Jack Langerman
Explaining Human Preferences via Metrics for Structured 3D Reconstruction
Mar 11, 2025



Abstract:"What cannot be measured cannot be improved" while likely never uttered by Lord Kelvin, summarizes effectively the purpose of this work. This paper presents a detailed evaluation of automated metrics for evaluating structured 3D reconstructions. Pitfalls of each metric are discussed, and a thorough analyses through the lens of expert 3D modelers' preferences is presented. A set of systematic "unit tests" are proposed to empirically verify desirable properties, and context aware recommendations as to which metric to use depending on application are provided. Finally, a learned metric distilled from human expert judgments is proposed and analyzed.
Domain Adaptation of Networks for Camera Pose Estimation: Learning Camera Pose Estimation Without Pose Labels
Nov 29, 2021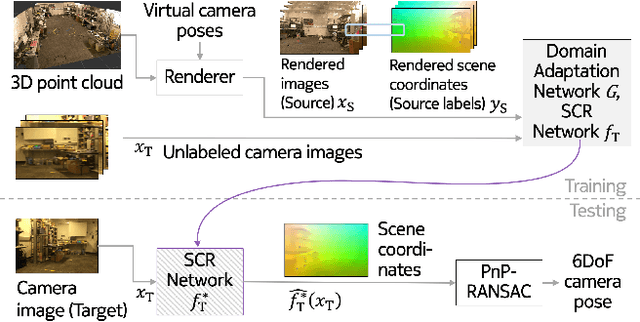
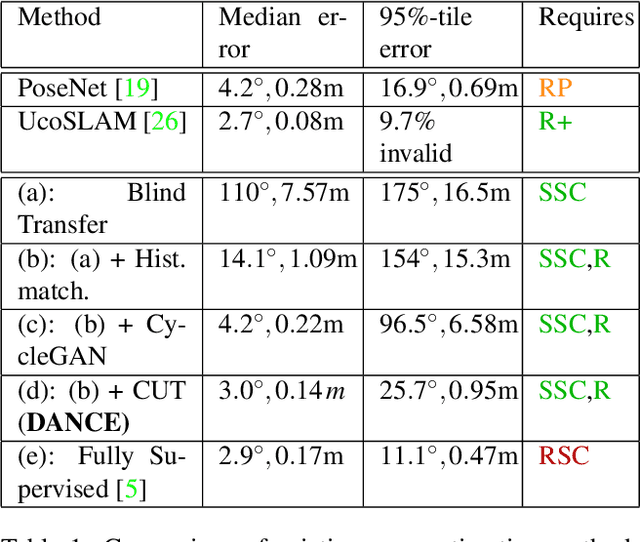
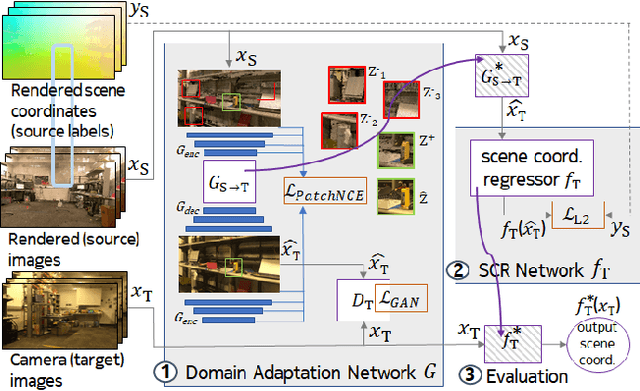

Abstract:One of the key criticisms of deep learning is that large amounts of expensive and difficult-to-acquire training data are required in order to train models with high performance and good generalization capabilities. Focusing on the task of monocular camera pose estimation via scene coordinate regression (SCR), we describe a novel method, Domain Adaptation of Networks for Camera pose Estimation (DANCE), which enables the training of models without access to any labels on the target task. DANCE requires unlabeled images (without known poses, ordering, or scene coordinate labels) and a 3D representation of the space (e.g., a scanned point cloud), both of which can be captured with minimal effort using off-the-shelf commodity hardware. DANCE renders labeled synthetic images from the 3D model, and bridges the inevitable domain gap between synthetic and real images by applying unsupervised image-level domain adaptation techniques (unpaired image-to-image translation). When tested on real images, the SCR model trained with DANCE achieved comparable performance to its fully supervised counterpart (in both cases using PnP-RANSAC for final pose estimation) at a fraction of the cost. Our code and dataset are available at https://github.com/JackLangerman/dance
Deep Mouse: An End-to-end Auto-context Refinement Framework for Brain Ventricle and Body Segmentation in Embryonic Mice Ultrasound Volumes
Oct 30, 2019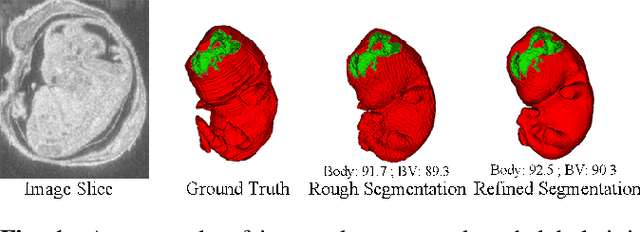
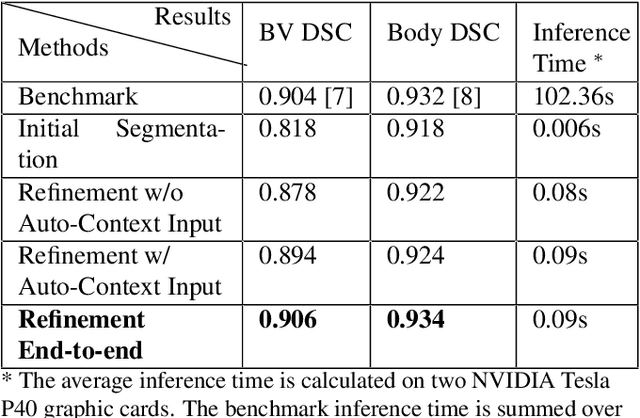
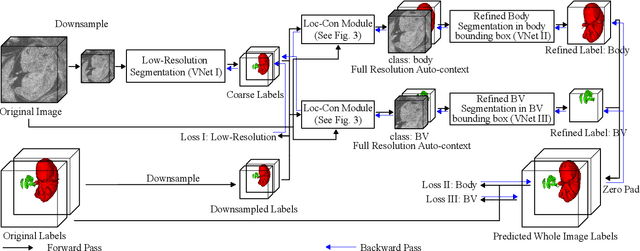
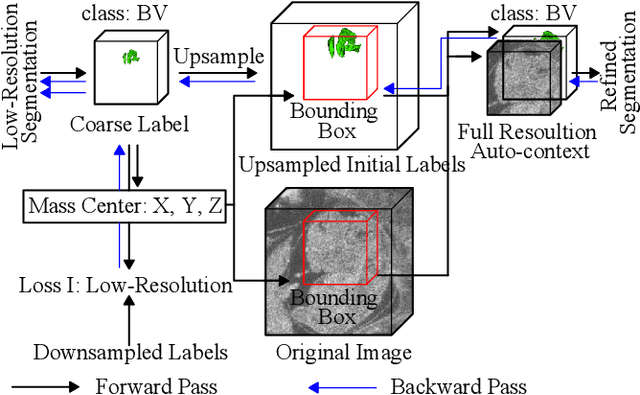
Abstract:High-frequency ultrasound (HFU) is well suited for imaging embryonic mice due to its noninvasive and real-time characteristics. However, manual segmentation of the brain ventricles (BVs) and body requires substantial time and expertise. This work proposes a novel deep learning based end-to-end auto-context refinement framework, consisting of two stages. The first stage produces a low resolution segmentation of the BV and body simultaneously. The resulting probability map for each object (BV or body) is then used to crop a region of interest (ROI) around the target object in both the original image and the probability map to provide context to the refinement segmentation network. Joint training of the two stages provides significant improvement in Dice Similarity Coefficient (DSC) over using only the first stage (0.818 to 0.906 for the BV, and 0.919 to 0.934 for the body). The proposed method significantly reduces the inference time (102.36 to 0.09 s/volume around 1000x faster) while slightly improves the segmentation accuracy over the previous methods using slide-window approaches.
Automatic Mouse Embryo Brain Ventricle & Body Segmentation and Mutant Classification From Ultrasound Data Using Deep Learning
Sep 23, 2019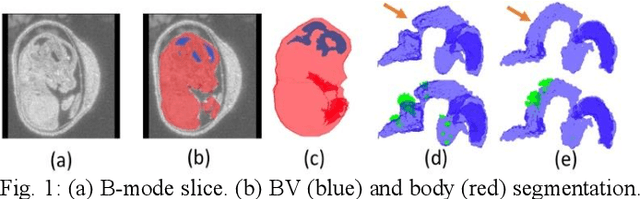
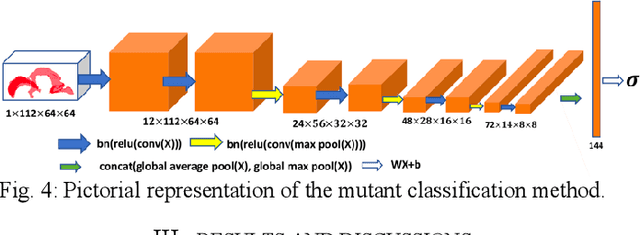
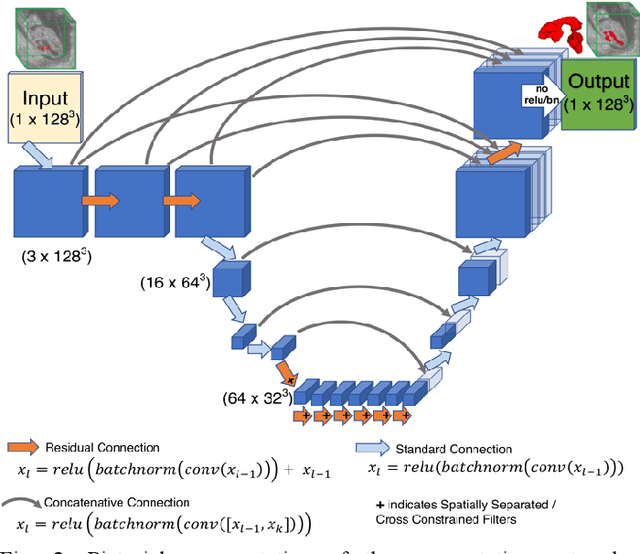
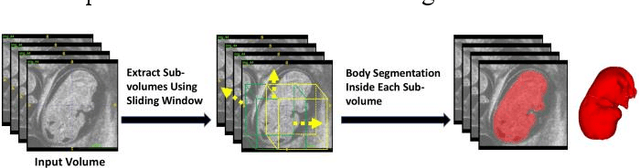

Abstract:High-frequency ultrasound (HFU) is well suited for imaging embryonic mice in vivo because it is non-invasive and real-time. Manual segmentation of the brain ventricles (BVs) and whole body from 3D HFU images is time-consuming and requires specialized training. This paper presents a deep-learning-based segmentation pipeline which automates several time-consuming, repetitive tasks currently performed to study genetic mutations in developing mouse embryos. Namely, the pipeline accurately segments the BV and body regions in 3D HFU images of mouse embryos, despite significant challenges due to position and shape variation of the embryos, as well as imaging artifacts. Based on the BV segmentation, a 3D convolutional neural network (CNN) is further trained to detect embryos with the Engrailed-1 (En1) mutation. The algorithms achieve 0.896 and 0.925 Dice Similarity Coefficient (DSC) for BV and body segmentation, respectively, and 95.8% accuracy on mutant classification. Through gradient based interrogation and visualization of the trained classifier, it is demonstrated that the model focuses on the morphological structures known to be affected by the En1 mutation.
Deep BV: A Fully Automated System for Brain Ventricle Localization and Segmentation in 3D Ultrasound Images of Embryonic Mice
Nov 05, 2018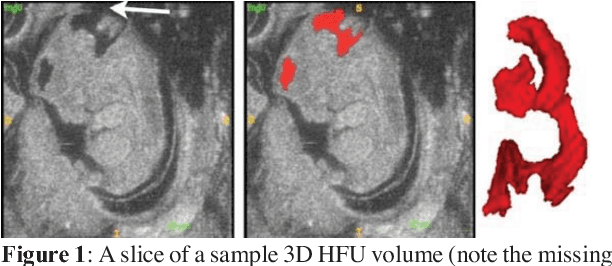
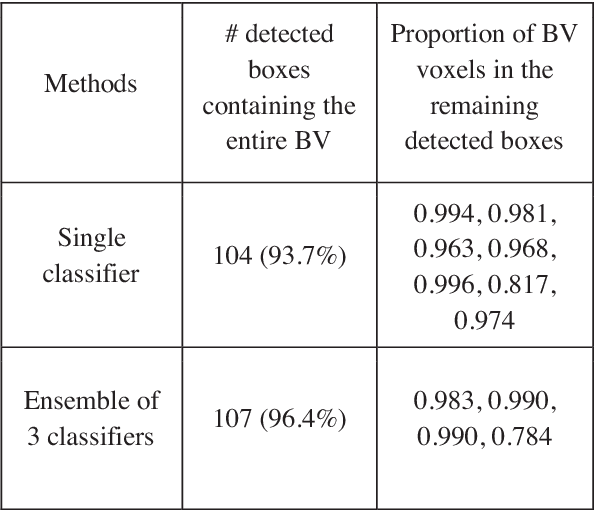
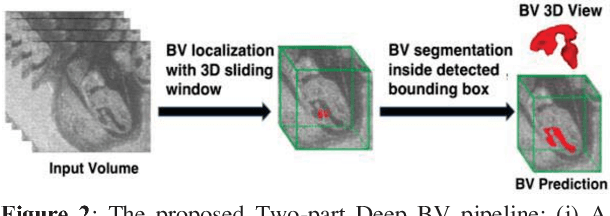
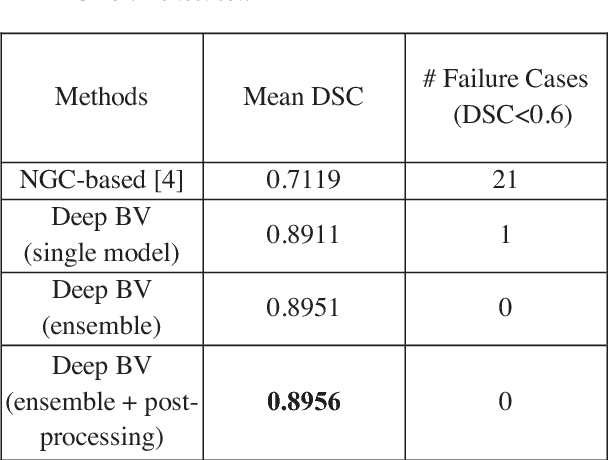
Abstract:Volumetric analysis of brain ventricle (BV) structure is a key tool in the study of central nervous system development in embryonic mice. High-frequency ultrasound (HFU) is the only non-invasive, real-time modality available for rapid volumetric imaging of embryos in utero. However, manual segmentation of the BV from HFU volumes is tedious, time-consuming, and requires specialized expertise. In this paper, we propose a novel deep learning based BV segmentation system for whole-body HFU images of mouse embryos. Our fully automated system consists of two modules: localization and segmentation. It first applies a volumetric convolutional neural network on a 3D sliding window over the entire volume to identify a 3D bounding box containing the entire BV. It then employs a fully convolutional network to segment the detected bounding box into BV and background. The system achieves a Dice Similarity Coefficient (DSC) of 0.8956 for BV segmentation on an unseen 111 HFU volume test set surpassing the previous state-of-the-art method (DSC of 0.7119) by a margin of 25%.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge