Guilherme Pombo
Bidirectional human-AI collaboration in brain tumour assessments improves both expert human and AI agent performance
Dec 13, 2025Abstract:The benefits of artificial intelligence (AI) human partnerships-evaluating how AI agents enhance expert human performance-are increasingly studied. Though rarely evaluated in healthcare, an inverse approach is possible: AI benefiting from the support of an expert human agent. Here, we investigate both human-AI clinical partnership paradigms in the magnetic resonance imaging-guided characterisation of patients with brain tumours. We reveal that human-AI partnerships improve accuracy and metacognitive ability not only for radiologists supported by AI, but also for AI agents supported by radiologists. Moreover, the greatest patient benefit was evident with an AI agent supported by a human one. Synergistic improvements in agent accuracy, metacognitive performance, and inter-rater agreement suggest that AI can create more capable, confident, and consistent clinical agents, whether human or model-based. Our work suggests that the maximal value of AI in healthcare could emerge not from replacing human intelligence, but from AI agents that routinely leverage and amplify it.
Generalizable automated ischaemic stroke lesion segmentation with vision transformers
Feb 10, 2025
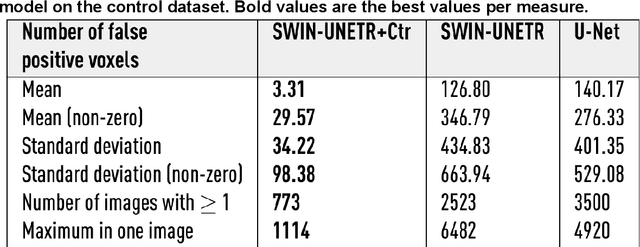
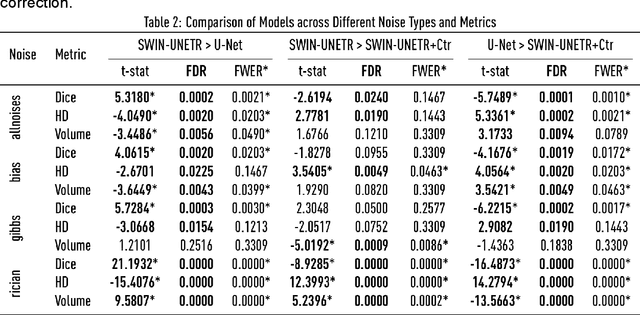
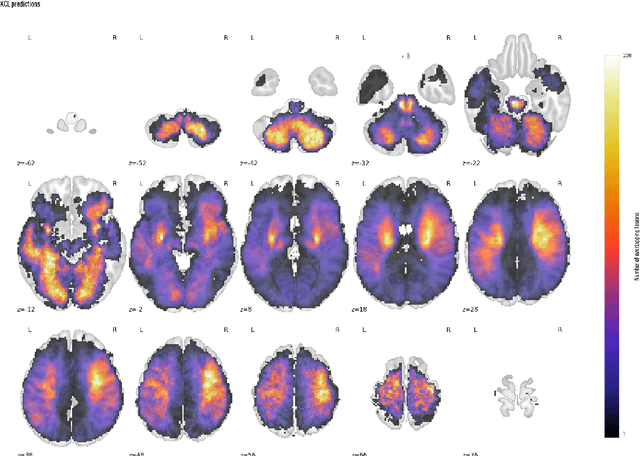
Abstract:Ischaemic stroke, a leading cause of death and disability, critically relies on neuroimaging for characterising the anatomical pattern of injury. Diffusion-weighted imaging (DWI) provides the highest expressivity in ischemic stroke but poses substantial challenges for automated lesion segmentation: susceptibility artefacts, morphological heterogeneity, age-related comorbidities, time-dependent signal dynamics, instrumental variability, and limited labelled data. Current U-Net-based models therefore underperform, a problem accentuated by inadequate evaluation metrics that focus on mean performance, neglecting anatomical, subpopulation, and acquisition-dependent variability. Here, we present a high-performance DWI lesion segmentation tool addressing these challenges through optimized vision transformer-based architectures, integration of 3563 annotated lesions from multi-site data, and algorithmic enhancements, achieving state-of-the-art results. We further propose a novel evaluative framework assessing model fidelity, equity (across demographics and lesion subtypes), anatomical precision, and robustness to instrumental variability, promoting clinical and research utility. This work advances stroke imaging by reconciling model expressivity with domain-specific challenges and redefining performance benchmarks to prioritize equity and generalizability, critical for personalized medicine and mechanistic research.
Deep generative computed perfusion-deficit mapping of ischaemic stroke
Feb 03, 2025



Abstract:Focal deficits in ischaemic stroke result from impaired perfusion downstream of a critical vascular occlusion. While parenchymal lesions are traditionally used to predict clinical deficits, the underlying pattern of disrupted perfusion provides information upstream of the lesion, potentially yielding earlier predictive and localizing signals. Such perfusion maps can be derived from routine CT angiography (CTA) widely deployed in clinical practice. Analysing computed perfusion maps from 1,393 CTA-imaged-patients with acute ischaemic stroke, we use deep generative inference to localise neural substrates of NIHSS sub-scores. We show that our approach replicates known lesion-deficit relations without knowledge of the lesion itself and reveals novel neural dependents. The high achieved anatomical fidelity suggests acute CTA-derived computed perfusion maps may be of substantial clinical-and-scientific value in rich phenotyping of acute stroke. Using only hyperacute imaging, deep generative inference could power highly expressive models of functional anatomical relations in ischaemic stroke within the pre-interventional window.
The legibility of the imaged human brain
Aug 23, 2023

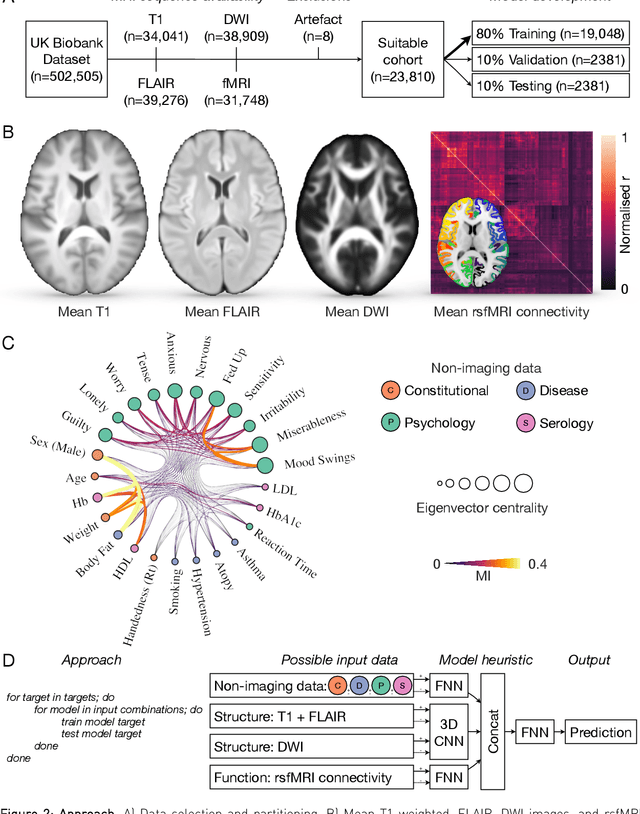

Abstract:Our knowledge of the organisation of the human brain at the population-level is yet to translate into power to predict functional differences at the individual-level, limiting clinical applications, and casting doubt on the generalisability of inferred mechanisms. It remains unknown whether the difficulty arises from the absence of individuating biological patterns within the brain, or from limited power to access them with the models and compute at our disposal. Here we comprehensively investigate the resolvability of such patterns with data and compute at unprecedented scale. Across 23810 unique participants from UK Biobank, we systematically evaluate the predictability of 25 individual biological characteristics, from all available combinations of structural and functional neuroimaging data. Over 4526 GPU*hours of computation, we train, optimize, and evaluate out-of-sample 700 individual predictive models, including multilayer perceptrons of demographic, psychological, serological, chronic morbidity, and functional connectivity characteristics, and both uni- and multi-modal 3D convolutional neural network models of macro- and micro-structural brain imaging. We find a marked discrepancy between the high predictability of sex (balanced accuracy 99.7%), age (mean absolute error 2.048 years, R2 0.859), and weight (mean absolute error 2.609Kg, R2 0.625), for which we set new state-of-the-art performance, and the surprisingly low predictability of other characteristics. Neither structural nor functional imaging predicted individual psychology better than the coincidence of common chronic morbidity (p<0.05). Serology predicted common morbidity (p<0.05) and was best predicted by it (p<0.001), followed by structural neuroimaging (p<0.05). Our findings suggest either more informative imaging or more powerful models will be needed to decipher individual level characteristics from the brain.
The minimal computational substrate of fluid intelligence
Aug 14, 2023Abstract:The quantification of cognitive powers rests on identifying a behavioural task that depends on them. Such dependence cannot be assured, for the powers a task invokes cannot be experimentally controlled or constrained a priori, resulting in unknown vulnerability to failure of specificity and generalisability. Evaluating a compact version of Raven's Advanced Progressive Matrices (RAPM), a widely used clinical test of fluid intelligence, we show that LaMa, a self-supervised artificial neural network trained solely on the completion of partially masked images of natural environmental scenes, achieves human-level test scores a prima vista, without any task-specific inductive bias or training. Compared with cohorts of healthy and focally lesioned participants, LaMa exhibits human-like variation with item difficulty, and produces errors characteristic of right frontal lobe damage under degradation of its ability to integrate global spatial patterns. LaMa's narrow training and limited capacity -- comparable to the nervous system of the fruit fly -- suggest RAPM may be open to computationally simple solutions that need not necessarily invoke abstract reasoning.
Deep Variational Lesion-Deficit Mapping
May 27, 2023Abstract:Causal mapping of the functional organisation of the human brain requires evidence of \textit{necessity} available at adequate scale only from pathological lesions of natural origin. This demands inferential models with sufficient flexibility to capture both the observable distribution of pathological damage and the unobserved distribution of the neural substrate. Current model frameworks -- both mass-univariate and multivariate -- either ignore distributed lesion-deficit relations or do not model them explicitly, relying on featurization incidental to a predictive task. Here we initiate the application of deep generative neural network architectures to the task of lesion-deficit inference, formulating it as the estimation of an expressive hierarchical model of the joint lesion and deficit distributions conditioned on a latent neural substrate. We implement such deep lesion deficit inference with variational convolutional volumetric auto-encoders. We introduce a comprehensive framework for lesion-deficit model comparison, incorporating diverse candidate substrates, forms of substrate interactions, sample sizes, noise corruption, and population heterogeneity. Drawing on 5500 volume images of ischaemic stroke, we show that our model outperforms established methods by a substantial margin across all simulation scenarios, including comparatively small-scale and noisy data regimes. Our analysis justifies the widespread adoption of this approach, for which we provide an open source implementation: https://github.com/guilherme-pombo/vae_lesion_deficit
Equitable modelling of brain imaging by counterfactual augmentation with morphologically constrained 3D deep generative models
Nov 29, 2021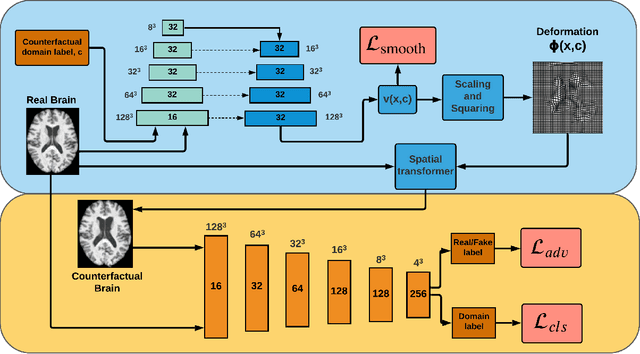
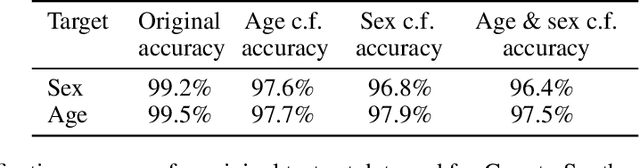

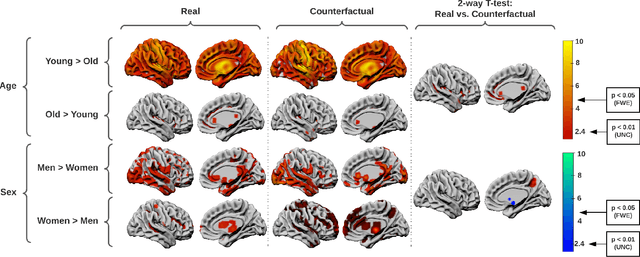
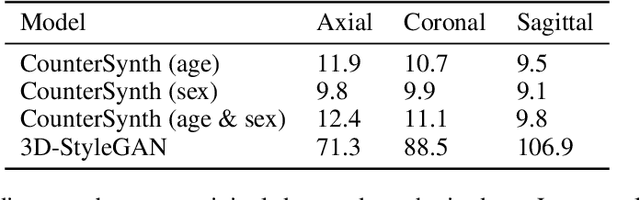
Abstract:We describe Countersynth, a conditional generative model of diffeomorphic deformations that induce label-driven, biologically plausible changes in volumetric brain images. The model is intended to synthesise counterfactual training data augmentations for downstream discriminative modelling tasks where fidelity is limited by data imbalance, distributional instability, confounding, or underspecification, and exhibits inequitable performance across distinct subpopulations. Focusing on demographic attributes, we evaluate the quality of synthesized counterfactuals with voxel-based morphometry, classification and regression of the conditioning attributes, and the Fr\'{e}chet inception distance. Examining downstream discriminative performance in the context of engineered demographic imbalance and confounding, we use UK Biobank magnetic resonance imaging data to benchmark CounterSynth augmentation against current solutions to these problems. We achieve state-of-the-art improvements, both in overall fidelity and equity. The source code for CounterSynth is available online.
Bayesian Volumetric Autoregressive generative models for better semisupervised learning
Jul 26, 2019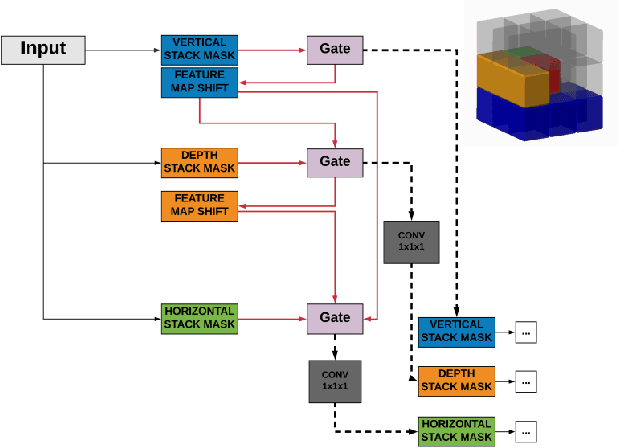
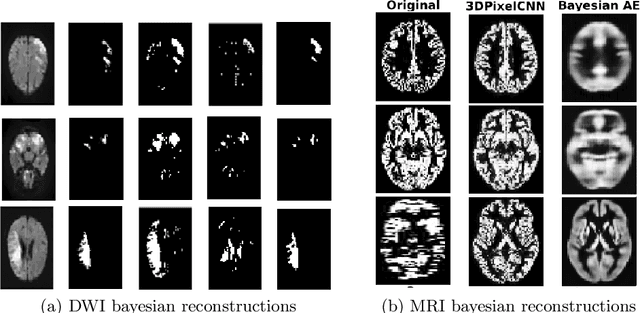

Abstract:Deep generative models are rapidly gaining traction in medical imaging. Nonetheless, most generative architectures struggle to capture the underlying probability distributions of volumetric data, exhibit convergence problems, and offer no robust indices of model uncertainty. By comparison, the autoregressive generative model PixelCNN can be extended to volumetric data with relative ease, it readily attempts to learn the true underlying probability distribution and it still admits a Bayesian reformulation that provides a principled framework for reasoning about model uncertainty. Our contributions in this paper are two fold: first, we extend PixelCNN to work with volumetric brain magnetic resonance imaging data. Second, we show that reformulating this model to approximate a deep Gaussian process yields a measure of uncertainty that improves the performance of semi-supervised learning, in particular classification performance in settings where the proportion of labelled data is low. We quantify this improvement across classification, regression, and semantic segmentation tasks, training and testing on clinical magnetic resonance brain imaging data comprising T1-weighted and diffusion-weighted sequences.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge