Emily B. Fox
Uncertainty Quantification for Distribution-to-Distribution Flow Matching in Scientific Imaging
Mar 23, 2026Abstract:Distribution-to-distribution generative models support scientific imaging tasks ranging from modeling cellular perturbation responses to translating medical images across conditions. Trustworthy generation requires both reliability (generalization across labs, devices, and experimental conditions) and accountability (detecting out-of-distribution cases where predictions may be unreliable). Uncertainty quantification (UQ) based approaches serve as promising candidates for these tasks, yet UQ for distribution-to-distribution generative models remains underexplored. We present a unified UQ framework, Bayesian Stochastic Flow Matching (BSFM), that disentangles aleatoric and epistemic uncertainty. The Stochastic Flow Matching (SFM) component augments deterministic flows with a diffusion term to improve model generalization to unseen scenarios. For UQ, we develop a scalable Bayesian approach -- MCD-Antithetic -- that combines Monte Carlo Dropout with sample-efficient antithetic sampling to produce effective anomaly scores for out-of-distribution detection. Experiments on cellular imaging (BBBC021, JUMP) and brain fMRI (Theory of Mind) across diverse scenarios show that SFM improves reliability while MCD-Antithetic enhances accountability.
CellFluxRL: Biologically-Constrained Virtual Cell Modeling via Reinforcement Learning
Mar 23, 2026Abstract:Building virtual cells with generative models to simulate cellular behavior in silico is emerging as a promising paradigm for accelerating drug discovery. However, prior image-based generative approaches can produce implausible cell images that violate basic physical and biological constraints. To address this, we propose to post-train virtual cell models with reinforcement learning (RL), leveraging biologically meaningful evaluators as reward functions. We design seven rewards spanning three categories-biological function, structural validity, and morphological correctness-and optimize the state-of-the-art CellFlux model to yield CellFluxRL. CellFluxRL consistently improves over CellFlux across all rewards, with further performance boosts from test-time scaling. Overall, our results present a virtual cell modeling framework that enforces physically-based constraints through RL, advancing beyond "visually realistic" generations towards "biologically meaningful" ones.
How Well Do Multimodal Models Reason on ECG Signals?
Feb 27, 2026Abstract:While multimodal large language models offer a promising solution to the "black box" nature of health AI by generating interpretable reasoning traces, verifying the validity of these traces remains a critical challenge. Existing evaluation methods are either unscalable, relying on manual clinician review, or superficial, utilizing proxy metrics (e.g. QA) that fail to capture the semantic correctness of clinical logic. In this work, we introduce a reproducible framework for evaluating reasoning in ECG signals. We propose decomposing reasoning into two distinct, components: (i) Perception, the accurate identification of patterns within the raw signal, and (ii) Deduction, the logical application of domain knowledge to those patterns. To evaluate Perception, we employ an agentic framework that generates code to empirically verify the temporal structures described in the reasoning trace. To evaluate Deduction, we measure the alignment of the model's logic against a structured database of established clinical criteria in a retrieval-based approach. This dual-verification method enables the scalable assessment of "true" reasoning capabilities.
Test-time regression: a unifying framework for designing sequence models with associative memory
Jan 21, 2025


Abstract:Sequences provide a remarkably general way to represent and process information. This powerful abstraction has placed sequence modeling at the center of modern deep learning applications, inspiring numerous architectures from transformers to recurrent networks. While this fragmented development has yielded powerful models, it has left us without a unified framework to understand their fundamental similarities and explain their effectiveness. We present a unifying framework motivated by an empirical observation: effective sequence models must be able to perform associative recall. Our key insight is that memorizing input tokens through an associative memory is equivalent to performing regression at test-time. This regression-memory correspondence provides a framework for deriving sequence models that can perform associative recall, offering a systematic lens to understand seemingly ad-hoc architectural choices. We show numerous recent architectures -- including linear attention models, their gated variants, state-space models, online learners, and softmax attention -- emerge naturally as specific approaches to test-time regression. Each architecture corresponds to three design choices: the relative importance of each association, the regressor function class, and the optimization algorithm. This connection leads to new understanding: we provide theoretical justification for QKNorm in softmax attention, and we motivate higher-order generalizations of softmax attention. Beyond unification, our work unlocks decades of rich statistical tools that can guide future development of more powerful yet principled sequence models.
Learning Explainable Treatment Policies with Clinician-Informed Representations: A Practical Approach
Nov 26, 2024



Abstract:Digital health interventions (DHIs) and remote patient monitoring (RPM) have shown great potential in improving chronic disease management through personalized care. However, barriers like limited efficacy and workload concerns hinder adoption of existing DHIs; while limited sample sizes and lack of interpretability limit the effectiveness and adoption of purely black-box algorithmic DHIs. In this paper, we address these challenges by developing a pipeline for learning explainable treatment policies for RPM-enabled DHIs. We apply our approach in the real-world setting of RPM using a DHI to improve glycemic control of youth with type 1 diabetes. Our main contribution is to reveal the importance of clinical domain knowledge in developing state and action representations for effective, efficient, and interpretable targeting policies. We observe that policies learned from clinician-informed representations are significantly more efficacious and efficient than policies learned from black-box representations. This work emphasizes the importance of collaboration between ML researchers and clinicians for developing effective DHIs in the real world.
CriticAL: Critic Automation with Language Models
Nov 10, 2024Abstract:Understanding the world through models is a fundamental goal of scientific research. While large language model (LLM) based approaches show promise in automating scientific discovery, they often overlook the importance of criticizing scientific models. Criticizing models deepens scientific understanding and drives the development of more accurate models. Automating model criticism is difficult because it traditionally requires a human expert to define how to compare a model with data and evaluate if the discrepancies are significant--both rely heavily on understanding the modeling assumptions and domain. Although LLM-based critic approaches are appealing, they introduce new challenges: LLMs might hallucinate the critiques themselves. Motivated by this, we introduce CriticAL (Critic Automation with Language Models). CriticAL uses LLMs to generate summary statistics that capture discrepancies between model predictions and data, and applies hypothesis tests to evaluate their significance. We can view CriticAL as a verifier that validates models and their critiques by embedding them in a hypothesis testing framework. In experiments, we evaluate CriticAL across key quantitative and qualitative dimensions. In settings where we synthesize discrepancies between models and datasets, CriticAL reliably generates correct critiques without hallucinating incorrect ones. We show that both human and LLM judges consistently prefer CriticAL's critiques over alternative approaches in terms of transparency and actionability. Finally, we show that CriticAL's critiques enable an LLM scientist to improve upon human-designed models on real-world datasets.
How to Build the Virtual Cell with Artificial Intelligence: Priorities and Opportunities
Sep 18, 2024

Abstract:The cell is arguably the smallest unit of life and is central to understanding biology. Accurate modeling of cells is important for this understanding as well as for determining the root causes of disease. Recent advances in artificial intelligence (AI), combined with the ability to generate large-scale experimental data, present novel opportunities to model cells. Here we propose a vision of AI-powered Virtual Cells, where robust representations of cells and cellular systems under different conditions are directly learned from growing biological data across measurements and scales. We discuss desired capabilities of AI Virtual Cells, including generating universal representations of biological entities across scales, and facilitating interpretable in silico experiments to predict and understand their behavior using Virtual Instruments. We further address the challenges, opportunities and requirements to realize this vision including data needs, evaluation strategies, and community standards and engagement to ensure biological accuracy and broad utility. We envision a future where AI Virtual Cells help identify new drug targets, predict cellular responses to perturbations, as well as scale hypothesis exploration. With open science collaborations across the biomedical ecosystem that includes academia, philanthropy, and the biopharma and AI industries, a comprehensive predictive understanding of cell mechanisms and interactions is within reach.
Hybrid Square Neural ODE Causal Modeling
Feb 27, 2024Abstract:Hybrid models combine mechanistic ODE-based dynamics with flexible and expressive neural network components. Such models have grown rapidly in popularity, especially in scientific domains where such ODE-based modeling offers important interpretability and validated causal grounding (e.g., for counterfactual reasoning). The incorporation of mechanistic models also provides inductive bias in standard blackbox modeling approaches, critical when learning from small datasets or partially observed, complex systems. Unfortunately, as hybrid models become more flexible, the causal grounding provided by the mechanistic model can quickly be lost. We address this problem by leveraging another common source of domain knowledge: ranking of treatment effects for a set of interventions, even if the precise treatment effect is unknown. We encode this information in a causal loss that we combine with the standard predictive loss to arrive at a hybrid loss that biases our learning towards causally valid hybrid models. We demonstrate our ability to achieve a win-win -- state-of-the-art predictive performance and causal validity -- in the challenging task of modeling glucose dynamics during exercise.
Automated Statistical Model Discovery with Language Models
Feb 27, 2024



Abstract:Statistical model discovery involves a challenging search over a vast space of models subject to domain-specific modeling constraints. Efficiently searching over this space requires human expertise in modeling and the problem domain. Motivated by the domain knowledge and programming capabilities of large language models (LMs), we introduce a method for language model driven automated statistical model discovery. We cast our automated procedure within the framework of Box's Loop: the LM iterates between proposing statistical models represented as probabilistic programs, acting as a modeler, and critiquing those models, acting as a domain expert. By leveraging LMs, we do not have to define a domain-specific language of models or design a handcrafted search procedure, key restrictions of previous systems. We evaluate our method in three common settings in probabilistic modeling: searching within a restricted space of models, searching over an open-ended space, and improving classic models under natural language constraints (e.g., this model should be interpretable to an ecologist). Our method matches the performance of previous systems, identifies models on par with human expert designed models, and extends classic models in interpretable ways. Our results highlight the promise of LM driven model discovery.
Interpretable Mechanistic Representations for Meal-level Glycemic Control in the Wild
Dec 06, 2023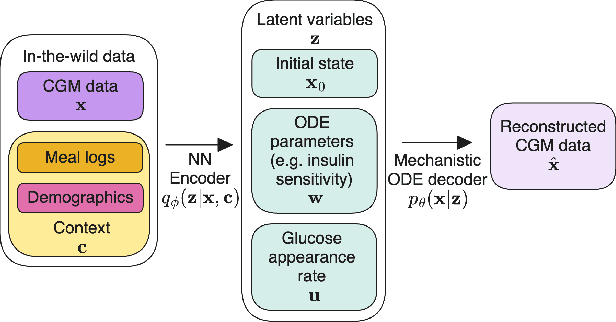
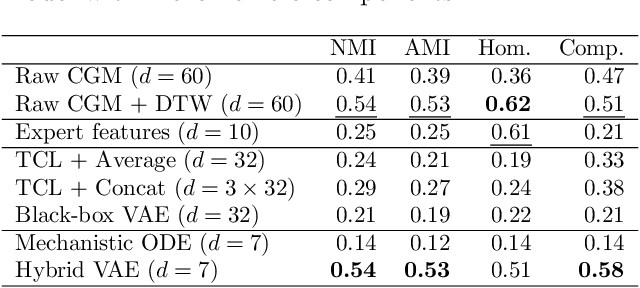
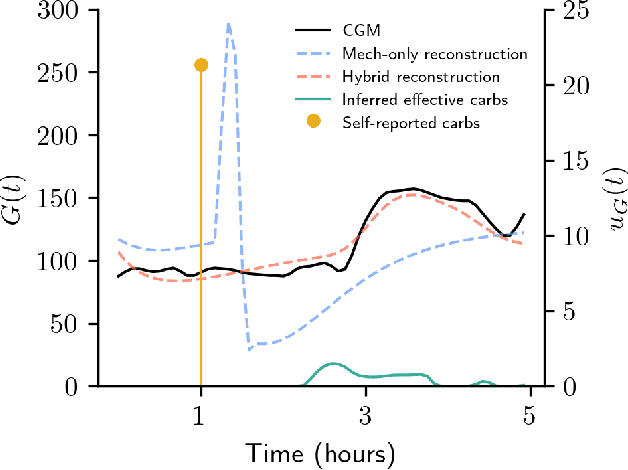
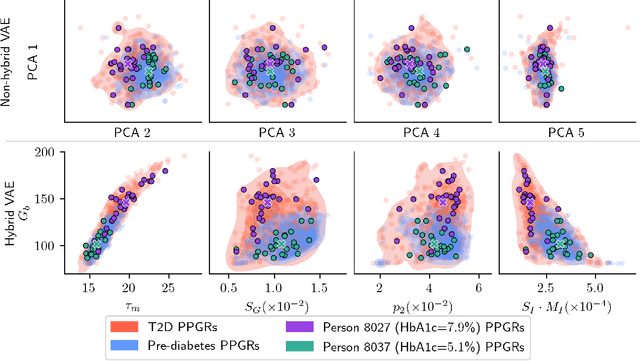
Abstract:Diabetes encompasses a complex landscape of glycemic control that varies widely among individuals. However, current methods do not faithfully capture this variability at the meal level. On the one hand, expert-crafted features lack the flexibility of data-driven methods; on the other hand, learned representations tend to be uninterpretable which hampers clinical adoption. In this paper, we propose a hybrid variational autoencoder to learn interpretable representations of CGM and meal data. Our method grounds the latent space to the inputs of a mechanistic differential equation, producing embeddings that reflect physiological quantities, such as insulin sensitivity, glucose effectiveness, and basal glucose levels. Moreover, we introduce a novel method to infer the glucose appearance rate, making the mechanistic model robust to unreliable meal logs. On a dataset of CGM and self-reported meals from individuals with type-2 diabetes and pre-diabetes, our unsupervised representation discovers a separation between individuals proportional to their disease severity. Our embeddings produce clusters that are up to 4x better than naive, expert, black-box, and pure mechanistic features. Our method provides a nuanced, yet interpretable, embedding space to compare glycemic control within and across individuals, directly learnable from in-the-wild data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge