically, generating interesting methodological ideas on the use machine learning and feature extraction methods. However, practical impact is much more limited and, eventually, one could not tell which of these approaches are the most efficient. While over 90\% of these works make use of ADNI an objective comparison between approaches is impossible due to variations in the subjects included, image pre-processing, performance metrics and cross-validation procedures. In this paper, we propose a framework for reproducible classification experiments using multimodal MRI and PET data from ADNI. The core components are: 1) code to automatically convert the full ADNI database into BIDS format; 2) a modular architecture based on Nipype in order to easily plug-in different classification and feature extraction tools; 3) feature extraction pipelines for MRI and PET data; 4) baseline classification approaches for unimodal and multimodal features. This provides a flexible framework for benchmarking different feature extraction and classification tools in a reproducible manner. We demonstrate its use on all (1519) baseline T1 MR images and all (1102) baseline FDG PET images from ADNI 1, GO and 2 with SPM-based feature extraction pipelines and three different classification techniques (linear SVM, anatomically regularized SVM and multiple kernel learning SVM). The highest accuracies achieved were: 91% for AD vs CN, 83% for MCIc vs CN, 75% for MCIc vs MCInc, 94% for AD-A$\beta$+ vs CN-A$\beta$- and 72% for MCIc-A$\beta$+ vs MCInc-A$\beta$+. The code is publicly available at https://gitlab.icm-institute.org/aramislab/AD-ML (depends on the Clinica software platform, publicly available at http://www.clinica.run).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge