Xurui Jin
Explaining Graph Neural Networks via Non-parametric Subgraph Matching
Jan 07, 2023



Abstract:The great success in graph neural networks (GNNs) provokes the question about explainability: Which fraction of the input graph is the most determinant of the prediction? Particularly, parametric explainers prevail in existing approaches because of their stronger capability to decipher the black-box (i.e., the target GNN). In this paper, based on the observation that graphs typically share some joint motif patterns, we propose a novel non-parametric subgraph matching framework, dubbed MatchExplainer, to explore explanatory subgraphs. It couples the target graph with other counterpart instances and identifies the most crucial joint substructure by minimizing the node corresponding-based distance. Moreover, we note that present graph sampling or node-dropping methods usually suffer from the false positive sampling problem. To ameliorate that issue, we design a new augmentation paradigm named MatchDrop. It takes advantage of MatchExplainer to fix the most informative portion of the graph and merely operates graph augmentations on the rest less informative part. We conduct extensive experiments on both synthetic and real-world datasets and show the effectiveness of our MatchExplainer by outperforming all parametric baselines with significant margins. Additional results also demonstrate that our MatchDrop is a general scheme to be equipped with GNNs for enhanced performance.
A Score-based Geometric Model for Molecular Dynamics Simulations
Apr 19, 2022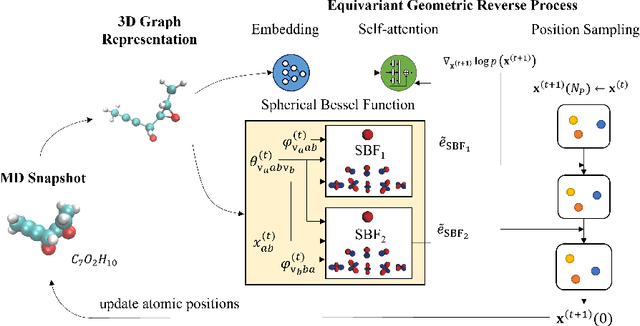
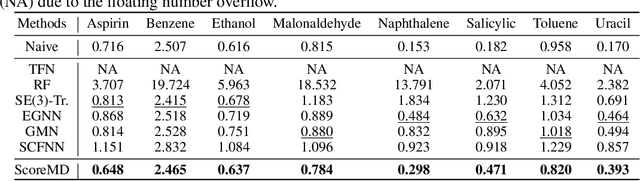
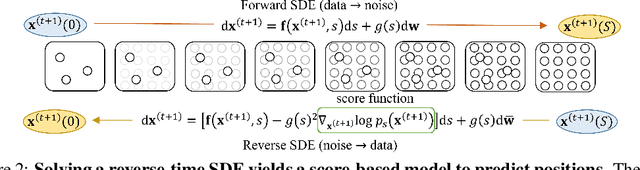

Abstract:Molecular dynamics (MD) has long been the \emph{de facto} choice for modeling complex atomistic systems from first principles, and recently deep learning become a popular way to accelerate it. Notwithstanding, preceding approaches depend on intermediate variables such as the potential energy or force fields to update atomic positions, which requires additional computations to perform back-propagation. To waive this requirement, we propose a novel model called ScoreMD by directly estimating the gradient of the log density of molecular conformations. Moreover, we analyze that diffusion processes highly accord with the principle of enhanced sampling in MD simulations, and is therefore a perfect match to our sequential conformation generation task. That is, ScoreMD perturbs the molecular structure with a conditional noise depending on atomic accelerations and employs conformations at previous timeframes as the prior distribution for sampling. Another challenge of modeling such a conformation generation process is that the molecule is kinetic instead of static, which no prior studies strictly consider. To solve this challenge, we introduce a equivariant geometric Transformer as a score function in the diffusion process to calculate the corresponding gradient. It incorporates the directions and velocities of atomic motions via 3D spherical Fourier-Bessel representations. With multiple architectural improvements, we outperforms state-of-the-art baselines on MD17 and isomers of C7O2H10. This research provides new insights into the acceleration of new material and drug discovery.
Prompt-Guided Injection of Conformation to Pre-trained Protein Model
Feb 07, 2022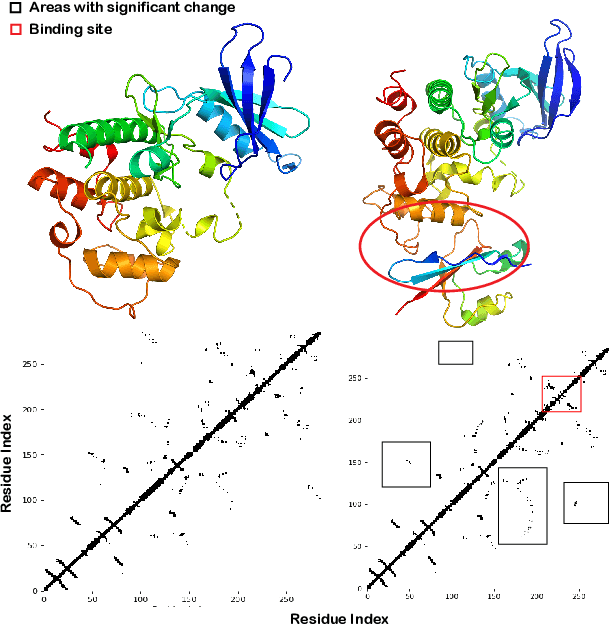
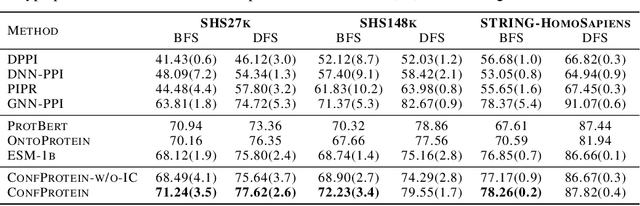
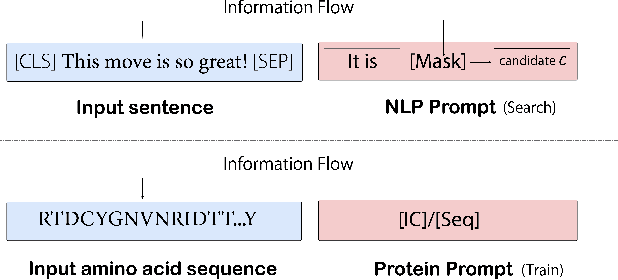

Abstract:Pre-trained protein models (PTPMs) represent a protein with one fixed embedding and thus are not capable for diverse tasks. For example, protein structures can shift, namely protein folding, between several conformations in various biological processes. To enable PTPMs to produce task-aware representations, we propose to learn interpretable, pluggable and extensible protein prompts as a way of injecting task-related knowledge into PTPMs. In this regard, prior PTPM optimization with the masked language modeling task can be interpreted as learning a sequence prompt (Seq prompt) that enables PTPMs to capture the sequential dependency between amino acids. To incorporate conformational knowledge to PTPMs, we propose an interaction-conformation prompt (IC prompt) that is learned through back-propagation with the protein-protein interaction task. As an instantiation, we present a conformation-aware pre-trained protein model that learns both sequence and interaction-conformation prompts in a multi-task setting. We conduct comprehensive experiments on nine protein datasets. Results confirm our expectation that using the sequence prompt does not hurt PTPMs' performance on sequence-related tasks while incorporating the interaction-conformation prompt significantly improves PTPMs' performance on tasks where conformational knowledge counts. We also show the learned prompts can be combined and extended to deal with new complex tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge