Chih-chan Tien
Bidirectional Hierarchical Protein Multi-Modal Representation Learning
Apr 07, 2025Abstract:Protein representation learning is critical for numerous biological tasks. Recently, large transformer-based protein language models (pLMs) pretrained on large scale protein sequences have demonstrated significant success in sequence-based tasks. However, pLMs lack structural information. Conversely, graph neural networks (GNNs) designed to leverage 3D structural information have shown promising generalization in protein-related prediction tasks, but their effectiveness is often constrained by the scarcity of labeled structural data. Recognizing that sequence and structural representations are complementary perspectives of the same protein entity, we propose a multimodal bidirectional hierarchical fusion framework to effectively merge these modalities. Our framework employs attention and gating mechanisms to enable effective interaction between pLMs-generated sequential representations and GNN-extracted structural features, improving information exchange and enhancement across layers of the neural network. Based on the framework, we further introduce local Bi-Hierarchical Fusion with gating and global Bi-Hierarchical Fusion with multihead self-attention approaches. Through extensive experiments on a diverse set of protein-related tasks, our method demonstrates consistent improvements over strong baselines and existing fusion techniques in a variety of protein representation learning benchmarks, including react (enzyme/EC classification), model quality assessment (MQA), protein-ligand binding affinity prediction (LBA), protein-protein binding site prediction (PPBS), and B cell epitopes prediction (BCEs). Our method establishes a new state-of-the-art for multimodal protein representation learning, emphasizing the efficacy of BIHIERARCHICAL FUSION in bridging sequence and structural modalities.
Entropy-Reinforced Planning with Large Language Models for Drug Discovery
Jun 11, 2024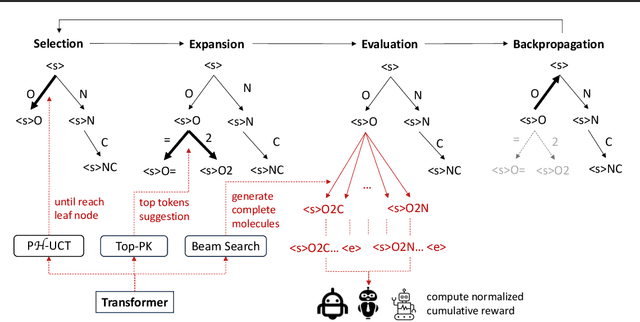
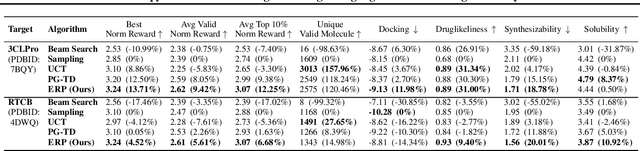
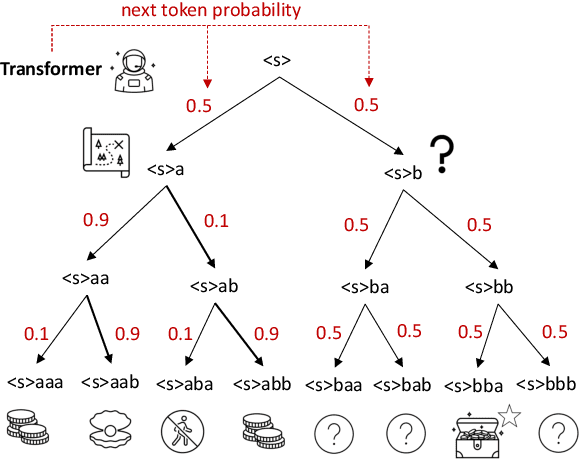
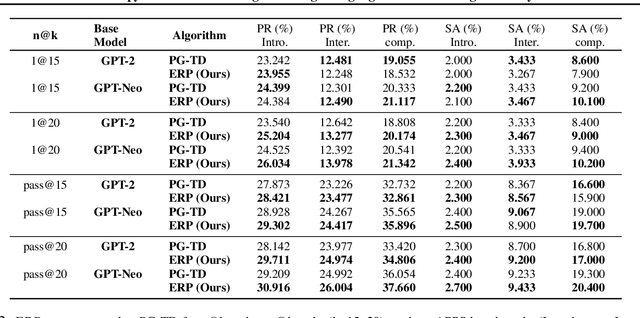
Abstract:The objective of drug discovery is to identify chemical compounds that possess specific pharmaceutical properties toward a binding target. Existing large language models (LLMS) can achieve high token matching scores in terms of likelihood for molecule generation. However, relying solely on LLM decoding often results in the generation of molecules that are either invalid due to a single misused token, or suboptimal due to unbalanced exploration and exploitation as a consequence of the LLMs prior experience. Here we propose ERP, Entropy-Reinforced Planning for Transformer Decoding, which employs an entropy-reinforced planning algorithm to enhance the Transformer decoding process and strike a balance between exploitation and exploration. ERP aims to achieve improvements in multiple properties compared to direct sampling from the Transformer. We evaluated ERP on the SARS-CoV-2 virus (3CLPro) and human cancer cell target protein (RTCB) benchmarks and demonstrated that, in both benchmarks, ERP consistently outperforms the current state-of-the-art algorithm by 1-5 percent, and baselines by 5-10 percent, respectively. Moreover, such improvement is robust across Transformer models trained with different objectives. Finally, to further illustrate the capabilities of ERP, we tested our algorithm on three code generation benchmarks and outperformed the current state-of-the-art approach as well. Our code is publicly available at: https://github.com/xuefeng-cs/ERP.
Transformers are efficient hierarchical chemical graph learners
Oct 02, 2023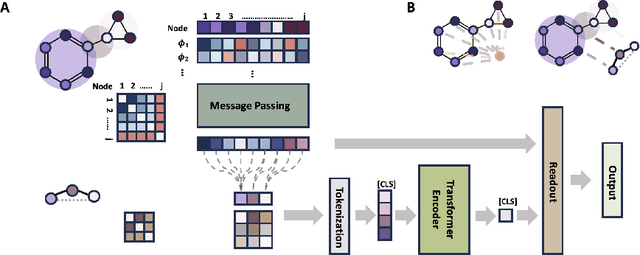
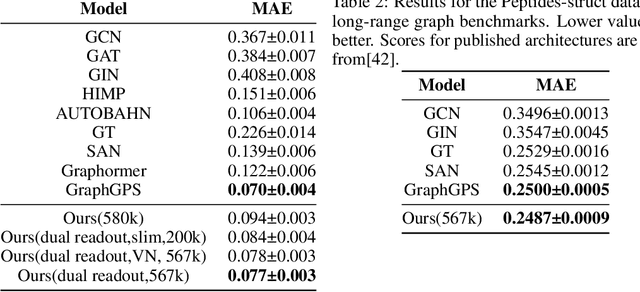

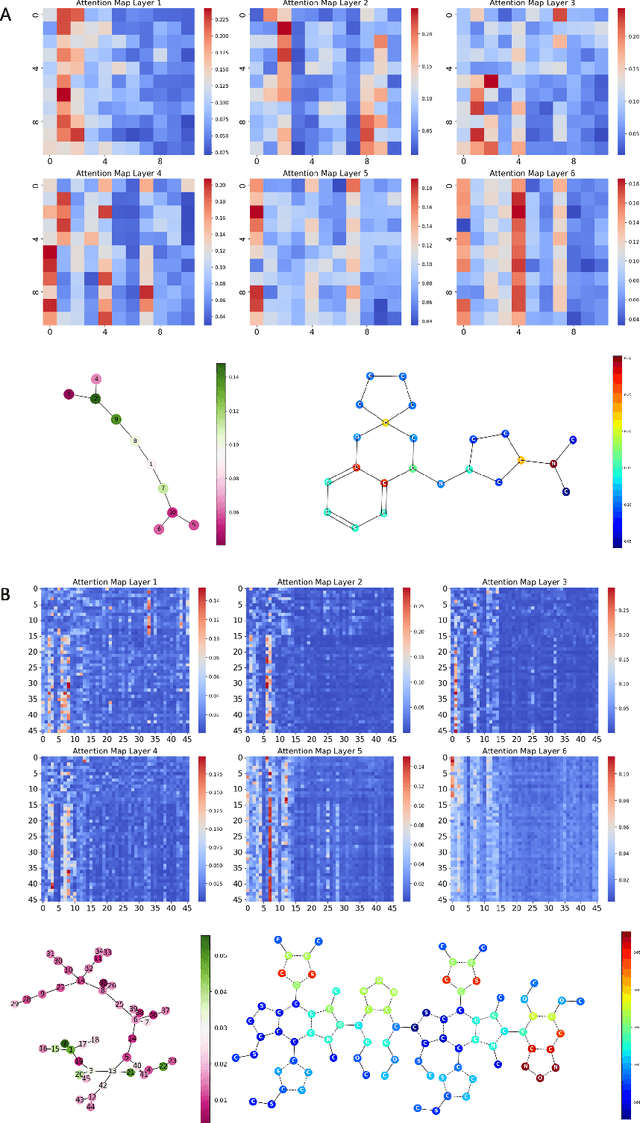
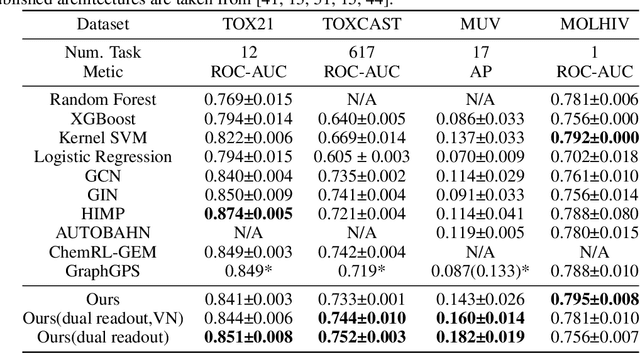
Abstract:Transformers, adapted from natural language processing, are emerging as a leading approach for graph representation learning. Contemporary graph transformers often treat nodes or edges as separate tokens. This approach leads to computational challenges for even moderately-sized graphs due to the quadratic scaling of self-attention complexity with token count. In this paper, we introduce SubFormer, a graph transformer that operates on subgraphs that aggregate information by a message-passing mechanism. This approach reduces the number of tokens and enhances learning long-range interactions. We demonstrate SubFormer on benchmarks for predicting molecular properties from chemical structures and show that it is competitive with state-of-the-art graph transformers at a fraction of the computational cost, with training times on the order of minutes on a consumer-grade graphics card. We interpret the attention weights in terms of chemical structures. We show that SubFormer exhibits limited over-smoothing and avoids over-squashing, which is prevalent in traditional graph neural networks.
Bilingual alignment transfers to multilingual alignment for unsupervised parallel text mining
Apr 15, 2021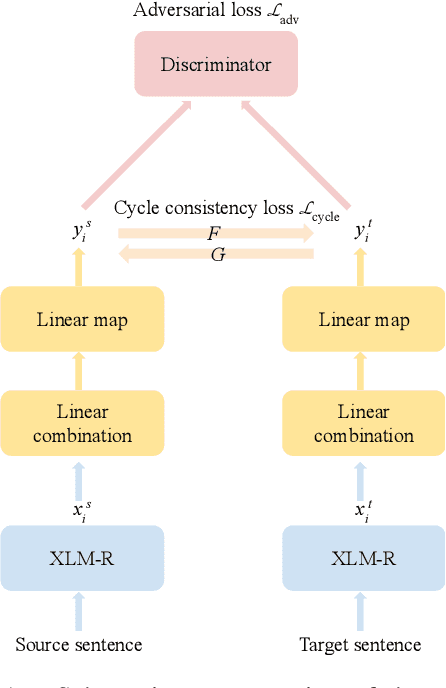
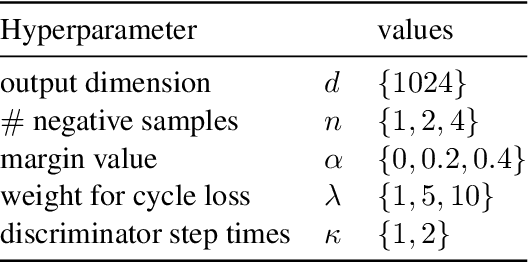

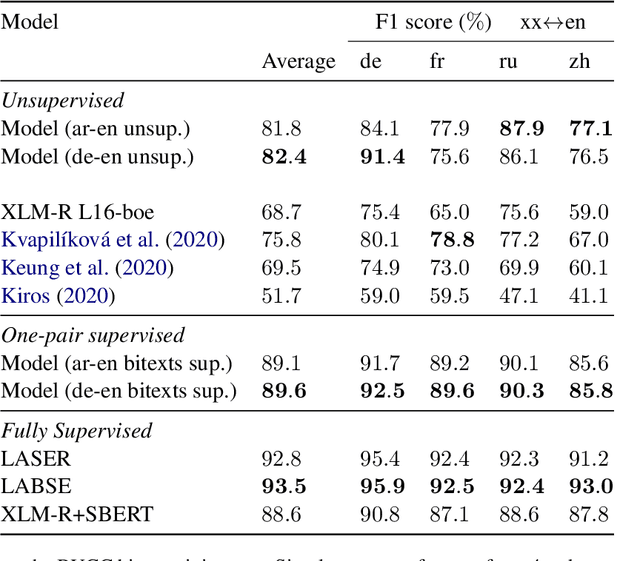
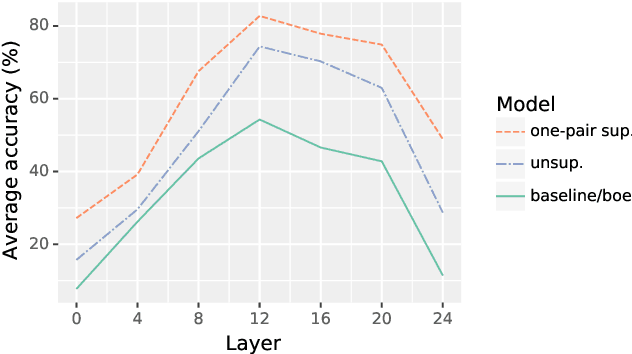
Abstract:This work presents methods for learning cross-lingual sentence representations using paired or unpaired bilingual texts. We hypothesize that the cross-lingual alignment strategy is transferable, and therefore a model trained to align only two languages can encode multilingually more aligned representations. And such transfer from bilingual alignment to multilingual alignment is a dual-pivot transfer from two pivot languages to other language pairs. To study this theory, we train an unsupervised model with unpaired sentences and another single-pair supervised model with bitexts, both based on the unsupervised language model XLM-R. The experiments evaluate the models as universal sentence encoders on the task of unsupervised bitext mining on two datasets, where the unsupervised model reaches the state of the art of unsupervised retrieval, and the alternative single-pair supervised model approaches the performance of multilingually supervised models. The results suggest that bilingual training techniques as proposed can be applied to get sentence representations with higher multilingual alignment.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge