Yunsoo Kim
HH-SAE: Discovering and Steering Hierarchical Knowledge of Complex Manifolds
May 11, 2026Abstract:Rare semantic innovations in high-dimensional, mission-critical domains are often obscured by dense background contexts, a challenge we define as \textit{feature density conflict}. We introduce the \textbf{Hybrid Hierarchical SAE (HH-SAE)} to resolve this by factorizing manifolds into a nested hierarchy of \textbf{Contextual} ($L_0$), \textbf{Atomic} ($f_1$), and \textbf{Compository} ($f_2$) tiers. Evaluating across disparate manifolds, HH-SAE demonstrates superior resolution by \textbf{``fracturing'' administrative clinical labels into physiological modes} and achieving a peak \textbf{cross-domain zero-shot AUC of 0.9156 in fraud detection}. Path ablation confirms the architecture's structural necessity, revealing a 13.46\% utility collapse when contextual subtraction is removed. Finally, knowledge-steered synthesis achieves a +9.9\% AUPRC lift over state-of-the-art generators, proving that HH-SAE effectively prioritizes high-order mechanistic innovation over environmental proxies to enable high-precision discovery in high-stakes environments.
A Regime Theory of Controller Class Selection for LLM Action Decisions
May 07, 2026Abstract:Deployed language and vision-language models must decide, on each input, whether to answer directly, retrieve evidence, defer to a stronger model, or abstain. Contrary to the common monotonicity intuition, greater per-input expressivity is not uniformly beneficial in finite samples: under identical strict cross-validation, different benchmarks prefer different controller classes. This reflects a finite-sample limitation of instance-level uncertainty signals, which can be exhausted at a distribution-dependent scale. We organize controllers into a nested lattice of four classes: fixed actions, partition routers, instance-level controllers, and prior-gated controllers, ordered by complexity. We prove a regime theory that turns three data-estimable bottlenecks into a class choice: how much improvement is possible beyond the best fixed action, whether there are enough samples for instance-level controllers to make reliable decisions, and how much improvement a coarse partition router can recover when instance-level signal is unreliable. The resulting Bernstein-tight threshold has a matching information-theoretic lower bound, and strict nested cross-validation provably selects a near-best class. Across SMS-Spam, HallusionBench, A-OKVQA, and FOLIO, the predicted class matches the empirical winner; the prior-gated controller wins on TextVQA when OCR tokens supply a label-free prediction-time prior. Code is available at https://github.com/Anonymous-Awesome-Submissions/Regime-Theory.
E2EGS: Event-to-Edge Gaussian Splatting for Pose-Free 3D Reconstruction
Mar 16, 2026Abstract:The emergence of neural radiance fields (NeRF) and 3D Gaussian splatting (3DGS) has advanced novel view synthesis (NVS). These methods, however, require high-quality RGB inputs and accurate corresponding poses, limiting robustness under real-world conditions such as fast camera motion or adverse lighting. Event cameras, which capture brightness changes at each pixel with high temporal resolution and wide dynamic range, enable precise sensing of dynamic scenes and offer a promising solution. However, existing event-based NVS methods either assume known poses or rely on depth estimation models that are bounded by their initial observations, failing to generalize as the camera traverses previously unseen regions. We present E2EGS, a pose-free framework operating solely on event streams. Our key insight is that edge information provides rich structural cues essential for accurate trajectory estimation and high-quality NVS. To extract edges from noisy event streams, we exploit the distinct spatio-temporal characteristics of edges and non-edge regions. The event camera's movement induces consistent events along edges, while non-edge regions produce sparse noise. We leverage this through a patch-based temporal coherence analysis that measures local variance to extract edges while robustly suppressing noise. The extracted edges guide structure-aware Gaussian initialization and enable edge-weighted losses throughout initialization, tracking, and bundle adjustment. Extensive experiments on both synthetic and real datasets demonstrate that E2EGS achieves superior reconstruction quality and trajectory accuracy, establishing a fully pose-free paradigm for event-based 3D reconstruction.
LoV3D: Grounding Cognitive Prognosis Reasoning in Longitudinal 3D Brain MRI via Regional Volume Assessments
Mar 14, 2026Abstract:Longitudinal brain MRI is essential for characterizing the progression of neurological diseases such as Alzheimer's disease assessment. However, current deep-learning tools fragment this process: classifiers reduce a scan to a label, volumetric pipelines produce uninterpreted measurements, and vision-language models (VLMs) may generate fluent but potentially hallucinated conclusions. We present LoV3D, a pipeline for training 3D vision-language models, which reads longitudinal T1-weighted brain MRI, produces a region-level anatomical assessment, conducts longitudinal comparison with the prior scan, and finally outputs a three-class diagnosis (Cognitively Normal, Mild Cognitive Impairment, or Dementia) along with a synthesized diagnostic summary. The stepped pipeline grounds the final diagnosis by enforcing label consistency, longitudinal coherence, and biological plausibility, thereby reducing the risks of hallucinations. The training process introduces a clinically-weighted Verifier that scores candidate outputs automatically against normative references derived from standardized volume metrics, driving Direct Preference Optimization without a single human annotation. On a subject-level held-out ADNI test set (479 scans, 258 subjects), LoV3D achieves 93.7% three-class diagnostic accuracy (+34.8% over the no-grounding baseline), 97.2% on two-class diagnosis accuracy (+4% over the SOTA) and 82.6% region-level anatomical classification accuracy (+33.1% over VLM baselines). Zero-shot transfer yields 95.4% on MIRIAD (100% Dementia recall) and 82.9% three-class accuracy on AIBL, confirming high generalizability across sites, scanners, and populations. Code is available at https://github.com/Anonymous-TEVC/LoV-3D.
Paper Title: LoV3D: Grounding Cognitive Prognosis Reasoning in Longitudinal 3D Brain MRI via Regional Volume Assessments
Mar 12, 2026Abstract:Longitudinal brain MRI is essential for characterizing the progression of neurological diseases such as Alzheimer's disease assessment. However, current deep-learning tools fragment this process: classifiers reduce a scan to a label, volumetric pipelines produce uninterpreted measurements, and vision-language models (VLMs) may generate fluent but potentially hallucinated conclusions. We present LoV3D, a pipeline for training 3D vision-language models, which reads longitudinal T1-weighted brain MRI, produces a region-level anatomical assessment, conducts longitudinal comparison with the prior scan, and finally outputs a three-class diagnosis (Cognitively Normal, Mild Cognitive Impairment, or Dementia) along with a synthesized diagnostic summary. The stepped pipeline grounds the final diagnosis by enforcing label consistency, longitudinal coherence, and biological plausibility, thereby reducing the risks of hallucinations. The training process introduces a clinically-weighted Verifier that scores candidate outputs automatically against normative references derived from standardized volume metrics, driving Direct Preference Optimization without a single human annotation. On a subject-level held-out ADNI test set (479 scans, 258 subjects), LoV3D achieves 93.7% three-class diagnostic accuracy (+34.8% over the no-grounding baseline), 97.2% on two-class diagnosis accuracy (+4% over the SOTA) and 82.6% region-level anatomical classification accuracy (+33.1% over VLM baselines). Zero-shot transfer yields 95.4% on MIRIAD (100% Dementia recall) and 82.9% three-class accuracy on AIBL, confirming high generalizability across sites, scanners, and populations. Code is available at https://github.com/Anonymous-TEVC/LoV-3D.
BioHopR: A Benchmark for Multi-Hop, Multi-Answer Reasoning in Biomedical Domain
May 28, 2025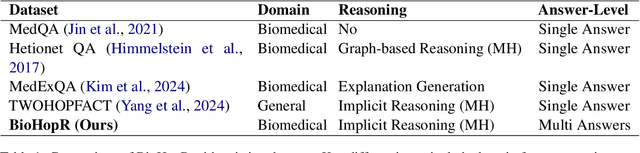
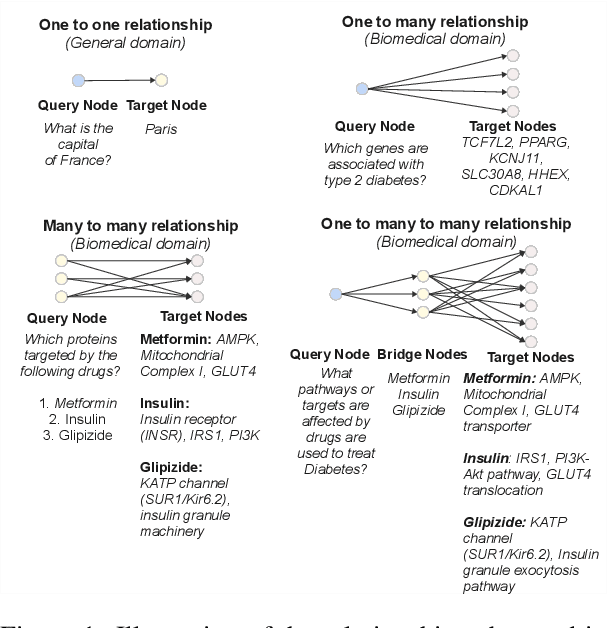

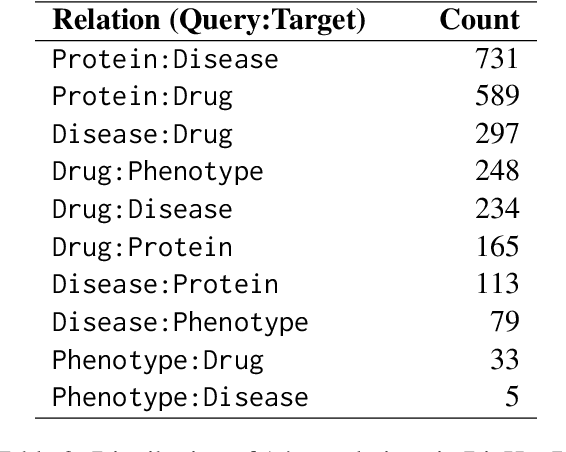
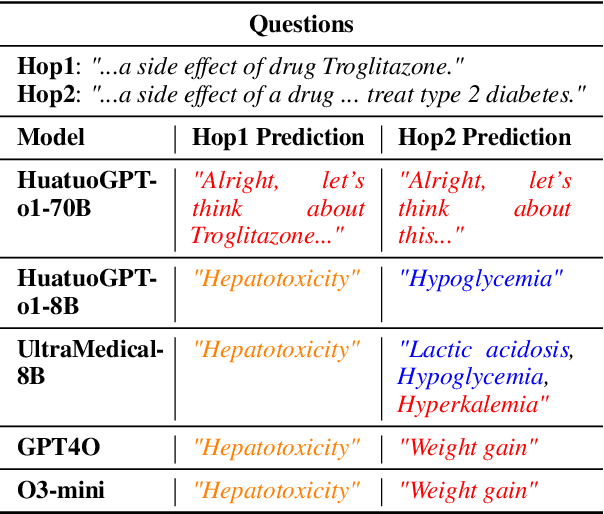
Abstract:Biomedical reasoning often requires traversing interconnected relationships across entities such as drugs, diseases, and proteins. Despite the increasing prominence of large language models (LLMs), existing benchmarks lack the ability to evaluate multi-hop reasoning in the biomedical domain, particularly for queries involving one-to-many and many-to-many relationships. This gap leaves the critical challenges of biomedical multi-hop reasoning underexplored. To address this, we introduce BioHopR, a novel benchmark designed to evaluate multi-hop, multi-answer reasoning in structured biomedical knowledge graphs. Built from the comprehensive PrimeKG, BioHopR includes 1-hop and 2-hop reasoning tasks that reflect real-world biomedical complexities. Evaluations of state-of-the-art models reveal that O3-mini, a proprietary reasoning-focused model, achieves 37.93% precision on 1-hop tasks and 14.57% on 2-hop tasks, outperforming proprietary models such as GPT4O and open-source biomedical models including HuatuoGPT-o1-70B and Llama-3.3-70B. However, all models exhibit significant declines in multi-hop performance, underscoring the challenges of resolving implicit reasoning steps in the biomedical domain. By addressing the lack of benchmarks for multi-hop reasoning in biomedical domain, BioHopR sets a new standard for evaluating reasoning capabilities and highlights critical gaps between proprietary and open-source models while paving the way for future advancements in biomedical LLMs.
Look & Mark: Leveraging Radiologist Eye Fixations and Bounding boxes in Multimodal Large Language Models for Chest X-ray Report Generation
May 28, 2025Abstract:Recent advancements in multimodal Large Language Models (LLMs) have significantly enhanced the automation of medical image analysis, particularly in generating radiology reports from chest X-rays (CXR). However, these models still suffer from hallucinations and clinically significant errors, limiting their reliability in real-world applications. In this study, we propose Look & Mark (L&M), a novel grounding fixation strategy that integrates radiologist eye fixations (Look) and bounding box annotations (Mark) into the LLM prompting framework. Unlike conventional fine-tuning, L&M leverages in-context learning to achieve substantial performance gains without retraining. When evaluated across multiple domain-specific and general-purpose models, L&M demonstrates significant gains, including a 1.2% improvement in overall metrics (A.AVG) for CXR-LLaVA compared to baseline prompting and a remarkable 9.2% boost for LLaVA-Med. General-purpose models also benefit from L&M combined with in-context learning, with LLaVA-OV achieving an 87.3% clinical average performance (C.AVG)-the highest among all models, even surpassing those explicitly trained for CXR report generation. Expert evaluations further confirm that L&M reduces clinically significant errors (by 0.43 average errors per report), such as false predictions and omissions, enhancing both accuracy and reliability. These findings highlight L&M's potential as a scalable and efficient solution for AI-assisted radiology, paving the way for improved diagnostic workflows in low-resource clinical settings.
IHC-LLMiner: Automated extraction of tumour immunohistochemical profiles from PubMed abstracts using large language models
Apr 01, 2025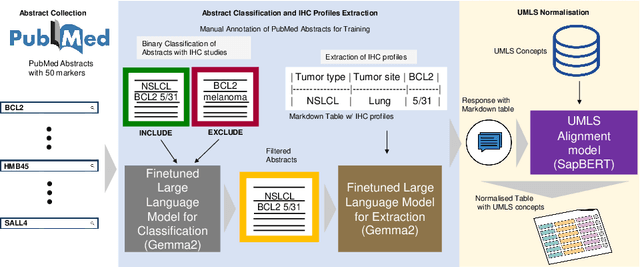
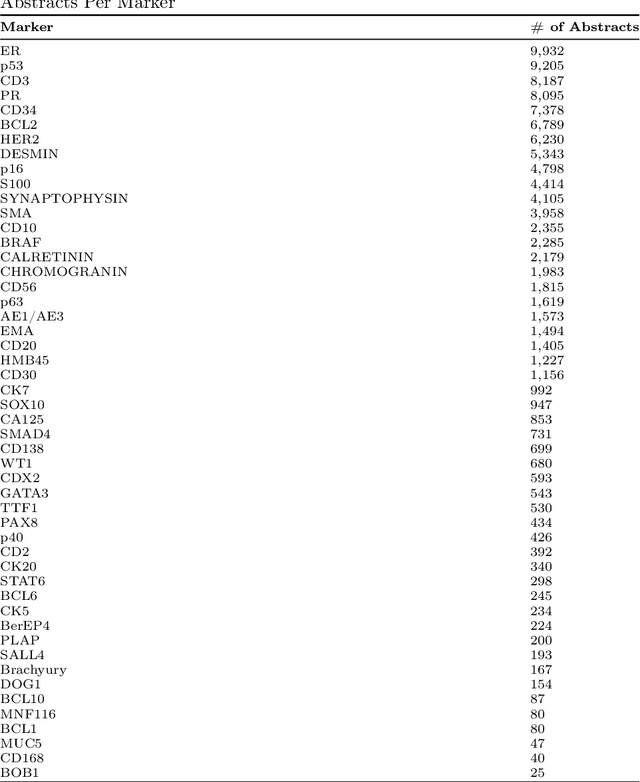
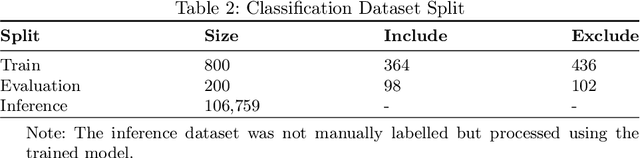
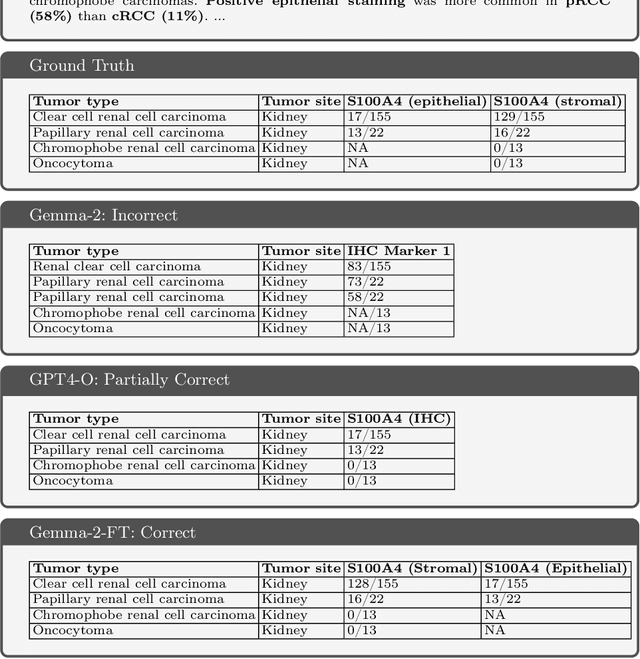
Abstract:Immunohistochemistry (IHC) is essential in diagnostic pathology and biomedical research, offering critical insights into protein expression and tumour biology. This study presents an automated pipeline, IHC-LLMiner, for extracting IHC-tumour profiles from PubMed abstracts, leveraging advanced biomedical text mining. There are two subtasks: abstract classification (include/exclude as relevant) and IHC-tumour profile extraction on relevant included abstracts. The best-performing model, "Gemma-2 finetuned", achieved 91.5% accuracy and an F1 score of 91.4, outperforming GPT4-O by 9.5% accuracy with 5.9 times faster inference time. From an initial dataset of 107,759 abstracts identified for 50 immunohistochemical markers, the classification task identified 30,481 relevant abstracts (Include) using the Gemma-2 finetuned model. For IHC-tumour profile extraction, the Gemma-2 finetuned model achieved the best performance with 63.3% Correct outputs. Extracted IHC-tumour profiles (tumour types and markers) were normalised to Unified Medical Language System (UMLS) concepts to ensure consistency and facilitate IHC-tumour profile landscape analysis. The extracted IHC-tumour profiles demonstrated excellent concordance with available online summary data and provided considerable added value in terms of both missing IHC-tumour profiles and quantitative assessments. Our proposed LLM based pipeline provides a practical solution for large-scale IHC-tumour profile data mining, enhancing the accessibility and utility of such data for research and clinical applications as well as enabling the generation of quantitative and structured data to support cancer-specific knowledge base development. Models and training datasets are available at https://github.com/knowlab/IHC-LLMiner.
Harnessing Knowledge Retrieval with Large Language Models for Clinical Report Error Correction
Jun 21, 2024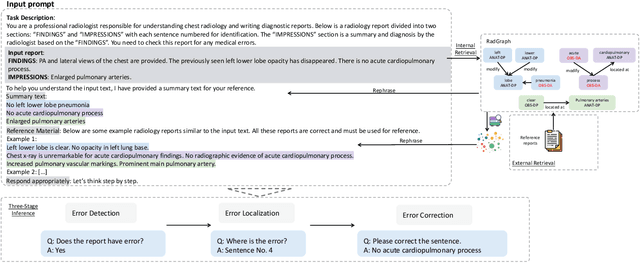



Abstract:This study proposes an approach for error correction in clinical radiology reports, leveraging large language models (LLMs) and retrieval-augmented generation (RAG) techniques. The proposed framework employs internal and external retrieval mechanisms to extract relevant medical entities and relations from the report and external knowledge sources. A three-stage inference process is introduced, decomposing the task into error detection, localization, and correction subtasks, which enhances the explainability and performance of the system. The effectiveness of the approach is evaluated using a benchmark dataset created by corrupting real-world radiology reports with realistic errors, guided by domain experts. Experimental results demonstrate the benefits of the proposed methods, with the combination of internal and external retrieval significantly improving the accuracy of error detection, localization, and correction across various state-of-the-art LLMs. The findings contribute to the development of more robust and reliable error correction systems for clinical documentation.
Chain-of-Though (CoT) prompting strategies for medical error detection and correction
Jun 13, 2024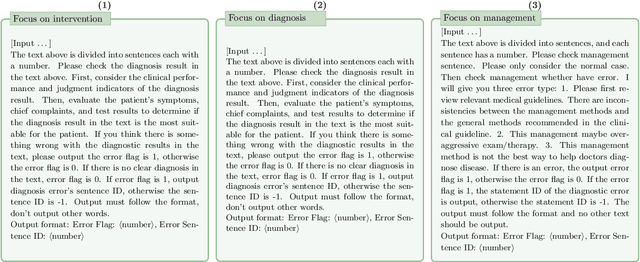
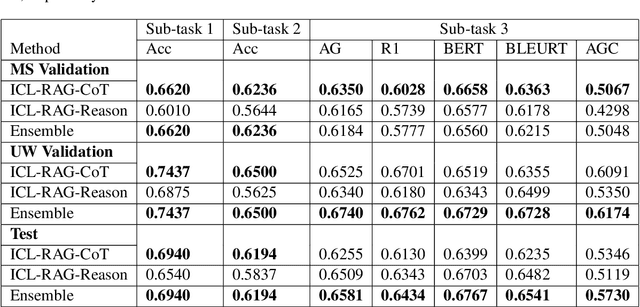

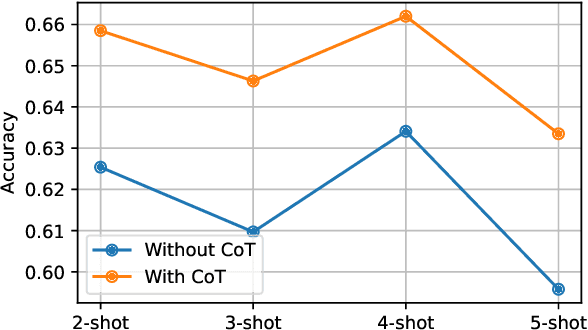
Abstract:This paper describes our submission to the MEDIQA-CORR 2024 shared task for automatically detecting and correcting medical errors in clinical notes. We report results for three methods of few-shot In-Context Learning (ICL) augmented with Chain-of-Thought (CoT) and reason prompts using a large language model (LLM). In the first method, we manually analyse a subset of train and validation dataset to infer three CoT prompts by examining error types in the clinical notes. In the second method, we utilise the training dataset to prompt the LLM to deduce reasons about their correctness or incorrectness. The constructed CoTs and reasons are then augmented with ICL examples to solve the tasks of error detection, span identification, and error correction. Finally, we combine the two methods using a rule-based ensemble method. Across the three sub-tasks, our ensemble method achieves a ranking of 3rd for both sub-task 1 and 2, while securing 7th place in sub-task 3 among all submissions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge