Ananth Reddy Bhimireddy
Few-Shot Transfer Learning to improve Chest X-Ray pathology detection using limited triplets
Apr 16, 2022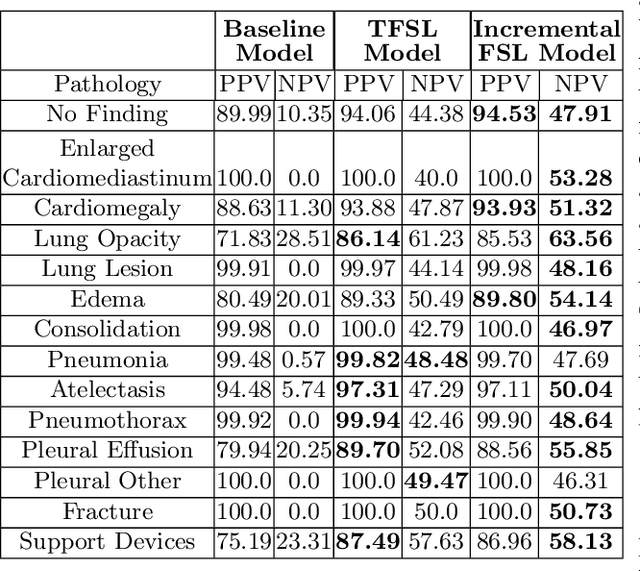
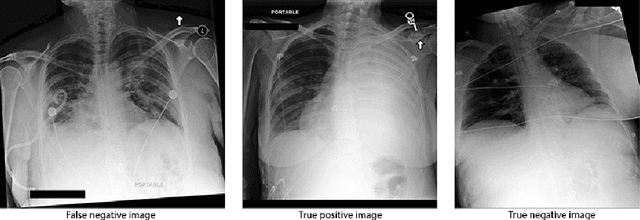
Abstract:Deep learning approaches applied to medical imaging have reached near-human or better-than-human performance on many diagnostic tasks. For instance, the CheXpert competition on detecting pathologies in chest x-rays has shown excellent multi-class classification performance. However, training and validating deep learning models require extensive collections of images and still produce false inferences, as identified by a human-in-the-loop. In this paper, we introduce a practical approach to improve the predictions of a pre-trained model through Few-Shot Learning (FSL). After training and validating a model, a small number of false inference images are collected to retrain the model using \textbf{\textit{Image Triplets}} - a false positive or false negative, a true positive, and a true negative. The retrained FSL model produces considerable gains in performance with only a few epochs and few images. In addition, FSL opens rapid retraining opportunities for human-in-the-loop systems, where a radiologist can relabel false inferences, and the model can be quickly retrained. We compare our retrained model performance with existing FSL approaches in medical imaging that train and evaluate models at once.
Reading Race: AI Recognises Patient's Racial Identity In Medical Images
Jul 21, 2021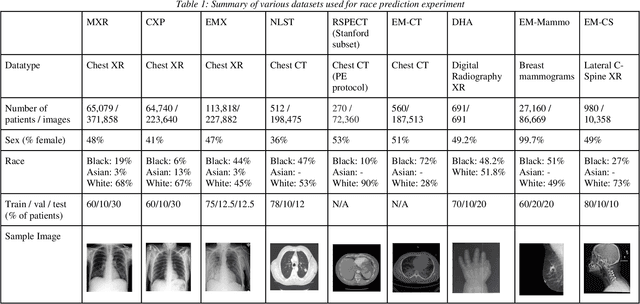
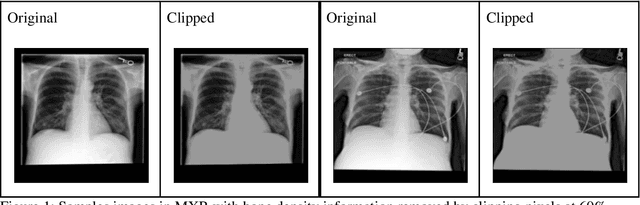
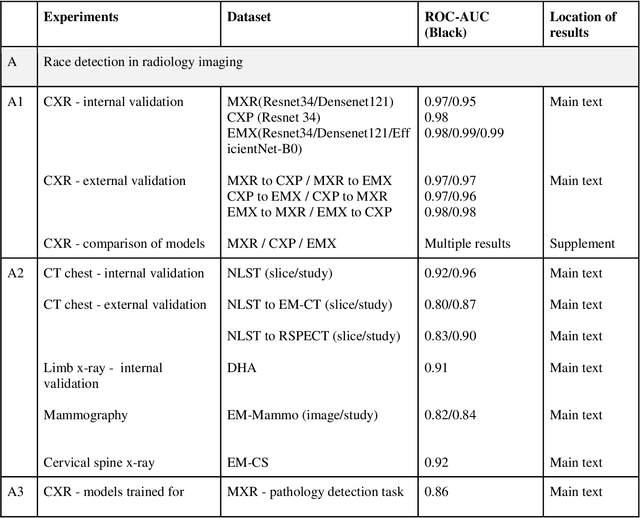
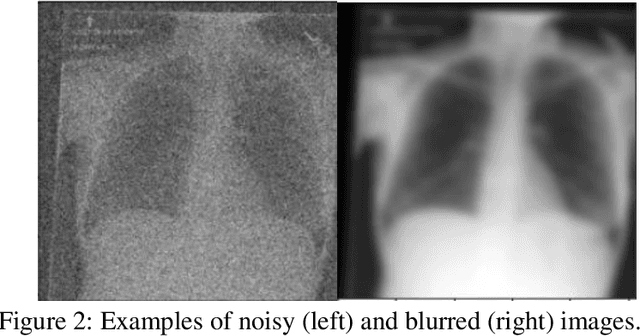
Abstract:Background: In medical imaging, prior studies have demonstrated disparate AI performance by race, yet there is no known correlation for race on medical imaging that would be obvious to the human expert interpreting the images. Methods: Using private and public datasets we evaluate: A) performance quantification of deep learning models to detect race from medical images, including the ability of these models to generalize to external environments and across multiple imaging modalities, B) assessment of possible confounding anatomic and phenotype population features, such as disease distribution and body habitus as predictors of race, and C) investigation into the underlying mechanism by which AI models can recognize race. Findings: Standard deep learning models can be trained to predict race from medical images with high performance across multiple imaging modalities. Our findings hold under external validation conditions, as well as when models are optimized to perform clinically motivated tasks. We demonstrate this detection is not due to trivial proxies or imaging-related surrogate covariates for race, such as underlying disease distribution. Finally, we show that performance persists over all anatomical regions and frequency spectrum of the images suggesting that mitigation efforts will be challenging and demand further study. Interpretation: We emphasize that model ability to predict self-reported race is itself not the issue of importance. However, our findings that AI can trivially predict self-reported race -- even from corrupted, cropped, and noised medical images -- in a setting where clinical experts cannot, creates an enormous risk for all model deployments in medical imaging: if an AI model secretly used its knowledge of self-reported race to misclassify all Black patients, radiologists would not be able to tell using the same data the model has access to.
Multi-label natural language processing to identify diagnosis and procedure codes from MIMIC-III inpatient notes
Mar 17, 2020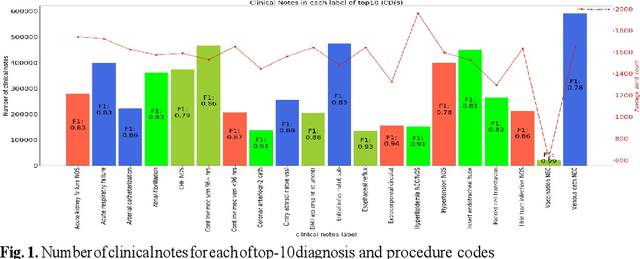
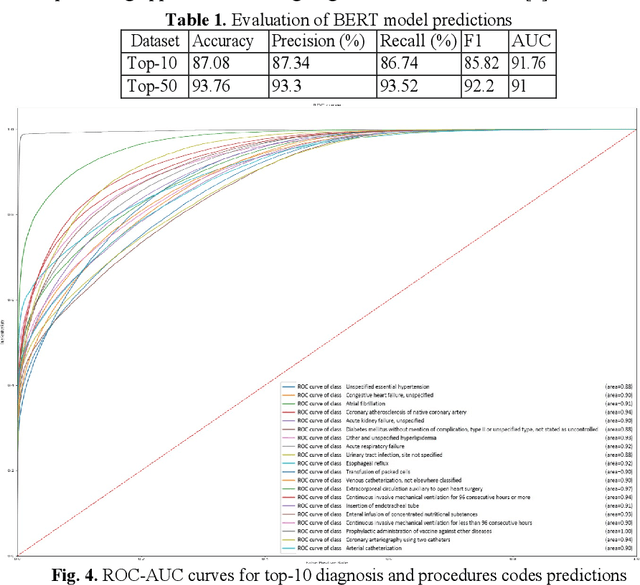
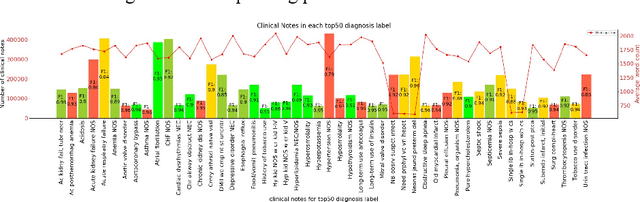
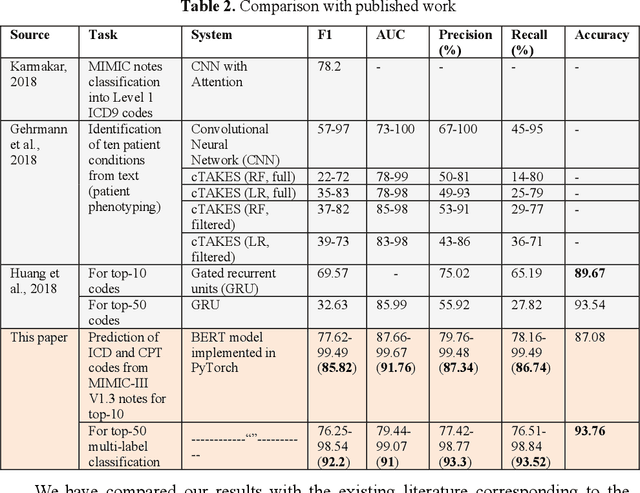
Abstract:In the United States, 25% or greater than 200 billion dollars of hospital spending accounts for administrative costs that involve services for medical coding and billing. With the increasing number of patient records, manual assignment of the codes performed is overwhelming, time-consuming and error-prone, causing billing errors. Natural language processing can automate the extraction of codes/labels from unstructured clinical notes, which can aid human coders to save time, increase productivity, and verify medical coding errors. Our objective is to identify appropriate diagnosis and procedure codes from clinical notes by performing multi-label classification. We used de-identified data of critical care patients from the MIMIC-III database and subset the data to select the ten (top-10) and fifty (top-50) most common diagnoses and procedures, which covers 47.45% and 74.12% of all admissions respectively. We implemented state-of-the-art Bidirectional Encoder Representations from Transformers (BERT) to fine-tune the language model on 80% of the data and validated on the remaining 20%. The model achieved an overall accuracy of 87.08%, an F1 score of 85.82%, and an AUC of 91.76% for top-10 codes. For the top-50 codes, our model achieved an overall accuracy of 93.76%, an F1 score of 92.24%, and AUC of 91%. When compared to previously published research, our model outperforms in predicting codes from the clinical text. We discuss approaches to generalize the knowledge discovery process of our MIMIC-BERT to other clinical notes. This can help human coders to save time, prevent backlogs, and additional costs due to coding errors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge