Trey Mullikin
Brain Tumor Segmentation (BraTS) Challenge 2024: Meningioma Radiotherapy Planning Automated Segmentation
May 28, 2024Abstract:The 2024 Brain Tumor Segmentation Meningioma Radiotherapy (BraTS-MEN-RT) challenge aims to advance automated segmentation algorithms using the largest known multi-institutional dataset of radiotherapy planning brain MRIs with expert-annotated target labels for patients with intact or post-operative meningioma that underwent either conventional external beam radiotherapy or stereotactic radiosurgery. Each case includes a defaced 3D post-contrast T1-weighted radiotherapy planning MRI in its native acquisition space, accompanied by a single-label "target volume" representing the gross tumor volume (GTV) and any at-risk post-operative site. Target volume annotations adhere to established radiotherapy planning protocols, ensuring consistency across cases and institutions. For pre-operative meningiomas, the target volume encompasses the entire GTV and associated nodular dural tail, while for post-operative cases, it includes at-risk resection cavity margins as determined by the treating institution. Case annotations were reviewed and approved by expert neuroradiologists and radiation oncologists. Participating teams will develop, containerize, and evaluate automated segmentation models using this comprehensive dataset. Model performance will be assessed using the lesion-wise Dice Similarity Coefficient and the 95% Hausdorff distance. The top-performing teams will be recognized at the Medical Image Computing and Computer Assisted Intervention Conference in October 2024. BraTS-MEN-RT is expected to significantly advance automated radiotherapy planning by enabling precise tumor segmentation and facilitating tailored treatment, ultimately improving patient outcomes.
A personalized Uncertainty Quantification framework for patient survival models: estimating individual uncertainty of patients with metastatic brain tumors in the absence of ground truth
Nov 28, 2023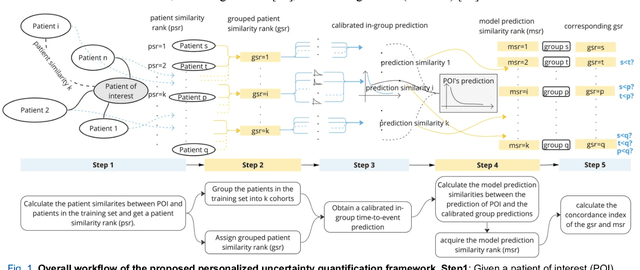
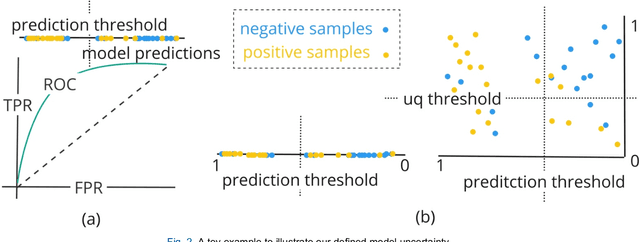
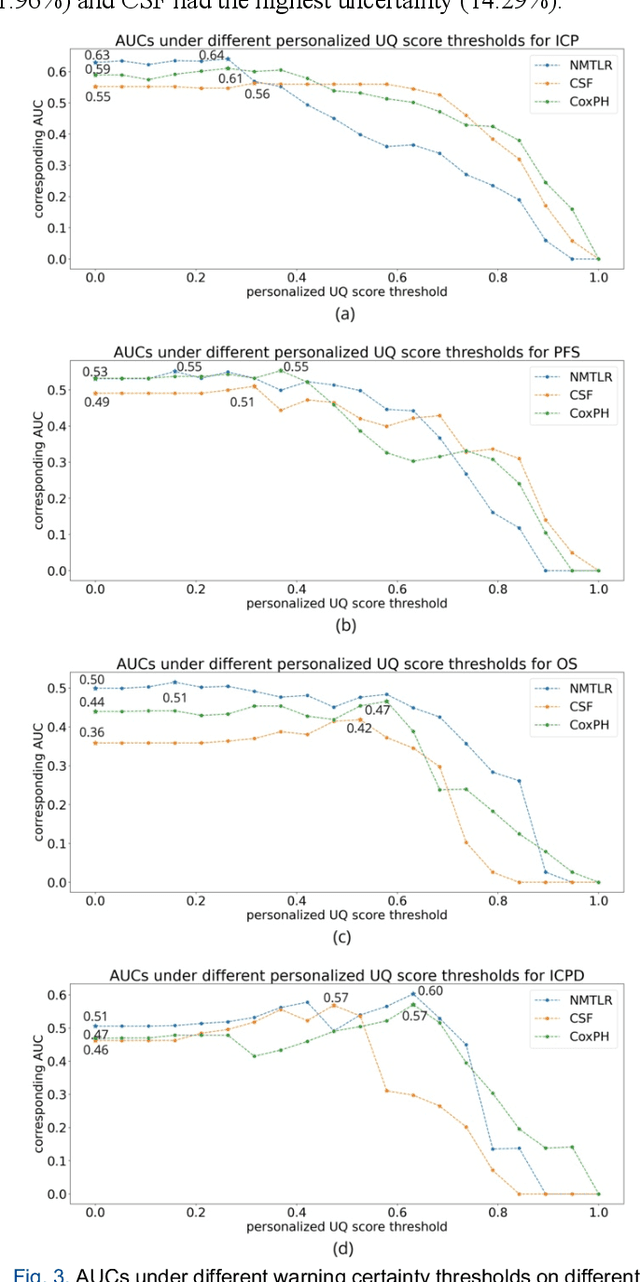
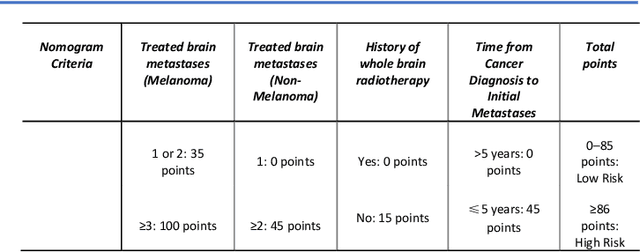
Abstract:TodevelopanovelUncertaintyQuantification (UQ) framework to estimate the uncertainty of patient survival models in the absence of ground truth, we developed and evaluated our approach based on a dataset of 1383 patients treated with stereotactic radiosurgery (SRS) for brain metastases between January 2015 and December 2020. Our motivating hypothesis is that a time-to-event prediction of a test patient on inference is more certain given a higher feature-space-similarity to patients in the training set. Therefore, the uncertainty for a particular patient-of-interest is represented by the concordance index between a patient similarity rank and a prediction similarity rank. Model uncertainty was defined as the increased percentage of the max uncertainty-constrained-AUC compared to the model AUC. We evaluated our method on multiple clinically-relevant endpoints, including time to intracranial progression (ICP), progression-free survival (PFS) after SRS, overall survival (OS), and time to ICP and/or death (ICPD), on a variety of both statistical and non-statistical models, including CoxPH, conditional survival forest (CSF), and neural multi-task linear regression (NMTLR). Our results show that all models had the lowest uncertainty on ICP (2.21%) and the highest uncertainty (17.28%) on ICPD. OS models demonstrated high variation in uncertainty performance, where NMTLR had the lowest uncertainty(1.96%)and CSF had the highest uncertainty (14.29%). In conclusion, our method can estimate the uncertainty of individual patient survival modeling results. As expected, our data empirically demonstrate that as model uncertainty measured via our technique increases, the similarity between a feature-space and its predicted outcome decreases.
Quantifying U-Net Uncertainty in Multi-Parametric MRI-based Glioma Segmentation by Spherical Image Projection
Oct 12, 2022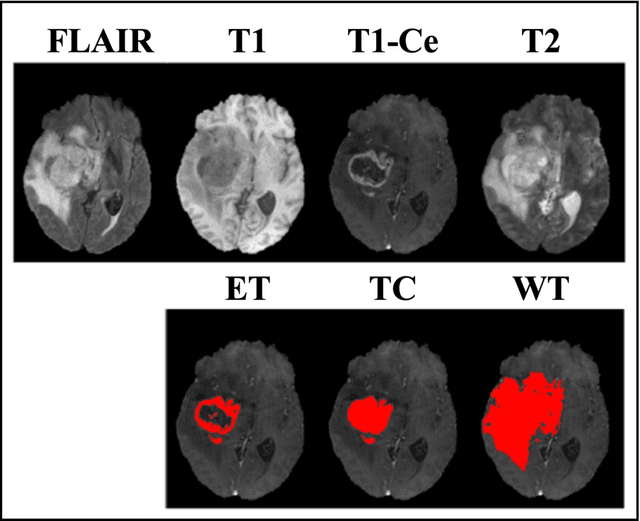
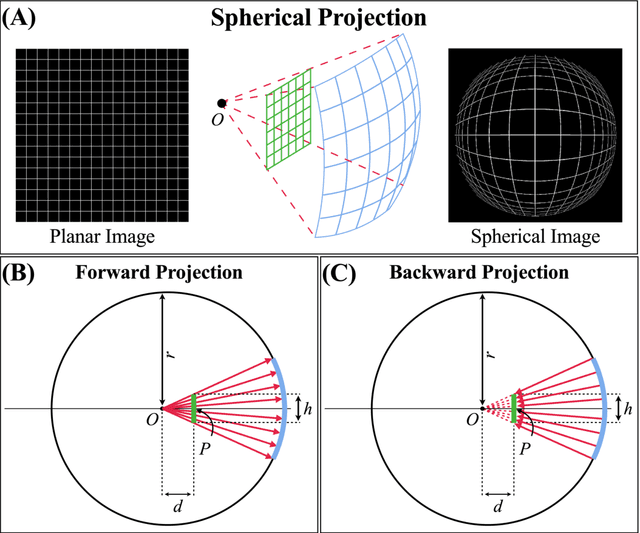
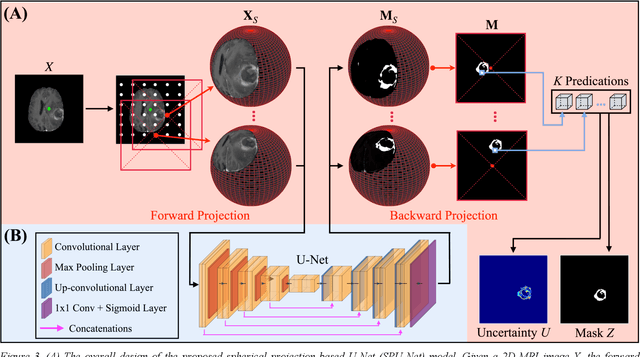
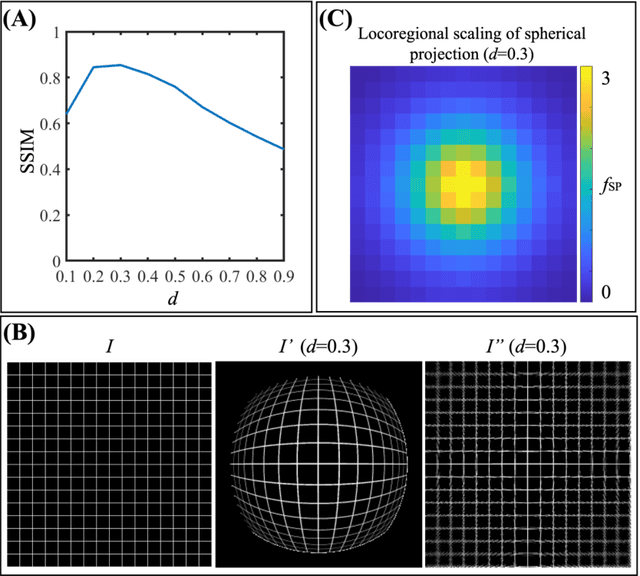
Abstract:Purpose: To develop a U-Net segmentation uncertainty quantification method based on spherical image projection of multi-parametric MRI (MP-MRI) in glioma segmentation. Methods: The projection of planar MRI onto a spherical surface retains global anatomical information. By incorporating such image transformation in our proposed spherical projection-based U-Net (SPU-Net) segmentation model design, multiple segmentation predictions can be obtained for a single MRI. The final segmentation is the average of all predictions, and the variation can be shown as an uncertainty map. An uncertainty score was introduced to compare the uncertainty measurements' performance. The SPU-Net model was implemented on 369 glioma patients with MP-MRI scans. Three SPU-Nets were trained to segment enhancing tumor (ET), tumor core (TC), and whole tumor (WT), respectively. The SPU-Net was compared with (1) classic U-Net with test-time augmentation (TTA) and (2) linear scaling-based U-Net (LSU-Net) in both segmentation accuracy (Dice coefficient) and uncertainty (uncertainty map and uncertainty score). Results: The SPU-Net achieved low uncertainty for correct segmentation predictions (e.g., tumor interior or healthy tissue interior) and high uncertainty for incorrect results (e.g., tumor boundaries). This model could allow the identification of missed tumor targets or segmentation errors in U-Net. The SPU-Net achieved the highest uncertainty scores for 3 targets (ET/TC/WT): 0.826/0.848/0.936, compared to 0.784/0.643/0.872 for the U-Net with TTA and 0.743/0.702/0.876 for the LSU-Net. The SPU-Net also achieved statistically significantly higher Dice coefficients. Conclusion: The SPU-Net offers a powerful tool to quantify glioma segmentation uncertainty while improving segmentation accuracy. The proposed method can be generalized to other medical image-related deep-learning applications for uncertainty evaluation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge