Vivek Muthurangu
Training deep learning based dynamic MR image reconstruction using synthetic fractals
Mar 31, 2026Abstract:Purpose: To investigate whether synthetically generated fractal data can be used to train deep learning (DL) models for dynamic MRI reconstruction, thereby avoiding the privacy, licensing, and availability limitations associated with cardiac MR training datasets. Methods: A training dataset was generated using quaternion Julia fractals to produce 2D+time images. Multi-coil MRI acquisition was simulated to generate paired fully sampled and radially undersampled k-space data. A 3D UNet deep artefact suppression model was trained using these fractal data (F-DL) and compared with an identical model trained on cardiac MRI data (CMR-DL). Both models were evaluated on prospectively acquired radial real-time cardiac MRI from 10 patients. Reconstructions were compared against compressed sensing(CS) and low-rank deep image prior (LR-DIP). All reconstrctuions were ranked for image quality, while ventricular volumes and ejection fraction were compared with reference breath-hold cine MRI. Results: There was no significant difference in qualitative ranking between F-DL and CMR-DL (p=0.9), while both outperformed CS and LR-DIP (p<0.001). Ventricular volumes and function derived from F-DL were similar to CMR-DL, showing no significant bias and accptable limits of agreement compared to reference cine imaging. However, LR-DIP had a signifcant bias (p=0.016) and wider lmits of agreement. Conclusion: DL models trained using synthetic fractal data can reconstruct real-time cardiac MRI with image quality and clinical measurements comparable to models trained on true cardiac MRI data. Fractal training data provide an open, scalable alternative to clinical datasets and may enable development of more generalisable DL reconstruction models for dynamic MRI.
MultiFlow: A unified deep learning framework for multi-vessel classification, segmentation and clustering of phase-contrast MRI validated on a multi-site single ventricle patient cohort
Feb 17, 2025



Abstract:This study presents a unified deep learning (DL) framework, MultiFlowSeg, for classification and segmentation of velocity-encoded phase-contrast magnetic resonance imaging data, and MultiFlowDTC for temporal clustering of flow phenotypes. Applied to the FORCE registry of Fontan procedure patients, MultiFlowSeg achieved 100% classification accuracy for the aorta, SVC, and IVC, and 94% for the LPA and RPA. It demonstrated robust segmentation with a median Dice score of 0.91 (IQR: 0.86-0.93). The automated pipeline processed registry data, achieving high segmentation success despite challenges like poor image quality and dextrocardia. Temporal clustering identified five distinct patient subgroups, with significant differences in clinical outcomes, including ejection fraction, exercise tolerance, liver disease, and mortality. These results demonstrate the potential of combining DL and time-varying flow data for improved CHD prognosis and personalized care.
Image2Flow: A hybrid image and graph convolutional neural network for rapid patient-specific pulmonary artery segmentation and CFD flow field calculation from 3D cardiac MRI data
Feb 28, 2024



Abstract:Computational fluid dynamics (CFD) can be used for evaluation of hemodynamics. However, its routine use is limited by labor-intensive manual segmentation, CFD mesh creation, and time-consuming simulation. This study aims to train a deep learning model to both generate patient-specific volume-meshes of the pulmonary artery from 3D cardiac MRI data and directly estimate CFD flow fields. This study used 135 3D cardiac MRIs from both a public and private dataset. The pulmonary arteries in the MRIs were manually segmented and converted into volume-meshes. CFD simulations were performed on ground truth meshes and interpolated onto point-point correspondent meshes to create the ground truth dataset. The dataset was split 85/10/15 for training, validation and testing. Image2Flow, a hybrid image and graph convolutional neural network, was trained to transform a pulmonary artery template to patient-specific anatomy and CFD values. Image2Flow was evaluated in terms of segmentation and accuracy of CFD predicted was assessed using node-wise comparisons. Centerline comparisons of Image2Flow and CFD simulations performed using machine learning segmentation were also performed. Image2Flow achieved excellent segmentation accuracy with a median Dice score of 0.9 (IQR: 0.86-0.92). The median node-wise normalized absolute error for pressure and velocity magnitude was 11.98% (IQR: 9.44-17.90%) and 8.06% (IQR: 7.54-10.41), respectively. Centerline analysis showed no significant difference between the Image2Flow and conventional CFD simulated on machine learning-generated volume-meshes. This proof-of-concept study has shown it is possible to simultaneously perform patient specific volume-mesh based segmentation and pressure and flow field estimation. Image2Flow completes segmentation and CFD in ~205ms, which ~7000 times faster than manual methods, making it more feasible in a clinical environment.
Investigating the use of publicly available natural videos to learn Dynamic MR image reconstruction
Nov 23, 2023Abstract:Purpose: To develop and assess a deep learning (DL) pipeline to learn dynamic MR image reconstruction from publicly available natural videos (Inter4K). Materials and Methods: Learning was performed for a range of DL architectures (VarNet, 3D UNet, FastDVDNet) and corresponding sampling patterns (Cartesian, radial, spiral) either from true multi-coil cardiac MR data (N=692) or from pseudo-MR data simulated from Inter4K natural videos (N=692). Real-time undersampled dynamic MR images were reconstructed using DL networks trained with cardiac data and natural videos, and compressed sensing (CS). Differences were assessed in simulations (N=104 datasets) in terms of MSE, PSNR, and SSIM and prospectively for cardiac (short axis, four chambers, N=20) and speech (N=10) data in terms of subjective image quality ranking, SNR and Edge sharpness. Friedman Chi Square tests with post-hoc Nemenyi analysis were performed to assess statistical significance. Results: For all simulation metrics, DL networks trained with cardiac data outperformed DL networks trained with natural videos, which outperformed CS (p<0.05). However, in prospective experiments DL reconstructions using both training datasets were ranked similarly (and higher than CS) and presented no statistical differences in SNR and Edge Sharpness for most conditions. Additionally, high SSIM was measured between the DL methods with cardiac data and natural videos (SSIM>0.85). Conclusion: The developed pipeline enabled learning dynamic MR reconstruction from natural videos preserving DL reconstruction advantages such as high quality fast and ultra-fast reconstructions while overcoming some limitations (data scarcity or sharing). The natural video dataset, code and pre-trained networks are made readily available on github. Key Words: real-time; dynamic MRI; deep learning; image reconstruction; machine learning;
Deep Learning Pipeline for Preprocessing and Segmenting Cardiac Magnetic Resonance of Single Ventricle Patients from an Image Registry
Mar 21, 2023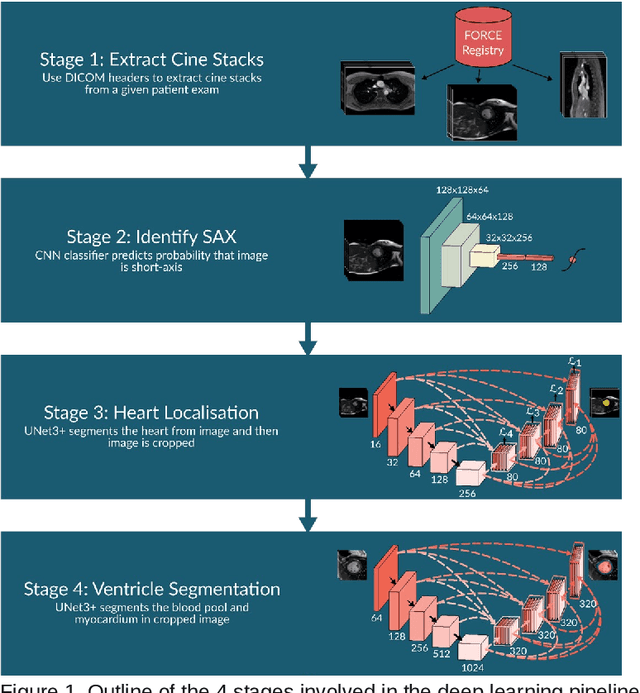
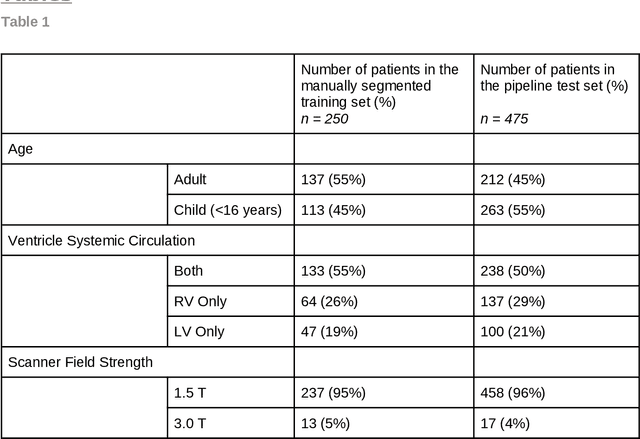
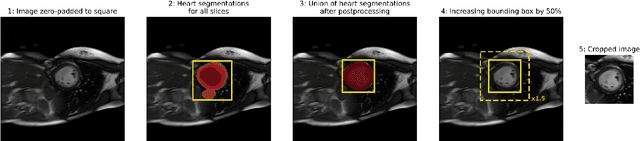
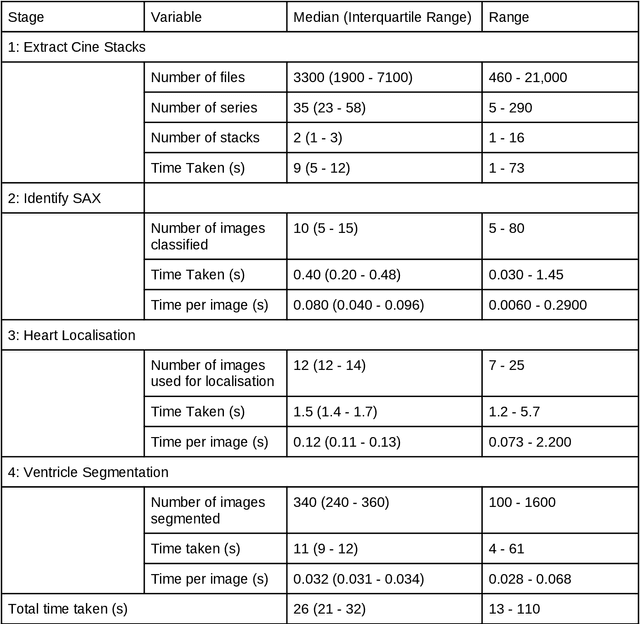
Abstract:Purpose: To develop and evaluate an end-to-end deep learning pipeline for segmentation and analysis of cardiac magnetic resonance images to provide core-lab processing for a multi-centre registry of Fontan patients. Materials and Methods: This retrospective study used training (n = 175), validation (n = 25) and testing (n = 50) cardiac magnetic resonance image exams collected from 13 institutions in the UK, US and Canada. The data was used to train and evaluate a pipeline containing three deep-learning models. The pipeline's performance was assessed on the Dice and IoU score between the automated and reference standard manual segmentation. Cardiac function values were calculated from both the automated and manual segmentation and evaluated using Bland-Altman analysis and paired t-tests. The overall pipeline was further evaluated qualitatively on 475 unseen patient exams. Results: For the 50 testing dataset, the pipeline achieved a median Dice score of 0.91 (0.89-0.94) for end-diastolic volume, 0.86 (0.82-0.89) for end-systolic volume, and 0.74 (0.70-0.77) for myocardial mass. The deep learning-derived end-diastolic volume, end-systolic volume, myocardial mass, stroke volume and ejection fraction had no statistical difference compared to the same values derived from manual segmentation with p values all greater than 0.05. For the 475 unseen patient exams, the pipeline achieved 68% adequate segmentation in both systole and diastole, 26% needed minor adjustments in either systole or diastole, 5% needed major adjustments, and the cropping model only failed in 0.4%. Conclusion: Deep learning pipeline can provide standardised 'core-lab' segmentation for Fontan patients. This pipeline can now be applied to the >4500 cardiac magnetic resonance exams currently in the FORCE registry as well as any new patients that are recruited.
GLADE: Gradient Loss Augmented Degradation Enhancement for Unpaired Super-Resolution of Anisotropic MRI
Mar 21, 2023Abstract:We present a novel approach to synthesise high-resolution isotropic 3D abdominal MR images, from anisotropic 3D images in an unpaired fashion. Using a modified CycleGAN architecture with a gradient mapping loss, we leverage disjoint patches from the high-resolution (in-plane) data of an anisotropic volume to enforce the network generator to increase the resolution of the low-resolution (through-plane) slices. This will enable accelerated whole-abdomen scanning with high-resolution isotropic images within short breath-hold times.
Deep neural networks for fast acquisition of aortic 3D pressure and velocity flow fields
Aug 25, 2022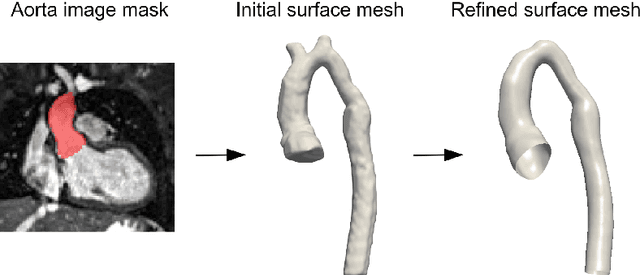
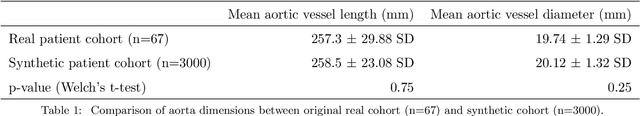

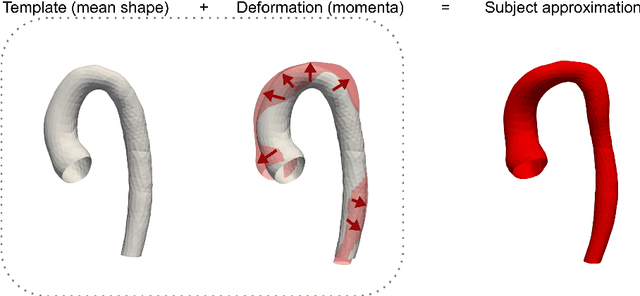
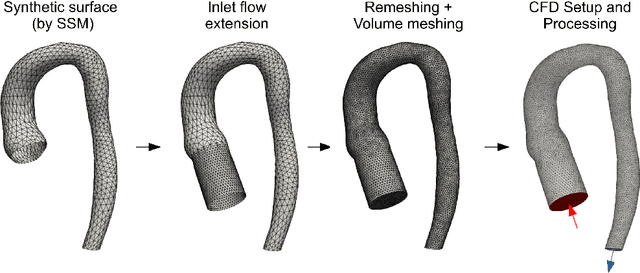
Abstract:Computational fluid dynamics (CFD) can be used to simulate vascular haemodynamics and analyse potential treatment options. CFD has shown to be beneficial in improving patient outcomes. However, the implementation of CFD for routine clinical use is yet to be realised. Barriers for CFD include high computational resources, specialist experience needed for designing simulation set-ups, and long processing times. The aim of this study was to explore the use of machine learning (ML) to replicate conventional aortic CFD with automatic and fast regression models. Data used to train/test the model comprised of 3,000 CFD simulations performed on synthetically generated 3D aortic shapes. These subjects were generated from a statistical shape model (SSM) built on real patient-specific aortas (N=67). Inference performed on 200 test shapes resulted in average errors of 6.01% +/-3.12 SD and 3.99% +/-0.93 SD for pressure and velocity, respectively. Our ML-based models performed CFD in ~0.075 seconds (4,000x faster than the solver). This study shows that results from conventional vascular CFD can be reproduced using ML at a much faster rate, in an automatic process, and with high accuracy.
FReSCO: Flow Reconstruction and Segmentation for low latency Cardiac Output monitoring using deep artifact suppression and segmentation
Mar 25, 2022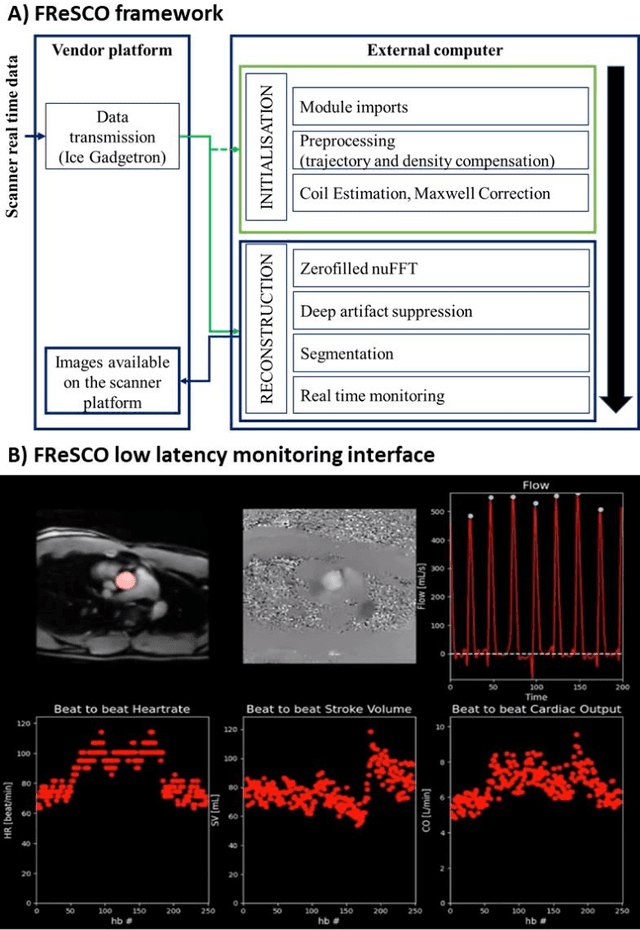
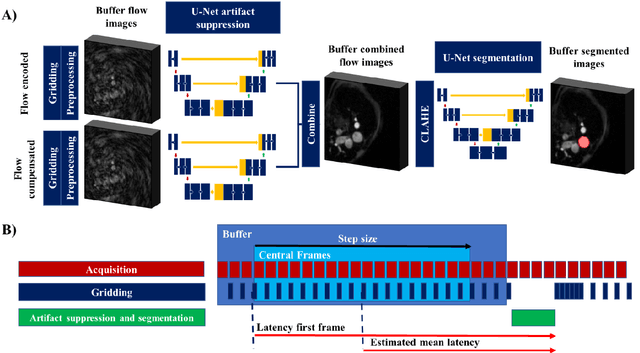
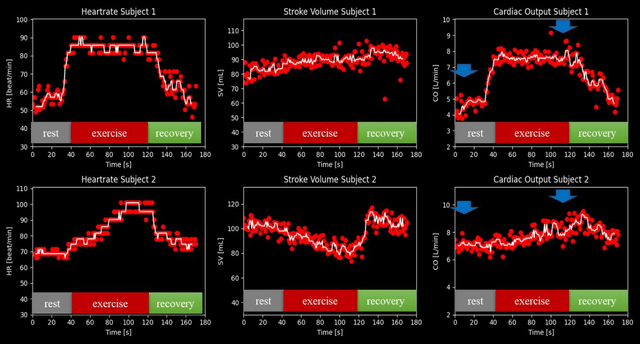
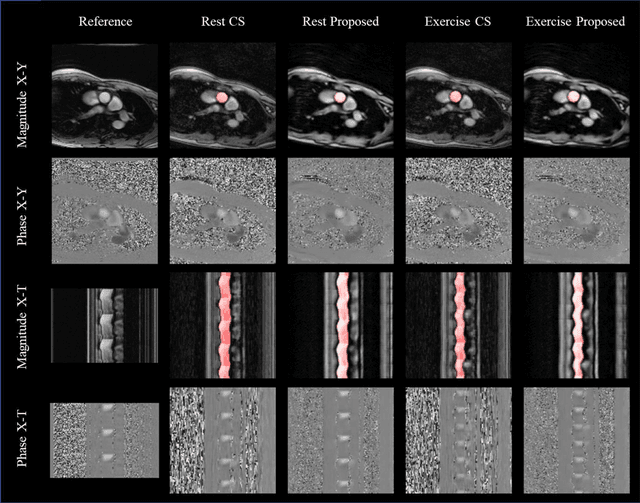
Abstract:Purpose: Real-time monitoring of cardiac output (CO) requires low latency reconstruction and segmentation of real-time phase contrast MR (PCMR), which has previously been difficult to perform. Here we propose a deep learning framework for 'Flow Reconstruction and Segmentation for low latency Cardiac Output monitoring' (FReSCO). Methods: Deep artifact suppression and segmentation U-Nets were independently trained. Breath hold spiral PCMR data (n=516) was synthetically undersampled using a variable density spiral sampling pattern and gridded to create aliased data for training of the artifact suppression U-net. A subset of the data (n=96) was segmented and used to train the segmentation U-net. Real-time spiral PCMR was prospectively acquired and then reconstructed and segmented using the trained models (FReSCO) at low latency at the scanner in 10 healthy subjects during rest, exercise and recovery periods. CO obtained via FReSCO was compared to a reference rest CO and rest and exercise Compressed Sensing (CS) CO. Results: FReSCO was demonstrated prospectively at the scanner. Beat-to-beat heartrate, stroke volume and CO could be visualized with a mean latency of 622ms. No significant differences were noted when compared to reference at rest (Bias = -0.21+-0.50 L/min, p=0.246) or CS at peak exercise (Bias=0.12+-0.48 L/min, p=0.458). Conclusion: FReSCO was successfully demonstrated for real-time monitoring of CO during exercise and could provide a convenient tool for assessment of the hemodynamic response to a range of stressors.
Machine Learning in Magnetic Resonance Imaging: Image Reconstruction
Dec 09, 2020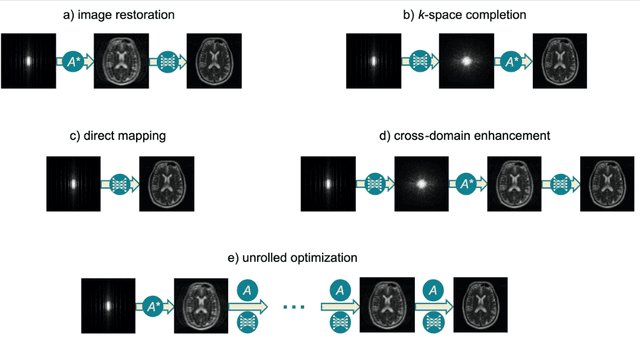
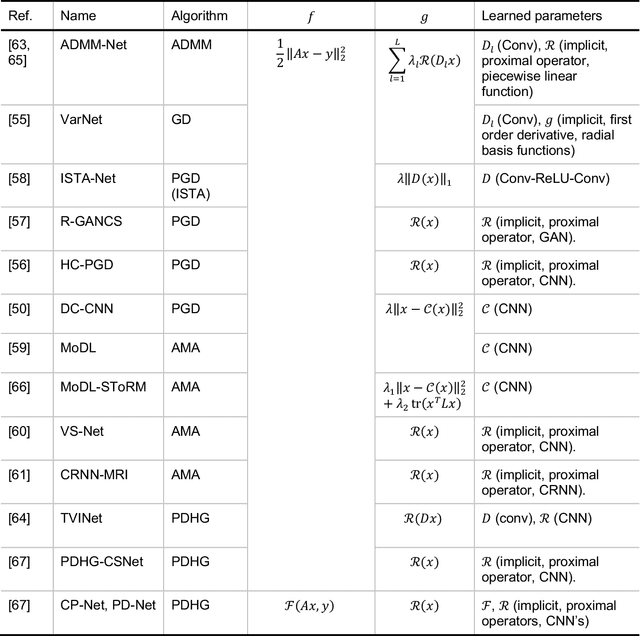
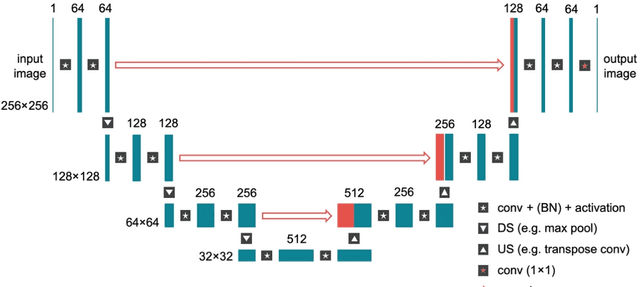
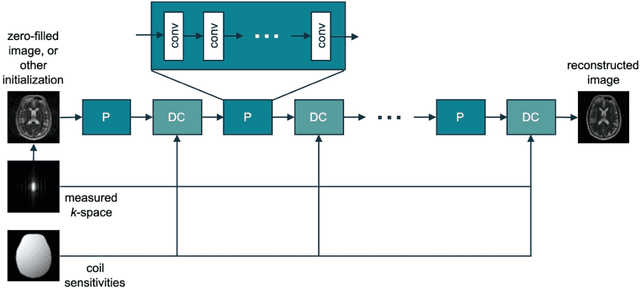
Abstract:Magnetic Resonance Imaging (MRI) plays a vital role in diagnosis, management and monitoring of many diseases. However, it is an inherently slow imaging technique. Over the last 20 years, parallel imaging, temporal encoding and compressed sensing have enabled substantial speed-ups in the acquisition of MRI data, by accurately recovering missing lines of k-space data. However, clinical uptake of vastly accelerated acquisitions has been limited, in particular in compressed sensing, due to the time-consuming nature of the reconstructions and unnatural looking images. Following the success of machine learning in a wide range of imaging tasks, there has been a recent explosion in the use of machine learning in the field of MRI image reconstruction. A wide range of approaches have been proposed, which can be applied in k-space and/or image-space. Promising results have been demonstrated from a range of methods, enabling natural looking images and rapid computation. In this review article we summarize the current machine learning approaches used in MRI reconstruction, discuss their drawbacks, clinical applications, and current trends.
Rapid Whole-Heart CMR with Single Volume Super-resolution
Dec 22, 2019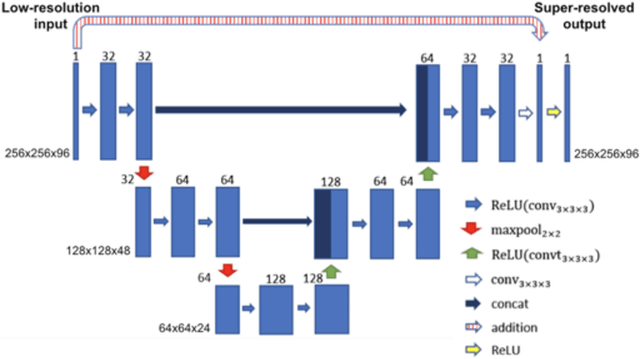
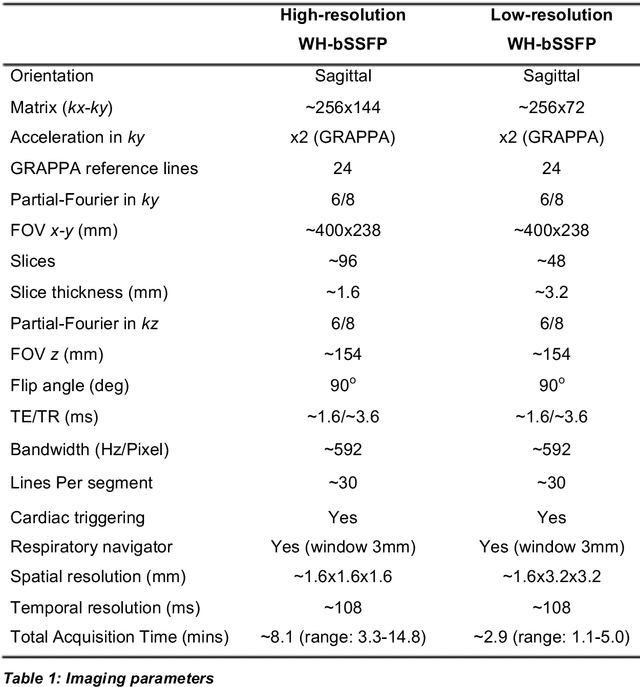
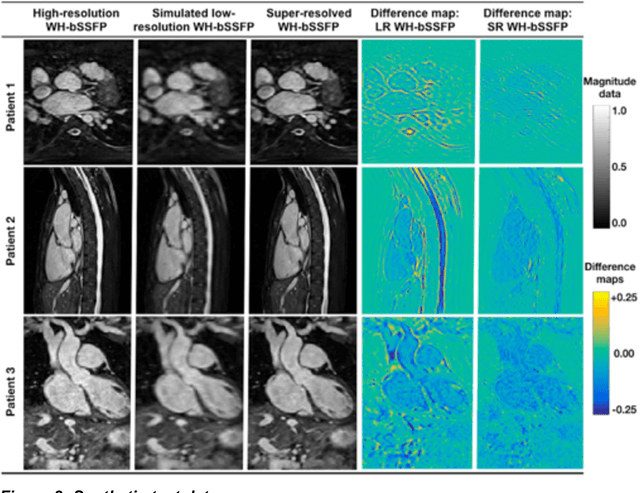
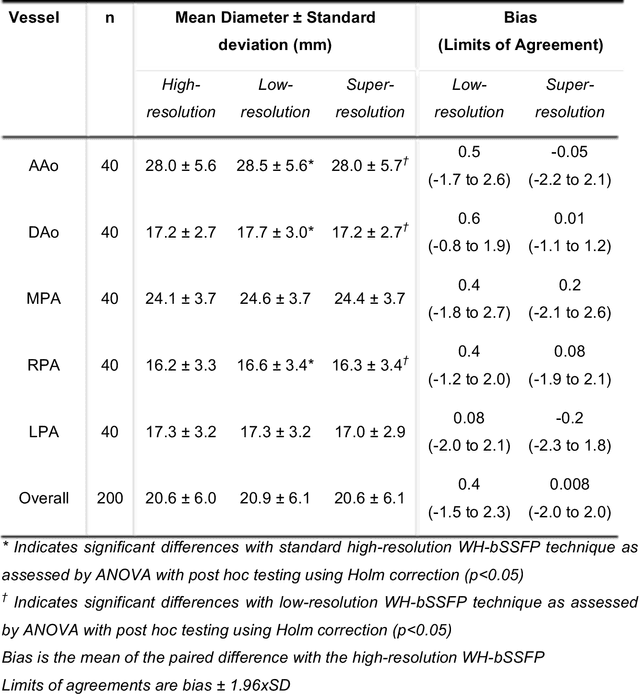
Abstract:Background: Three-dimensional, whole heart, balanced steady state free precession (WH-bSSFP) sequences provide delineation of intra-cardiac and vascular anatomy. However, they have long acquisition times. Here, we propose significant speed ups using a deep learning single volume super resolution reconstruction, to recover high resolution features from rapidly acquired low resolution WH-bSSFP images. Methods: A 3D residual U-Net was trained using synthetic data, created from a library of high-resolution WH-bSSFP images by simulating 0.5 slice resolution and 0.5 phase resolution. The trained network was validated with synthetic test data, as well as prospective low-resolution data. Results: Synthetic low-resolution data had significantly better image quality after super-resolution reconstruction. Qualitative image scores showed super-resolved images had better edge sharpness, fewer residual artefacts and less image distortion than low-resolution images, with similar scores to high-resolution data. Quantitative image scores showed super-resolved images had significantly better edge sharpness than low-resolution or high-resolution images, with significantly better signal-to-noise ratio than high-resolution data. Vessel diameters measurements showed over-estimation in the low-resolution measurements, compared to the high-resolution data. No significant differences and no bias was found in the super-resolution measurements. Conclusion: This paper demonstrates the potential of using a residual U-Net for super-resolution reconstruction of rapidly acquired low-resolution whole heart bSSFP data within a clinical setting. The resulting network can be applied very quickly, making these techniques particularly appealing within busy clinical workflow. Thus, we believe that this technique may help speed up whole heart CMR in clinical practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge